Subfamily: Betarhabdovirinae
Genus: Betanucleorhabdovirus
Distinguishing features
Historically, the genera Nucleorhabdovirus and Cytorhabdovirus were established based on the sites of virus replication and morphogenesis, with nucleorhabdoviruses replicating and maturing in the nuclei of infected cells. Recently, a reclassification became necessary as phylogenetic analyses of new plant rhabdovirus genomes consistently showed that nucleorhabdoviruses did not form a monophyletic clade upon analysis of complete L protein sequence alignments (Kuhn et al., 2020).
Based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L protein sequences, viruses classified in the genus Betanucleorhabdovirus form a monophyletic cluster clearly distinguished from alpha-, delta- and gammanucleorhabdoviruses and from dichorhaviruses, as well as from other plant rhabdoviruses.
Virion
Morphology
Enveloped virions are bacilliform, 45–100 nm in diameter and 130–300 nm long (Goodin and Jackson 2002, Jackson et al., 2005a).
Physicochemical and physical properties
Virus particles sediment at 800–1000 S in sucrose gradients and the buoyant density of virions is 1.18 g cm−3 in isopycnic sucrose gradients (Goodin and Jackson 2002).
Nucleic acid
The negative-sense, single-stranded RNA genome of 12.9–14.4 kb is unsegmented. Six to seven mRNAs, one for each of the encoded proteins identified in infected plants.
Proteins
N, P, M, G and L represent the five canonical rhabdovirus structural proteins. Sonchus yellow net virus (SYNV; species Betanucleorhabdovirus retesonchi) P has RNA silencing suppressor activity (Jackson et al., 2005a, Guo et al., 2013). The SYNV P3 protein has been shown by reverse genetics to facilitate cell-to-cell movement between plant cells (Wang et al., 2015); the P3 proteins of other nucleorhabdoviruses are thought to be movement proteins (Jackson et al., 2005a, Bandyopadhyay et al., 2010, Pappi et al., 2013).
SYNV N and P contain nuclear localisation sequences (NLS, or a karyophillic domain) and are independently imported into the nucleus, where they associate and move to a sub-nuclear location (Goodin et al., 2001). A distinct nuclear polymerase complex composed of N, P and L is present in the nuclei of infected cells (Wagner et al., 1996, Wagner and Jackson 1997, Martins et al., 1998).
Highly conserved regulatory regions separating the genes were identified in all betanucleorhabdoviruses, and all deduced viral protein sequences contain predicted mono- or bipartite nuclear localization signals, indicative of at least partial nuclear localization.
Lipids
The lipoprotein envelope is derived from the host plant or insect vector (Jackson et al., 2005a). Lipid composition is unknown.
Genome organisation and replication
The SYNV genome (13.7 kb) contains six genes in the order 3′-N-P-P3-M-G-L-5′. P3 (sc4) in SYNV has been shown to encode a protein involved in cell-to-cell movement. The SYNV 144 nt 3′-leader sequence is transcribed to produce a polyadenylated leader RNA, which localizes in the cytoplasm. The 5′-trailer RNA of 160 nt has extensive terminal complementarity with the leader sequence.
The genomes of Datura yellow vein virus (DYVV; species Betanucleorhabdovirus daturae), sowthistle yellow vein virus (SYVV; species Betanucleorhabdovirus venasonchi), blackcurrant-associated rhabdovirus (BCaRV; species Betanucleorhabdovirus ribes), Asclepias syriaca virus 2 (AscSyV2; species Betanucleorhabdovirus asclepiadis), Bacopa monnieri virus 2 (BmV2; species Betanucleorhabdovirus bacopae), cardamom vein clearing virus (CdVCV; species Betanucleorhabdovirus cardamomi), Cnidium virus 1 (CnV1; species Betanucleorhabdovirus cnidii), Plectranthus aromaticus virus 1 (PleArV1; species Betanucleorhabdovirus plectranthi), Picris betanucleorhabdovirus (PBRV1; species Betanucleorhabdovirus picridis), Rhododendon delavayi virus 1 (RhoDeV1; species Betanucleorhabdovirus rhododendri), Taraxacum betanucleorhabdovirus (TarBRV1; species Betanucleorhabdovirus taraxaci), tomato betanucleorhabdovirus 1 (TBRV1; species Betanucleorhabdovirus alphalycopersici), tomato betanucleorhabdovirus 1 (TBRV2; species Betaanucleorhabdovirus betalycopersici), Zhuye pepper nucleorhabdovirus (ZPNRV; species Betanucleorhabdovirus zanthoxyli) and bird’s-foot trefoil-associated virus 1 (BFTV1; species Betanucleorhabdovirus loti) have an organization similar to that of SYNV, whereas alfalfa-associated nucleorhabdovirus (AaNV; species Betanucleorhabdovirus medicagonis) and apple rootstock virus A (ApRVA; species Betanucleorhabdovirus mali) seem to harbour an additional ORF of unknown function (U) between the M and G genes (Figure 1 Betanucleorhabdovirus).
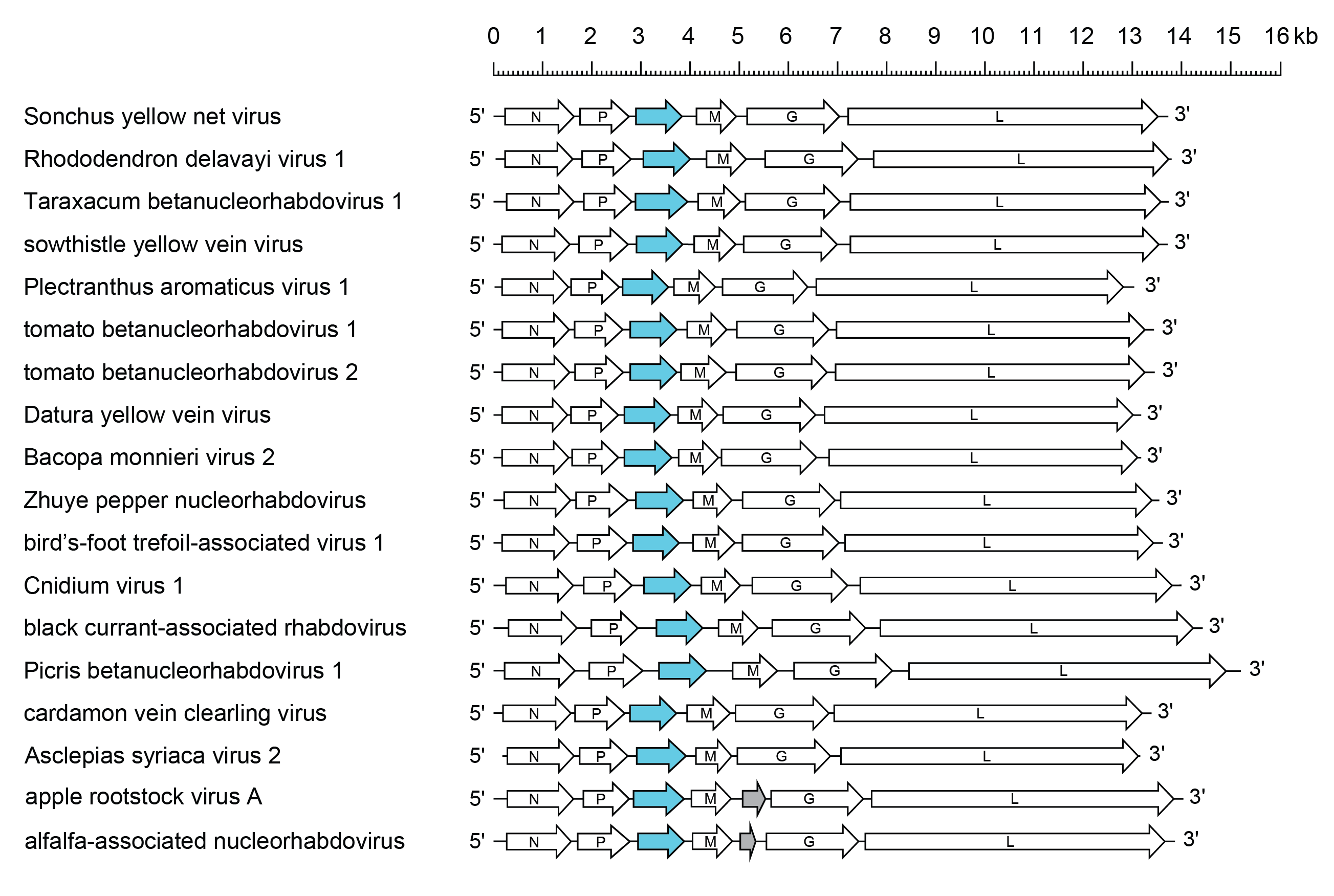 |
| Figure 1 Betanucleorhabdovirus. Schematic representation of a selection of betanucleorhabdovirus genomes shown in reverse (positive-sense) polarity. N, P, M, G and L represent ORFs encoding the structural proteins. ORFs encoding putative cell-to-cell movement proteins are highlighted (blue). Alfalfa-associated nucleorhabdovirus and apple rootstock virus A encode small proteins of unknown function between the M gene and G gene (grey). In addition to those genomes shown: Alpinia vein streaking virus, beet betanucleorhabdovirus 1, Paris yunnanensis rhabdovirus 1, and Sambucus betanucleorhabdoviruses 1–5 are similar to Sonchus yellow net virus. |
Betanucleorhabdoviruses replicate in the nuclei of plant cells, which become greatly enlarged and develop large granular nuclear inclusions that are thought to be sites of virus replication (Jackson et al., 2005a). In situ hybridisation analyses have shown that the viral genomic and anti-genomic RNAs of SYNV are highly expressed in subnuclear foci (Martins et al., 1998, Jackson et al., 2005a) and immunofluorescence studies have shown that the ribonucleoprotein (RNP) components N, P and L also accumulate in subnuclear foci (Martins et al., 1998, Deng et al., 2007). Viral proteins are synthesised from discrete polyadenylated mRNAs and reporter gene analyses have shown that they accumulate in subnuclear foci (Wagner and Jackson 1997, Martins et al., 1998). Virus morphogenesis occurs at the inner nuclear membrane and enveloped virus particles accumulate in perinuclear spaces (Martins et al., 1998, Jackson et al., 2005a). In protoplasts treated with the glycosylation inhibitor tunicamycin, morphogenesis is interrupted and nucleocapsids accumulate in the nucleoplasm (Jones and Jackson 1990).
Biology
A variety of dicot plants are susceptible to betanucleorhabdoviruses although each virus usually has a restricted host range (Goodin and Jackson 2002, Jackson et al., 2005a). SYNV and SYVV are transmitted by aphids, but no arthropod vectors have so far been identified for the other betanucleorhabdoviruses. SYNV and AaNV can also be transmitted mechanically from infected sap to Nicotiana benthamiana. In all carefully examined cases, viruses replicate in cells of the insect vector as well as in the plant host (Jackson et al., 2005a).
Species demarcation criteria
Viruses assigned to different species within the genus Betanucleorhabdovirus have several of the following characteristics: A) the nucleotide sequence identity of complete genomes is less than 75%; B) they occupy different ecological niches as evidenced by differences in hosts and/or arthropod vectors; and C) they can be clearly distinguished in serological tests or by nucleic acid hybridisation.
Betanucleorhabdovirus species are primarily differentiated by plant host range and vector specificity of the virus. Nucleic acid hybridisation has been used to provide confirmation of identification and serological criteria have enabled verification of common viruses that infect different hosts. However, no betanucleorhabdovirus species have been defined unambiguously using serology. Complete or coding-complete genome nucleotide sequences are available for all 14 viruses currently assigned to the genus. RT-PCR-based assays and fluorescent viral protein localisation have proven to be useful tools for species demarcation. Hybridisation using cloned probes and RT-PCR has been used to detect the viruses.
Related, unclassified viruses
| Virus name | Accession number | Virus abbreviation |
| Alfalfa nucleorhabdovirus 1 | OK514705* | AlNCRV1 |
| Artemisia betanucleorhabdovirus 1 | BK070483 | ArtBNRV1 |
| Artemisia betanucleorhabdovirus 2 | PX121438 | ArtBNRV2 |
| Boehmeria betanucleorhabdovirus 1 | BK070484 | BoeBNRV1 |
| Camellia betanucleorhabdovirus 1 | BK070485 | CamBNRV1 |
| canola betanucleorhabdovirus 1 | BK070486 | CanBNRV1 |
| Cardamine betanucleorhabdovirus 1 | BK070487 | CarBNRV1 |
| Chrysanthemum betanucleorhabdovirus 1 | BK070488 | ChrBNRV1 |
| Coptis betanucleorhabdovirus 1 | BK070489 | CopBNRV1 |
| Dendrobium betanucleorhabdovirus 1 | BK070490 | DenBNRV1 |
| dicot associated betanucleorhabdovirus 1 | BK070491 | DaBNRV1 |
| Eichornia betanucleorhabdovirus 1 | BK070492 | EicBNRV1 |
| Lysimachia betanucleorhabdovirus 1 | BK070493 | LysBNRV1 |
| Mentha betanucleorhabdovirus 1 | BK070494 | MenBNRV1 |
| Nekemias betanucleorhabdovirus 1 | BK070495 | NekBNRV1 |
| Nymphaea betanucleorhabdovirus 1 | BK070496 | NymBNRV1 |
| Osmolindsaea betanucleorhabdovirus 1 | BK070497 | OsmBNRV1 |
| Paedaria scandens nucleorhabdovirus | PV649689 | PaScNRV |
| Rhododendron betanucleorhabdovirus 2 | BK070498 | RhoBNRV1 |
| Ricotia betanucleorhabdovirus 1 | BK070499 | RicBNRV1 |
| Rorippa betanucleorhabdovirus 1 | BK070500 | RorBNRV1 |
| Salix betanucleorhabdovirus 1 | BK070501 | SalBNRV1 |
| Salvia betanucleorhabdovirus 1 | BK070502 | SalvBNRV1 |
| Stratiotes betanucleorhabdovirus 1 | BK070503 | StrBNRV1 |
| strawberry virus 5 | PV933990 | StrV5 |
| Thymus betanucleorhabdovirus 1 | BK070504 | ThyBNRV1 |
| Trichosanthes betanucleorhabdovirus 1 | BK070505 | TriBNRV1 |
| Viola betanucleorhabdovirus 1 | BK070506 | VioBNRV1 |
| Ziziphora betanucleorhabdovirus 1; | BK070507 | ZizBNRV1 |
Virus names and virus abbreviations are not official ICTV designations.
* Coding region sequence incomplete

