Subfamily: Alpharhabdovirinae
Genus: Caligrhavirus
Distinguishing features
Viruses assigned to the genus Caligrhavirus form a distinct monophyletic group based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L sequences. Viruses assigned to the genus have been detected in sea lice (Copepoda: Caligidae).
Virion
Morphology
For Lepeophtheirus salmonis rhabdovirus 127 (species Caligrhavirus lepeophtheirus), virus-like particles (bacilliform or bullet-shaped, 55 nm diameter and up to 425 nm in length) have been observed by transmission electron microscopy in thin sections of necrotic tissues of salmon louse samples; cross-striated nucleocapsids with helical symmetry were also observed (Økland et al., 2014).
Nucleic acid
Caligrhavirus genomes consist of a single molecule of negative-sense, single-stranded RNA and range from approximately 11.5–11.7 kb (Økland et al., 2014, Økland et al., 2018).
Proteins
N, P, M, G and L share sequence homology and/or structural characteristics with the cognate proteins of other rhabdoviruses. Caligus rogercresseyi rhabdovirus (CRogRV; species Caligrhavirus caligus) ORF U1 encodes a putative 15.3 kDa acidic protein of unknown function which has not yet been identified in infected cells (Økland et al., 2018).
Genome organisation and replication
Caligrhavirus genomes include five genes (N, P, M, G and L) encoding the structural proteins and concise intergenic regions (Figure 1 Caligrhavirus). In CRogRV, there is also an additional gene (U1) between the M and G genes which occurs as an independent transcriptional unit including consensus transcription initiation and transcription termination/polyadenylation sequences (Økland et al., 2014, Økland et al., 2018).
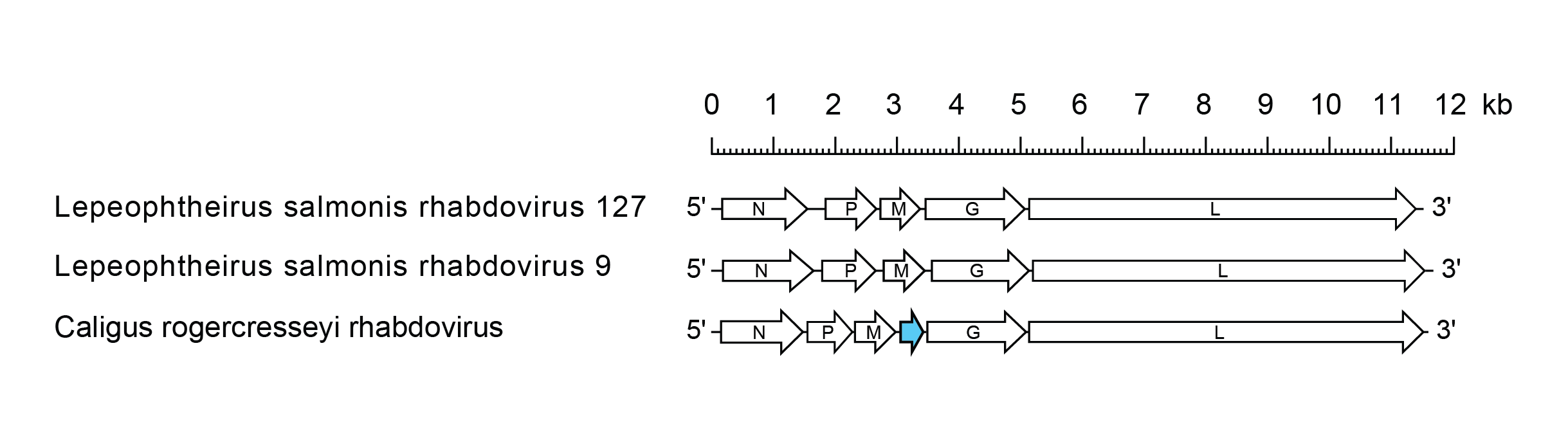 |
| Figure 1 Caligrhavirus. Schematic representation of caligrhavirus genomes shown in reverse (positive-sense) polarity. N, P, M, G and L represent ORFs encoding the structural proteins. ORF U1 (blue) of Caligus rogercresseyi rhabdovirus, encoding a protein of unknown function, is highlighted. |
Biology
Caligrhaviruses have been detected in sea lice in which they appear to cause active infections (Økland et al., 2014, Økland et al., 2018). Sea lice are crustaceans in the family Caligidae (Copepoda: Siphonostomatoida) and are marine ectoparasites that feed on the blood and tissues of marine fish. Each of the current members of the genus has been detected in sea lice infesting farmed Atlantic salmon (Salmo salar) from either Norway or Chile (Økland et al., 2014, Økland et al., 2018). No isolates are yet available for any of these viruses but they have been detected in necrotic tissues of salmon lice by electron microscopy, polymerase chain reaction or in situ hybridization (Økland et al., 2014, Økland et al., 2018).
Species demarcation criteria
Viruses assigned to different species within the genus have several of the following characteristics: A) minimum amino acid sequence divergence of 15% in the N proteins; B) minimum amino acid sequence divergence of 20% in the L proteins; C) minimum amino acid sequence divergence of 20% in the G proteins; D) significant differences in genome organization as evidenced by numbers and locations of ORFs; E) can be distinguished in virus neutralisation tests; and F) occupy different ecological niches as evidenced by differences in hosts and/or vectors.

