Subfamily: Alpharhabdovirinae
Genus: Tibrovirus
Distinguishing features
Viruses assigned to the genus Tibrovirus form a distinct monophyletic group based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L sequences. Several viruses assigned to the genus have been isolated from either biting midges (Culicoides spp.) or cattle; several other tibroviruses have been detected in humans. The tibrovirus clade is part of a larger phylogenetic group of arthropod-borne rhabdoviruses with large and complex genomes that also includes ephemeroviruses, hapaviruses and curioviruses. Tibroviruses share the common features of: i) two additional genes (U1 and U2) located between the M gene and G gene; and ii) an additional gene (U3), between the G gene and L gene, encoding a class 1a viroporin-like protein.
Virion
Morphology
Bullet-shaped virions of varying morphology have been reported for tibroviruses. Long narrow particles (375 nm × 50 nm) were observed by negative-stain of freshly prepared cell culture supernates of Tibrogargan virus (TIBV; species Tibrovirus tibrogargan) but similar samples stored for 1 week at 4 oC revealed shorter tapered bullet-shaped particles (125 nm × 75 nm) (Cybinski et al., 1980). Long bullet-shaped particles (250 nm × 70 nm) have also been observed in ultrathin sections of mosquito cells infected with Bivens Arm virus (BAV; species Tibrovirus tibrogargan) (Gibbs et al., 1989).
Nucleic acid
Tibrovirus genomes consist of a single molecule of negative-sense, single-stranded RNA and range from approximately 12.5–13.3 kb (Gubala et al., 2011, Grard et al., 2012, Stremlau et al., 2015, Walker et al., 2015, Wiley et al., 2017, Edridge et al., 2022).
Proteins
The N, P, M, G and L share sequence homology and/or structural characteristics with the cognate proteins of other rhabdoviruses (Gubala et al., 2011, Walker et al., 2015). The U1 genes encode proteins of unknown function that range from 170 to 216 amino acids (19.1–23.8 kDa); the U2 genes encode proteins of unknown function that range from 149 to 176 amino acids (17.1–20.1 kDa); the small viroporin-like proteins encoded in the U3 genes range from 92 to 125 amino acids (10.5–13.9 kDa) and feature an N-terminal domain containing large hydrophobic residues, a predicted hydrophobic transmembrane domain and a C-terminal domain that is rich in basic residues.
Genome organisation and replication
Tibrovirus genomes include five genes (N, P, M, G and L) encoding the structural proteins and multiple additional long ORFs including: i) two additional genes between the M and G genes encoding proteins of unknown function (U1 and U2); and ii) an additional gene between the G and L genes encoding a class 1a viroporin-like protein (U1) (Figure 1 Tibrovirus). All ORFs occur in discrete transcriptional units including consensus transcription initiation and transcription termination/polyadenylation sequences (Gubala et al., 2011, Walker et al., 2015).
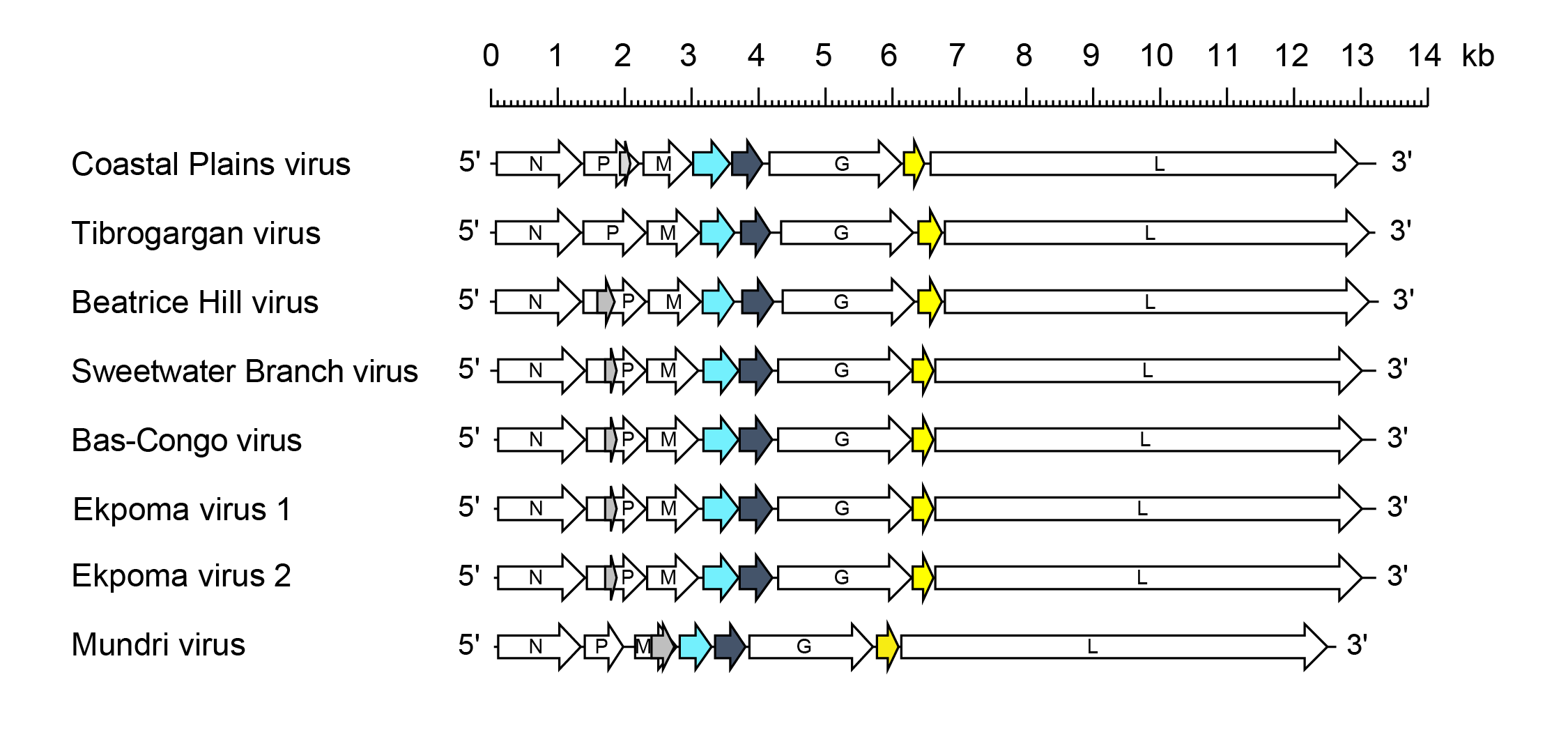 |
| Figure 1 Tibrovirus. Schematic representation of tibrovirus genomes shown in reverse (positive-sense) polarity. N, P, M, G and L represent ORFs encoding the structural proteins. ORF U1 (light blue), ORF U2 (dark grey) and ORF U3 (yellow) are highlighted. ORF U1 and ORF U2 encode small proteins of unknown function and ORF U3 encodes a small viroporin-like transmembrane protein. Alternative ORFs (≥180 nt) also occur in the P genes of most tibroviruses (grey) but it is not known if they are expressed. |
Biology
Several tibroviruses (TIBV, Coastal Plains virus [CPV; species Tibrovirus coastal], BAV, Sweetwater Branch virus [SWBV; species Tibrovirus sweetwater], Beatrice Hill virus [BHV; species Tibrovirus beatrice]) infect cattle and/or water buffalo and have been isolated from biting midges (Culicoides spp.). Tibrovirus infection in cattle has been reported in Australia, Papua New Guinea, the USA and the Caribbean (Cybinski et al., 1980, Cybinski and Gard 1986, Gibbs et al., 1989). There has been no report of disease in ruminants associated with tibrovirus infection. Several other tibroviruses (Bas-Congo virus (BASV; species Tibrovirus congo), Ekpoma virus 1 (EKV1; species Tibrovirus alphaekpoma) and Ekpoma virus 2 (EKV2; species Tibrovirus betaekpoma) have been identified in human sera by next-generation sequencing but infectious virus isolates have not been recovered. BASV was detected in an acute serum sample from the lone survivor of an outbreak of haemorrhagic fever in the Democratic Republic of Congo (Grard et al., 2012). EKV1 and EKV2 were detected in blood taken from apparently healthy individuals in Nigeria (Stremlau et al., 2015). Mundri virus (MUNV; species Tibrovirus mundri) was detected in the plasma and cerebrospinal fluid of a 15 year-old female human with new-onset nodding syndrome in South Sudan (Edridge et al., 2022). The role of tibroviruses in human disease is not currently clear.
Antigenicity
TIBV, CPV, BAV and SWBV partially cross-react in indirect immunofluorescence, complement-fixation and/or neutralisation tests (Cybinski et al., 1980, Cybinski and Gard 1986, Gibbs et al., 1989).
Species demarcation criteria
Viruses assigned to different species within the genus Tibrovirus have several of the following characteristics: A) minimum amino acid sequence divergence of 20% in the L proteins; B) minimum amino acid sequence divergence of 20% in the N proteins; C) can be distinguished in serological tests; and D) occupy different ecological niches as evidenced by differences in hosts and/or arthropod vectors.

