Subfamily: Alpharhabdovirinae
Genus: Ledantevirus
Distinguishing features
Viruses assigned to the genus Ledantevirus form a distinct monophyletic group based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L sequences. Many of the viruses in the genus have been isolated from bats and many or all appear to be transmitted by arthropods. There are three distinct phylogroups comprising: A) Fukuoka virus (FUKV; species Ledantevirus fukuoka), Barur virus (BARV; species Ledantevirus barur), Nishimuro virus (NISV; species Ledantevirus nishimuro), Nkolbisson virus (NKOV; species Ledantevirus nkolbisson), Longquan Niviventer coninga ledantevirus 1 (LNcoLV1; species Ledantevirus longquan), Tongren rhabd tick virus 2 (TrRTV2; species Ledantevirus tongren) and Yongjia tick virus 2 (YjTV2; species Ledantevirus yongjia); B) Le Dantec virus (LDV; species Ledantevirus ledantec), Keuraliba virus (KEUV; species Ledantevirus keuraliba), Kern Canyon virus (KCV; species Ledantevirus kern) and Vaprio virus (VAPV; species Ledantevirus vaprio); and C) Mount Elgon bat virus (MEBV; species Ledantevirus elgon), Taiyi bat virus (TYBV; species Ledantevirus taiyi); Kumasi rhabdovirus (KRV; species Ladantevirus kumasi), Wenzhou Rhinolophus pusillus ledantevirus 1 (WRpuLV-1; species Ledantevirus wenzhou), Oita virus (OITAV; species Ledantevirus oita), Fikirini rhabdovirus (FKRV; species Ledantevirus fikirini), Kolente virus (KOLEV; species Ledantevirus kolente), Wuhan louse fly virus 5 (WhLFV5; species Ledantevirus wuhan) and Kanyawara virus (KYAV; species Ledantevirus kanyawara). The genus is linked phylogenetically to viruses assigned to the genus Vesiculovirus (infecting mammals).
Virion
Morphology
For all members of the genus that have been characterised morphologically, virions are bullet-shaped. A detailed electron microscopic study of KCV revealed enveloped virions (mean dimensions 132 × 73 nm) in negative-stained preparations with prominent cross-striations, axial channels, and 8 nm surface projections evenly distributed at 7 nm intervals (Murphy and Fields 1967). MEBV displayed similar morphology under negative-stain but virions were longer (mean 230 nm × 70 nm) (Murphy et al., 1970). In ultrathin sections of mouse brain tissue, enveloped bullet-shaped OITAV virions (160–300 nm × 100 nm) have been reported (Iwasaki et al., 2004); enveloped bullet-shaped virions have also been reported from cell supernates or in ultrathin sections of infected tissue for NKOV (130–150 nm × 66 nm), LEDV (163 nm × 78 nm), VAPV, FUKV, NISV and KRV (Cropp et al., 1985, Kaneko et al., 1986, Ndiaye et al., 1987, Binger et al., 2015, Sakai et al., 2015, Lelli et al., 2018). Differences in reported particle dimensions are likely to result from factors such as variations in conditions for fixation and staining for electron microscopy, the presence of defective-interfering particles and end-to-end-fusion of virions.
Nucleic acid
Ledantevirus genomes consist of a single molecule of negative-sense, single-stranded RNA and range from approximately 10.8–11.5 kb.
Proteins
The N, P, M, G and L share sequence homology and/or structural characteristics with the cognate proteins of other rhabdoviruses. Other proteins are encoded in the genome but have not yet been identified in infected cells. The U1 ORFs present in LDV, KEUV, KCV and VAPV encode small basic proteins of 64 to 77 amino acids (7.3–8.8 kDa). The FUKV Mx ORF encodes a predicted double-membrane-spanning protein of 86 amino acids (10.4 kDa) (Blasdell et al., 2015, Walker et al., 2015, Lelli et al., 2018).
Genome organisation and replication
Ledantevirus genome organisations include five genes (N, P, M, G and L) encoding the structural proteins and concise intergenic regions. (Kading et al., 2013, Blasdell et al., 2015, Li et al., 2015, Sakai et al., 2015, Walker et al., 2015, Lelli et al., 2018). In LDV, KEUV, KCV and VAPV genomes, a small additional ORF (U1) between the G and L genes occurs as an independent transcriptional unit including consensus transcription initiation and transcription termination/polyadenylation sequences; in the FUKV genome, an alternative ORF (Mx) encoding a putative double-membrane-spanning protein occurs within the M gene (Figure 1 Ledantevirus).
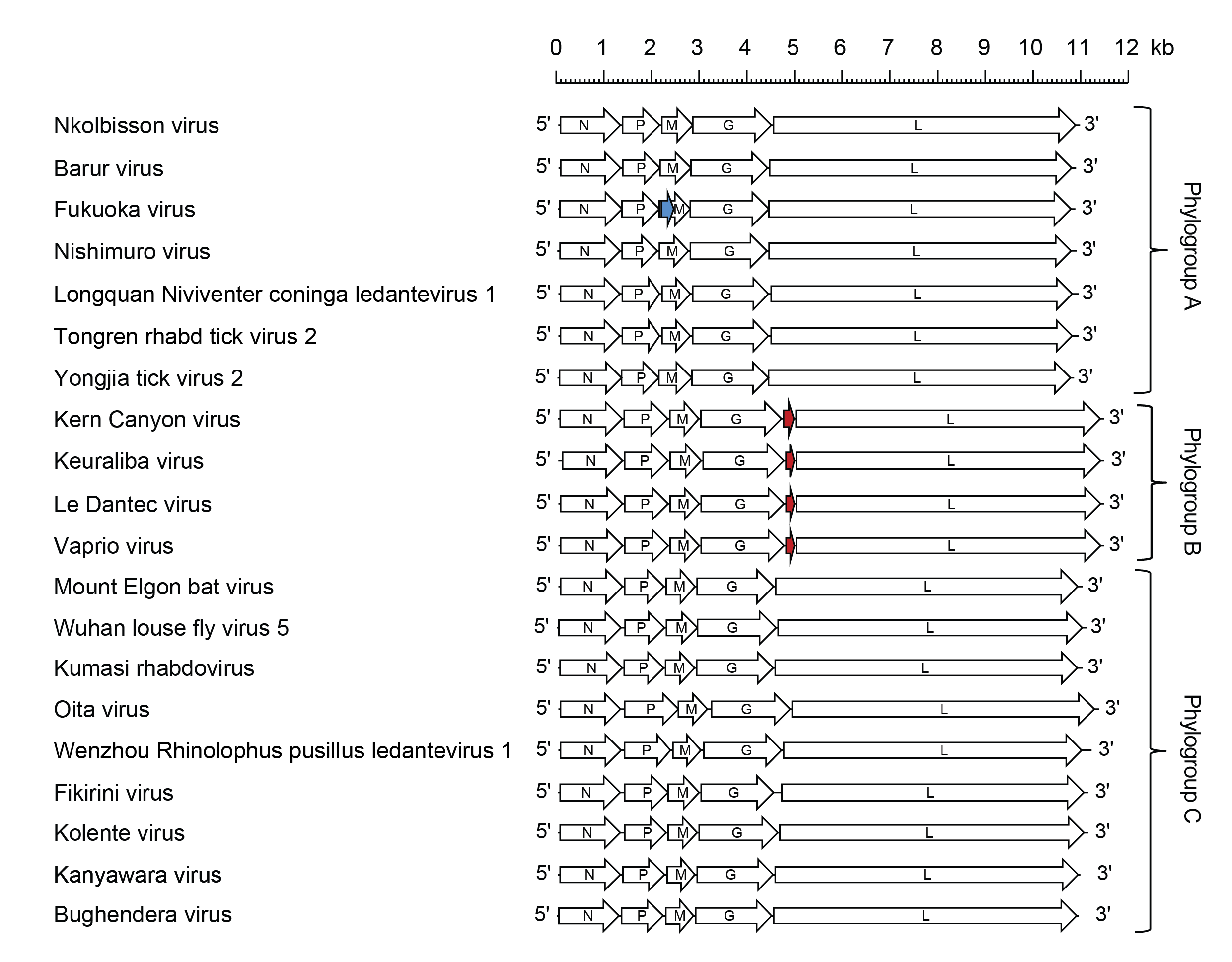 |
| Figure 1 Ledantevirus. Schematic representation of ledantevirus genomes shown in reverse (positive-sense) polarity. N, P, M, G and L represent ORFs encoding the structural proteins. The U1 ORFs of Kern Canyon virus, Keuraliba virus, Le Dantec virus and Vaprio virus (red brown) and the Fukuoka virus Mx ORF (blue) are highlighted. |
Biology
Viruses assigned to the genus infect mammals in Africa, Asia, Europe or North America. Many have been isolated from or detected in bats or arthropods feeding on bats (KCV, VAPV, KOLEV, FKRV, MEBV, TYBV, KRV, OITAV, WhLFV-5, KYAV, WRpuLV-1, LNco LV-1) (Johnson 1965, Murphy and Fields 1967, Metselaar et al., 1969, Iwasaki et al., 2004, Ghedin et al., 2013, Kading et al., 2013, Binger et al., 2015, Li et al., 2015, Goldberg et al., 2017, Wiley et al., 2017, Lelli et al., 2018). Others have been isolated from rodents (BARV, KEUV) or ungulates (FUKV, NISV) (Karabatsos 1985, Kaneko et al., 1986, Sakai et al., 2015). LDV was isolated from humans, initially from a young female with fever and hepatosplenomegaly and subsequently from an adult male with neurological signs; however, a causal relationship has not been established and its natural reservoir(s) and mode(s) of transmission are unknown (Woodruff et al., 1977, Cropp et al., 1985). NKOV has also been isolated from human serum (Ndiaye et al., 1987). Some of the viruses have been isolated from or detected in mosquitoes, midges, fleas, ticks or other arthropods (NKOV, KOLEV, FUKV, BARV, WhLFV5, YjTV-2, TrRTV2) and may be assumed to be arboviruses (Salaun et al., 1969, Johnson et al., 1977, Ndiaye et al., 1987, Ghedin et al., 2013, Li et al., 2015).
Antigenicity
Serological cross-reactions (complement-fixation, indirect fluorescence antibody or ELISA) have been reported between various members of the genus. These include between: LDV and KEUV; KCV, BARV, FUKV, NKOV and KEUV; KOLEV, BARV and FUKV; and FUKV, BARV and MEBV (Tesh et al., 1983, Cropp et al., 1985, Calisher et al., 1989, Ghedin et al., 2013).
Species demarcation criteria
Viruses assigned to different species within the genus Ledantevirus have several of the following characteristics: A) minimum amino acid sequence divergence of 7% in the L proteins; B) minimum amino acid sequence divergence of 15% in the G proteins; C) significant differences in genome organisation as evidenced by numbers and locations of ORFs; D) can be distinguished in serological tests; and E) occupy different ecological niches as evidenced by differences in hosts and/or arthropod vectors.
Related, unclassified viruses
| Virus name | Accession number | Virus abbreviation | |
| Jingmen bat ledantevirus 1 | OQ715726* | JmBLV1 | |
| Ordo virus | OR607590* | ORDOV | |
| Wufeng rodent ledantevirus 1 | OQ715719* | WfRRV1 |
Virus names and virus abbreviations are not official ICTV designations.
* Coding region sequence incomplete

