Family: Rhabdoviridae
Genus: Betaplatrhavirus
Distinguishing features
Viruses assigned to the genus Betaplatrhavirus form a distinct monophyletic group based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L sequences. Members of the genus have been detected in flatworms (phylum Platyhelminthes) or vertebrate tissue samples. They are distinct phylogenetically from rhabdoviruses assigned to the genera Alphaplatrhavirus and Gammaplatrhavirus.
Morphology
Viruses assigned to the genus have not yet been isolated or visualized by electron microscopy.
Nucleic acid
Betaplatrhavirus genomes consist of a single molecule of negative-sense, single-stranded RNA and range from approximately 9.9–12.6 kb (Shi et al., 2018, Dheilly et al., 2022, Gorbushin 2023).
Proteins
Betaplatrhavirus N, P, M, G and L proteins appear to be homologous with corresponding proteins of other rhabdoviruses but betaplatrhaviruses commonly lack the G protein. Betaplartrhavirus G proteins are class I transmembrane glycoproteins. Alignment of betaplatrhavirus G proteins with that of vesicular stomatitis Indiana virus (species Vesiculovirus indiana) indicates likely preservation of all 12 conserved cysteine residues that are typical of animal rhabdovirus G proteins (Walker and Kongsuwan 1999, Roche et al., 2006), and 2 additional conserved cysteine residues that are likely to form another disulphide bridge in the folded protein. Betaplatrhaviruses sometimes encode additional proteins.
Genome organisation and replication
Betaplatrhavirus genomes include the four genes (N, P, M and L) encoding the structural proteins, and sometimes also include the G gene (Figure 1 Betaplatrhavirus). An additional gene encoding a protein of unknown function occurs between G and L genes of Eptesicud fuscus rhabdovirus (species Betaplatrhavirus fuscus). An alternative long ORF occurs near the start of the N gene in Fujian dimarhabdovirus (species Betaplatrhavirus fujian) but it is not known if it is expressed in infected cells.
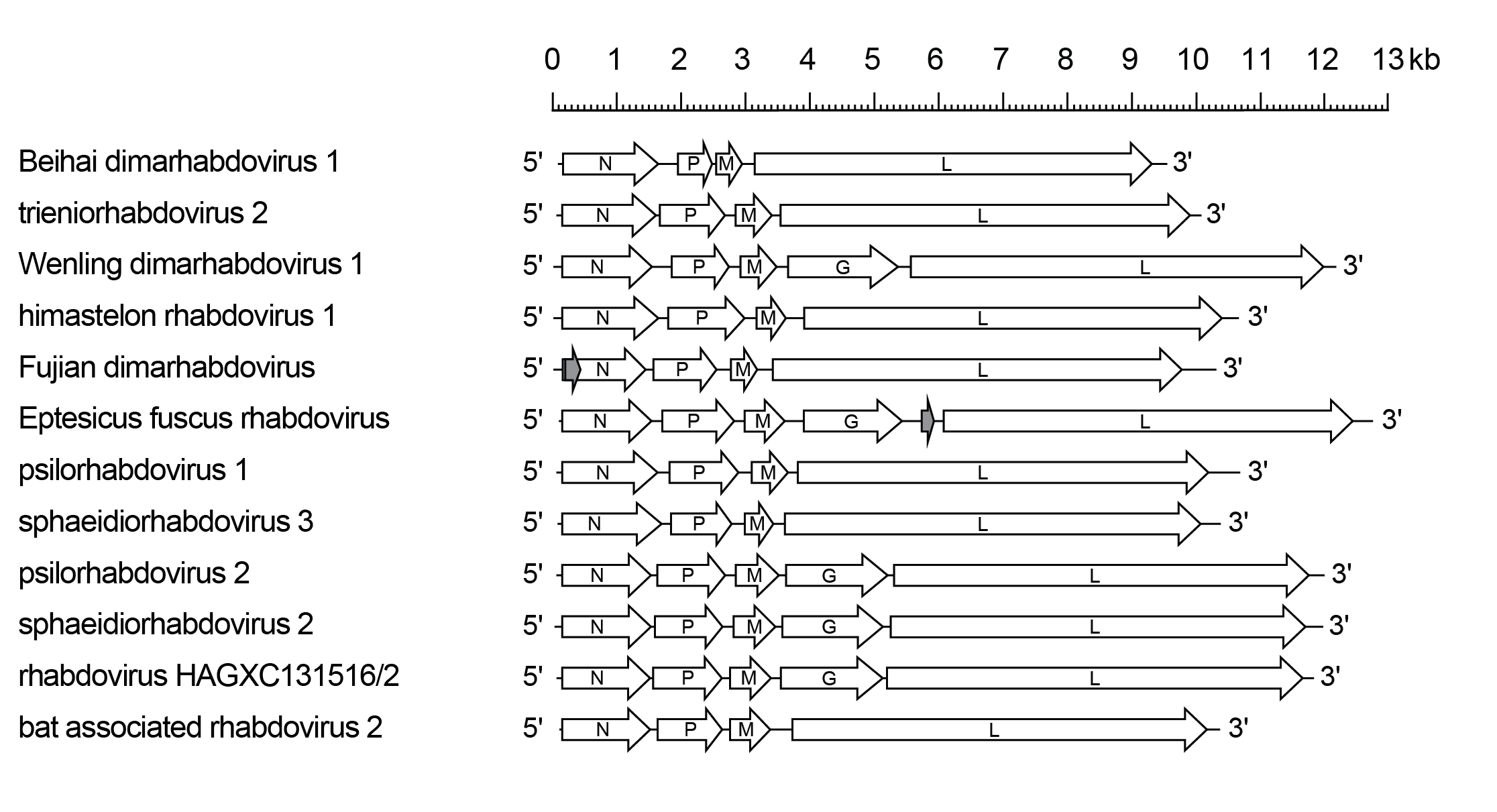 |
| Figure 1 Betaplatrhavirus. Schematic representation of betaplatrhavirus genomes in reverse (positive-sense) polarity. The five long open reading frames (ORFs) in the N, P, M, G and L genes are shown (open arrows). Additional and alternative ORFs are shown in grey. |
Biology
Betaplatrhaviruses have been detected by metagenomic sequencing of platyhelminths that parasitise invertebrate and vertebrate hosts (Dheilly et al., 2022). These include cestode worms (Triaenophorus nodulosus) and trematode worms (Psilotrema simillimum, Sphaeridiotrema pseudoglobulus and Himasthla elongata) (Dheilly et al., 2022, Gorbushin 2023). Other betaplatrhaviruses have been detected by metatranscriptomic sequencing of various visceral organs of bats, newts and fish (Shi et al., 2018). No isolates are currently available for any of these viruses. Available evidence suggests that those betaplatrhaviruses detected in vertebrates are likely due to platyhelminth infestation.
Species demarcation criteria
Viruses assigned to different species within the genus have several of the following characteristics: A) minimum amino acid sequence divergence of 10% in N proteins; B) minimum amino acid sequence divergence of 10% in the L proteins; C) minimum amino acid sequence divergence of 15% in G proteins; D) significant differences in genome organisation as evidenced by numbers and locations of ORFs; E) can be distinguished in virus neutralisation tests; and F) occupy different ecological niches as evidenced by differences in arthropod and/ or vertebrate hosts.

