Subfamily: Betarhabdovirinae
Genus: Gammacytorhabdovirus
Distinguishing features
Historically, the genera Nucleorhabdovirus and Cytorhabdovirus were established based on the sites of virus replication and morphogenesis, with cytorhabdoviruses replicating and maturing in the cytoplasm of infected cells. Recently, a reclassification became necessary as phylogenetic analyses of new plant rhabdovirus genomes consistently showed that cytorhabdoviruses did not form a monophyletic clade upon analysis of complete L protein sequence alignments (Rubino et al., 2025). Based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L protein sequences, viruses classified in the genus Gammacytorhabdovirus form a monophyletic cluster clearly distinguished from alpha- and betacytorhabdoviruses, as well as from other plant rhabdoviruses.
Virion
Morphology
Virions are likely to be similar to those reported for alpha- and betacytorhabdoviruses.
Nucleic acid
The negative-sense, single-stranded RNA genome of 10.7–12 kb is unsegmented.
Proteins
N, P, and L represent the three canonical structural proteins for gammacytorhabdoviruses.
Lipids
The lipoprotein envelope is derived from the host plant (Jackson et al., 2005a). Lipid composition is unknown.
Genome organisation and replication
All gammacyorhabdoviruses encode the canonical proteins N, P and L, and lack the G gene, while some of them also lack the M gene. All but two gammacytorhabdoviruses encode a movement protein (P3) and two of them harbor an additional gene between the M and L genes.
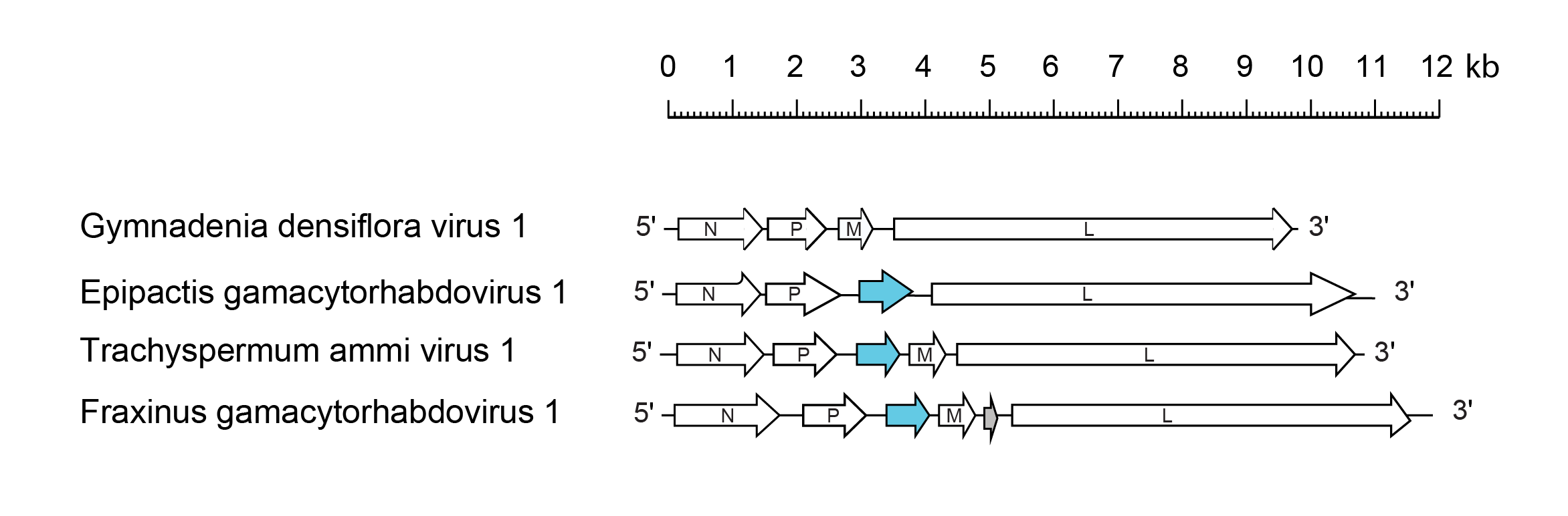 |
| Figure 1 Gammacytorhabdovirus. Schematic representation of a selection of gammacytorhabdovirus genomes shown in reverse (positive-sense) polarity. N, P, M, G and L represent ORFs encoding the structural proteins. ORFs encoding viral cell-to-cell movement proteins (blue) is shown. Other ORF encode a putative accessory proteins of unknown function (grey). In addition to those virus genomes shown: Argyranthemum gammacytorhabdovirus 1, carrot gammacytorhabdovirus 1, celery gammacytorhabdovirus 1, Coptis gammacytorhabdovirus 1, Cuscuta gammacytorhabdovirus 1, Cuscuta gammacytorhabdovirus 2, Cypripedium gammacytorhabdovirus 1, Heliosperma gammacytorhabdovirus 1, Hibiscus gammacytorhabdovirus 1, Lonas gammacytorhabdovirus 1, Lupinus gammacytorhabdovirus 1 and Silene gammacytorhabdovirus 1 are similar to Trachyspermum ammi virus 1; Fraxinus gammacytorhabdovirus 2 is similar to Fraxinus gammacytorhabdovirus 1; and Rhopalocnemis gammacytorhabdovirus 1 is similar to Epipactis gammacytorhabdovirus 1. |
Biology
Gammacytorhabdoviruses infect a wide range of monocot and dicot plants, and one was identified in a fungi; thus gammacytorhabdoviruses could be transmitted by fungal vectors (Bejerman et al., 20234).
Species demarcation criteria
Viruses assigned to different species within the genus Gammacytorhabdovirus have several of the following characteristics: A) nucleotide sequence identity less than 75% for the coding-complete genome sequence; B) amino acid sequence identity less than 86% in proteins encoded by all the cognate open reading frames; and C) they occupy different ecological niches as evidenced by differences in hosts and/or arthropod vectors.
Related, unclassified viruses
| Virus name | Accession number | Virus abbreviation |
| apple gammacytorhabdovirus 1 | PV979719 | AplGCRV1 |
| Bergerden gammacytorhabdovirus 4 | PX121446 | BerGCRV1 |
| Passiflora cytorhabdovirus | PV460965 | PaCRV |
Virus names and virus abbreviations are not official ICTV designations.

