Subfamily: Alpharhabdovirinae
Genus: Vesiculovirus
Distinguishing features
Viruses assigned to the genus Vesiculovirus form a distinct monophyletic group based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L sequences. Vesiculoviruses are transmitted by insects and infect mammals, birds and reptiles. Some vesiculoviruses can also be transmitted by direct contact. The vesiculovirus clade is part of a larger phylogenetic group of rhabdoviruses with relatively simple genomes that includes spriviviruses and perhabdoviruses infecting fish. Vesiculovirus genomes contain only five structural protein genes (3′-N-P-M-G-L-5′) and short intergenic regions. Some members of the genus also encode two small highly basic proteins (C and C′) in an alternative ORF within the P gene.
Virion
Morphology
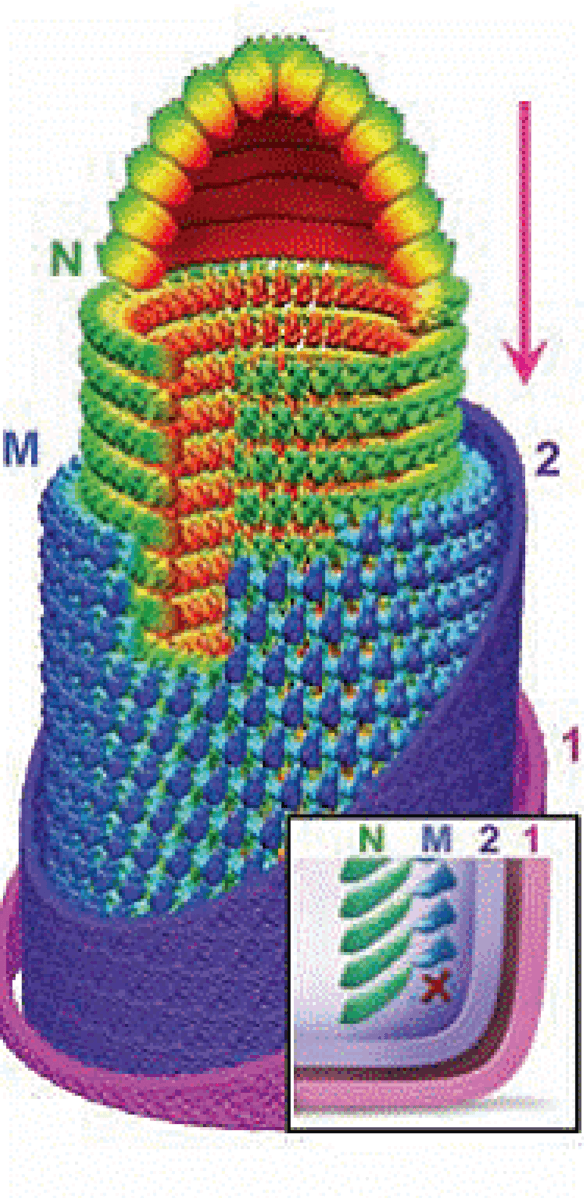
Virons are enveloped and bullet-shaped, approximately 190 nm in length and 85 nm in diameter. The three-dimensional structure of vesicular stomatitis Indiana virus (VSIV; species Vesiculovirus indiana) has been determined utilizing cryo-electron microscopy (Ge et al., 2010) (Figure 1 Vesiculovirus). The nucleocapsid forms a conical-shaped tip and a helical structure in the trunk. Each virion contains an outer helix of matrix protein (M) and an inner helix of nucleoprotein (N) and RNA. M has a hub domain with four contact sites that link to neighbouring M and N subunits, providing rigidity by clamping adjacent turns of the RNP. Interactions among N and M provide scaffolding and stability to the complex and are critical in maintaining the bullet shape. Two layers of lipids obtained from the host cell membrane with embedded trimers of the envelope glycoprotein (G) form the outer layers of the virion and mediate the interaction with cellular receptor(s).
|
| Figure 1 Vesiculovirus. Architecture of the vesicular stomatitis Indiana virus particle. Montage model of the tip and cryo-electron-microscopy map of the trunk. N is green, M is blue, and the inner (2) and outer (1) leaflets of the membrane are violet and pink. (Inset) illustration of the base region of the virion. X marks the absence of a turn of M helix below the lowest turn of the N helix (from: (Ge et al., 2010). Reproduced with permission from AAAS). |
Nucleic acid
Viruses contain a single molecule of linear, negative-sense, single-stranded RNA of approximately 10.7–11.3 kb (Walker et al., 2015). Defective-interfering RNAs are usually significantly shorter than full-length RNA and can be found packaged into small virions known as defective interfering (DI) particles (Huang et al., 1966).
Proteins
Vesiculoviruses have five structural proteins (designated N, P, M, G and L) that are common to all rhabdoviruses. For VSIV, the numbers of molecules per infectious virus particle have been estimated as: L (20–50); G (500–1,500); N (1,000–2,000); P (100–500); and M (1,500–4,000) (Cartwright et al., 1972, Thomas et al., 1985). Several vesiculoviruses (VSIV, vesicular stomatitis New Jersey virus (VSNJV; species Vesiculovirus newjersey), Jurona virus (JURV; species Vesiculovirus jurona), Cocal virus (COCV; species Vesiculovirus cocal) and Chandipura virus (CHPV; species Vesiculovirus chandipura) also have two small, highly basic proteins encoded in a second ORF within the P gene (Walker et al., 2015). In VSIV, an alternative ORF is expressed from alternative initiation codons as two proteins of 65 and 55 amino acids (C and C′, respectively) (Spiropoulou and Nichol 1993, Peluso et al., 1996). The roles of the nonstructural C and C′ proteins are unclear. Engineered viruses that do not express C proteins are indistinguishable from wild-type virus in protein synthesis, virus production and host-protein synthesis shut-off in tissue culture cells (Kretzschmar et al., 1996).
Genome organisation and replication
Total genomic RNA lengths range from approximately 10,700 bases in Carajas virus (CARV; species Vesiculovirus carajas) to 11,336 bases in field isolates of VSIV (Rodriguez et al., 2002, Walker et al., 2015) (Figure 2 Vesiculovirus). The VSIV genome organisation consists of a 47 nt leader sequence followed by the five structural protein genes in the order 3′-N-P-M-G-L-5′ and a 57–59 nt trailer sequence at the 5′-end. Most differences in length are found in non-translated regions, particularly in the M and G mRNAs (Rodriguez et al., 2002). Despite variability in lengths of the five mRNAs of VSIV, COCV and vesicular stomatitis Alagoas virus (VSAV; species Vesiculovirus alagoas) (all assigned to the Indiana serotype), four of the five predicted structural proteins (N, M, G and L) vary in length by only one amino acid residue.
Vesiculovirus replication is well studied and has served as model for replication of the rhabdoviruses (Banerjee 1987, Banerjee and Barik 1992). Transcription begins at a single entry site at the 3′-end of the genome with each gene expressed as a capped and polyadenylated monocistronic mRNA. The first transcript is the 3′-leader, which is neither capped nor polyadenylated. It is transported to the nucleus where it inhibits host cell transcription. The leader transcript is followed by the N mRNA, which is capped during synthesis by the virion polymerase complex (composed of N, P and L). The intergenic sequence (5′-AGUUUUUUUCAUA-3′) signals polyadenylation, termination and re-initiation of transcription in decreasing amounts as the polymerase complex moves away from the single entry site. Therefore, the gene order (3′-N>P>M>G>L-5′) provides an efficient way of regulating gene expression, by which proteins necessary in larger amounts (such as N) are located near the 3′-end and are transcribed in larger amounts and those needed in smaller amounts are located towards the 5′-end and are transcribed less frequently. Following translation of the mRNAs to yield the viral proteins, genome replication starts. In the replication process, the L protein initiates at the 3′-end of the genome, ignores all the signals for stop and polyadenylation of individual mRNAs and instead synthesises a full-length complementary anti-genome. This, in turn, serves as the template for the synthesis of a 45-nt minus sense leader RNA and for the synthesis of full-length progeny genomes. Full-length genomes can either serve as templates for secondary transcription or can be assembled into infectious particles. Factors determining the RdRP functions in transcription or replication modes are not fully understood but the proportion of N and available RNP templates are thought to be critical for determining these functions (Arnheiter et al., 1985). Following replication, further rounds of transcription (secondary transcription), translation and replication ensue.
 |
| Figure 2 Vesiculovirus. Schematic representation of vesiculovirus genomes shown in reverse (positive-sense) polarity. N, P, M, G and L represent ORFs encoding the structural proteins. Alternative ORFs (light blue and dark blue) in the P gene of some vesiculoviruses are highlighted. In vesicular stomatitis Indiana virus, the alternative ORF in the P gene encodes two small basic proteins (C and C′) that are expressed in infected cells. In Mediterranean bat vesiculovirus, there is an alternative ORV at the start of the G gene (grey). |
Biology
Vesiculoviruses infect mammals and are transmitted by insects and, therefore, are considered arboviruses (Rodriguez and Pauszek 2012). The natural cycle of vesiculoviruses infecting mammals remains largely unknown but these viruses are commonly found in insects of several species and serological evidence suggests they are capable of infecting wild mammals, birds and reptiles in endemic areas. Several vesiculoviruses (American bat vesiculovirus [ABVV; species Vesiculovirus eptesicus], Jinghong bat virus [JhBV; species Vesiculovirus rhinolophus], Mediterranean bat virus [MBV; species Vesiculovirus mediterranean], Yinshui bat virus [YSBV; species Vesiculovirus yinshui] and Wufeng Myotis altarium vesiculovirus 1 [WMalVV1; species Vesiculovirus wufeng]) have been detected by metagenomic sequencing in insectivorous bats (Rhinolophidae and Vespertilionidae). Mejal virus (MEJV; species Vesiculovirus mejal) was detected by metagenomic sequencing of streblid bat flies (Trichobius sp.) collected from Parnell's mustached bats (Pteronotus parnellii) sampled in Mexico (Ramírez-Martínez et al., 2021). Many vesiculoviruses have been isolated from phlebotomine sandflies in which vertical (transovarial) transmission has been demonstrated experimentally (Tesh et al., 1971, Tesh et al., 1972, Tesh and Modi 1983, Travassos da Rosa et al., 1984, Tesh et al., 1987, Comer et al., 1990). Vesiculoviruses can be transmitted both by insects and by contact. VSIV, VSNJV and VSAV are transmitted to cattle, horses and pigs by various blood-sucking insects found to be infected during epidemics, including phlebotomine sandflies, blackflies and biting midges, but also can be transmitted between mammals by direct contact (Rodriguez and Pauszek 2012). Experimental mechanical transmission also has been achieved by feeding vesiculovirus laboratory-infected grasshoppers to cattle. However, grasshoppers have never been shown to carry vesiculoviruses in nature. Vesiculoviruses are transmitted not only transovarially but also horizontally between infected and non-infected black-flies while co-feeding on mammalian hosts (Tesh et al., 2016). The latter means of transmission might explain the noticeable absence of viremic mammalian hosts for vesiculoviruses, an unusual feature for an arbovirus.
VSNJV, VSIV, COCV and VSAV are primarily veterinary pathogens, causing epizootics in cattle, pigs and horses in the Americas (Rodriguez and Pauszek 2012). Infected animals usually develop 3–5 days of fever with vesicular lesions on the tongue, buccal mucosa, teats, and coronary bands of the feet. The illness in bovine and swine clinically resembles foot-and-mouth disease, which presents an urgent diagnostic problem. Complications of infection in these animals include weight loss, dehydration, reduced milk production and mastitis in cows, and lameness. Clinical infection also occurs in humans and is usually characterised by a 3–4 day flu-like illness. Most reported cases have occurred in veterinarians and animal handlers in contact with sick animals. CHPV, Isfahan virus (ISFV; species Vesiculovirus isfahan) and Piry virus (PIRYV; species Vesiculovirus piry) have also been reported to cause a flu-like illness in humans (Tesh et al., 1977, Rodriguez and Pauszek 2012). Reports that CHPV also causes fatal encephalitis in humans have been questioned (Manghani et al., 1986, Sejvar 2006).
Antigenicity
Vesiculoviruses have been classified into serotypes based on their neutralisation pattern which is determined by epitopes located in the viral G (Kelley et al., 1972, Tesh et al., 1977, Lefrancois and Lyles 1982). G also determines protection in animals vaccinated either with inactivated whole virus or with subunit vaccines (Mackett et al., 1985, Yilma et al., 1985, Gearhart et al., 1987). Furthermore, recombinant viruses containing G of VSNJV, but all other proteins from VSIV, induced protective immune responses against challenge with VSNJV (Martinez et al., 2004). N is a cross-reactive antigen used in complement fixation tests that help define members of the genus (Kang and Prevec 1970). Weak serological cross-reactions may occur between viruses of different genera (Tesh et al., 1983, Calisher et al., 1989). One of the criteria used for vesiculovirus classification is cross-reactivity by complement fixation (Tesh et al., 1977).
In the case of vesicular stomatitis viruses, there are two major serotypes: New Jersey and Indiana. The Indiana serotype has been subdivided into four distinct serological complexes (Federer et al., 1967, Pauszek et al., 2008). Serotype Indiana 1 is classical VSIV found from northern South America to southern USA. The Indiana 2 serotype has COCV as the prototype virus which was originally isolated from mites collected from rice rats in Trinidad in 1961. Indiana 2 viruses cause disease in cattle and horses in Brazil and Argentina. The Indiana 3 subtype is represented by VSAV which was first isolated from a mule in Alagoas, Brazil, in 1964. This subtype is the most common cause of vesicular stomatitis in livestock in Brazil. The Indiana 4 subtype was originally identified as VSAV but was subsequently shown to be genetically distinct and named Morreton virus (MORV; species Vesiculovirus morreton) after the location in northeast Colombia from which it was first isolated from phlebotomine sandflies in 1986 (Tesh et al., 1987). MORV antibodies have been detected in animals and humans in Colombia but a direct association with disease has not yet been demonstrated (Tesh et al., 1987). Other vesiculoviruses infect mammals, including humans, (e.g., PIRYV, ISFV and CHPV, and some have been isolated from blood-sucking insects (e.g., CARV, Maraba virus (MARV; species Vesiculovirus maraba) and Radi virus (RADV; species Vesiculovirus radi) (de Souza et al., 2016).
Species demarcation criteria
Vesiculovirus species have been defined primarily by serological means coupled with phylogenetic analyses of the genomes. Biological characteristics such as host range and mechanisms of transmission are also used to distinguish viral species within the genus.
Viruses assigned to different species within the genus have several of the following characteristics: A) minimum amino acid sequence divergence of 20% in the L protein; B) minimum amino acid sequence divergence of 10% in the N protein; C) minimum amino acid sequence divergence of 15% in the G protein; D) can be distinguished in serological tests; and E) occupy different ecological niches as evidenced by differences in hosts and or arthropod vectors.
Related, unclassified viruses
| Virus name | Accession number | Virus abbreviation |
| Benxi bat virus | KX343071* | BxBV |
| Guampa virus | MN225577* | GMPV |
| Kannamangalam virus | JX276994* | KANV |
| Wuhan louse fly virus 11 | KM817658* | WhLFV-11 |
Virus names and virus abbreviations are not official ICTV designations.
* Coding region sequence incomplete