Subfamily: Betarhabdovirinae
Genus: Alphacytorhabdovirus
Distinguishing features
Historically, the genera Nucleorhabdovirus and Cytorhabdovirus were established based on the sites of virus replication and morphogenesis, with cytorhabdoviruses replicating and maturing in the cytoplasm of infected cells. Recently, a reclassification became necessary as phylogenetic analyses of new plant rhabdovirus genomes consistently showed that cytorhabdoviruses did not form a monophyletic clade upon analysis of complete L protein sequence alignments (Rubino et al., 2025). Based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L protein sequences, viruses classified in the genus Alphacytorhabdovirus form a monophyletic cluster clearly distinguished from beta- and gammacytorhabdoviruses, as well as from other plant rhabdoviruses.
Virion
Morphology
Enveloped virions are bacilliform, 60–75 nm in diameter and 200–350 nm long (Jackson et al., 2005a, Dietzgen, 2002).
Physicochemical and physical properties
The buoyant density in sucrose or potassium tartrate is 1.19–1.20 g cm−3 (Dietzgen, 2002). The lipid envelope is derived from the cytoplasmic membranes of plant or insect host cells (Jackson et al., 2005a).
Nucleic acid
The negative-sense, single-stranded RNA genome of 12.1–14.7 kb is unsegmented. Six to eight mRNAs, one for each of the encoded proteins, have been identified in infected plants.
Proteins
N, P, M, G and L represent the five canonical rhabdovirus structural proteins. P of lettuce necrotic yellows virus (LNYV; species Alphacytorhabdovirus lactucanecante) and alfalfa dwarf virus (ADV; species Alphacytorhabdovirus medicagonis) have RNA silencing suppressor activity (Mann et al., 2015, Bejerman et al., 2016, Mann et al., 2016b). The P3 proteins of LNYV and ADV (encoded in the 4b gene) localize to plasmodesmata and trans-complement movement of heterologous viruses between plant cells (Mann et al., 2016a). ADV, strawberry crinkle virus (SCV; species Alphacytorhabdovirus fragariarugosus) encode small proteins that have the structural characteristics of class 1a viroporins, similar to those detected commonly in mammalian rhabdoviruses.
Lipids
The lipoprotein envelope is derived from the host plant or the insect vector (Jackson et al., 2005a). Lipid composition is unknown.
Carbohydrates
LNYV G is glycosylated with a complex network of oligosaccharides containing N-acetylchitobiose, N-linked to asparagine residues. The carbohydrate binds to concanavalin A and can be removed by endoglycosidase F (Dietzgen & Francki, 1988).
Genome organisation and replication
The LNYV genome organisation (12.8 kb) is similar to that of Sonchus yellow net virus (SYNV, see genus Betanucleorhabdovirus). Preceded by a non-coding 84 nt leader sequence, the gene order is 3′-N-P-4b-M-G-L-5′ (Figure 1 Alphacytorhabdovirus). The N gene encodes the nucleoprotein (or nucleocapsid protein), and the P, M, G and L genes encode the phosphoprotein (polymerase cofactor), matrix protein, glycoprotein and RNA polymerase, respectively. The 4b gene encodes a cell-to-cell movement protein (P3). The intergenic regions contain highly conserved consensus sequences. The 5′-non-coding trailer sequence of 187 nt has extensive complementarity to the 3′-leader. Many alphacytorhabdoviruses harbor an overlapping ORF within the P gene; the function of the product of this ORF is unknown, and is not known if it is expressed. Moreover, many alphacytorhabdoviruses have an additional gene between the G and L genes that encodes a class 1a viroporin-like protein. One member also has an additional gene between the P3 and M genes, while other member have an additional gene after the L gene.
Alphacytorhabdoviruses replicate in the cytoplasm of infected cells in association with masses of thread-like structures (viroplasms) (Jackson et al., 2005a). Virions bud in association with the endoplasmic reticulum (ER) and accumulate in ER-derived vesicles (Chambers et al., 1965, Wolanski & Chambers 1971). A nuclear phase has been suggested but not proven in the replication of some alphacytorhabdoviruses (e.g., LNYV) (Wolanski & Chambers 1971). Evidence of the nuclear involvement in the replication of others is lacking. Endogenous transcriptase activity is readily detectable in alphacytorhabdovirus preparations (Francki & Randles, 1972).
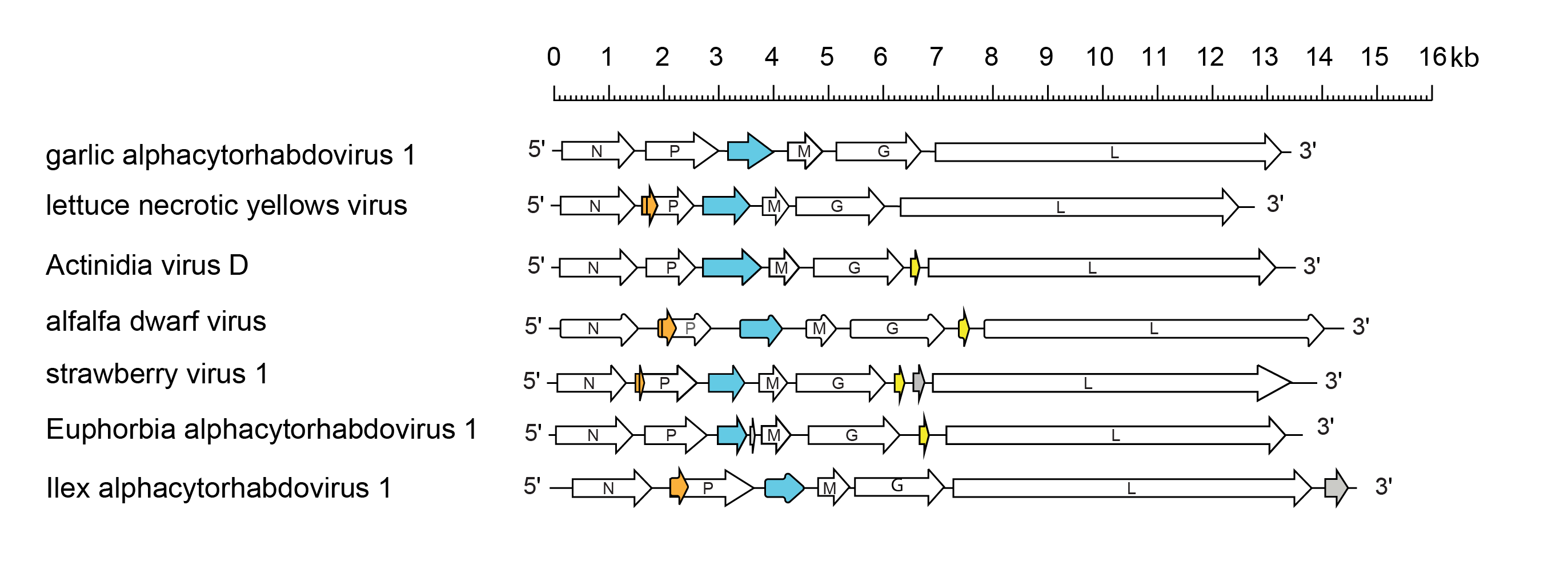 |
| Figure 1 Alphacytorhabdovirus. Schematic representation of a selection of alphacytorhabdovirus genomes shown in reverse (positive-sense) polarity. N, P, M, G and L represent ORFs encoding the structural proteins. ORFs encoding viral cell-to-cell movement proteins (blue) and predicted class 1a viroporin-like proteins (yellow) are shown. Other ORFs encode putative accessory proteins of unknown function, some of which occur as homologous sets (orange). ORFs with no obvious homologues are also shown (grey). In addition to those virus genomes shown: Artemisia alphacyotrhadovirus 1 is similar to garlic alphacytorhabdovirus 1; Arctium alphacytorhabdovirus 1, Baccharis alphacytorhabdovirus 1, blackcurrant rhabdovirus 2, cabbage cytorhabdovirus 1, Conopholis alphacytorhabdovirus 1, chrysanthemum alphacytorhabdovirus 1, Cnidium virus 2, Cynara alphacytorhabdovirus 1, Daphne virus 1, Geum alphacytorhabdovirus 1, Hedera alphacytorhabdovirus 1, lettuce yellow mottle virus, Morinda alphacytorhabdovirus 1, oak alphacytorhabdovirus 1, Phyllostachys alphacytorhabdovirus 1, Pogostemom alphacytorhabdovirus 1, Primula alphacytorhabdovirus 1, Primula alphacytorhabdovirus 2, Sambucus virus 1, strawberry virus 2, Tolmiea alphacytorhabdovirus 1, Trichosanthes associated virus 1, Trifolium pratense virus B and Wuhan insect virus 5 are similar to lettuce necrotic yellows virus; Utricularia alphacytorhabdovirus 1 and Wuhan insect virus 6 are similar to Actinidia virus D; Artemisia alphacytorhabdovirus 2, Artemisia alphacytorhabdovirus 3, Asclepias syriaca virus 1, Bacopa monieri virus 1, Cardamine alphacytorhabdovirus 1, Chelidonium yellow mottle associated virus, chrysanthemum yellow dwarf virus, coriander alphacytorhabdovirus 1, Ficus alphacytorhabdovirus 1, Glehnia littoralis virus 1, honeysuckle-associated cytorhabdovirus 1, Hyptis latent virus, Kenyan potato cytorhabdovirus, Medicago alphacytorhabdovirus 1, Mentha alphacytorhabdovirus 1, Nymphaea alba virus 1, Ocimum alphacytorhabdovirus 1, patchouli chlorosis associated cytorhabdovirus, Pastinaca cytorhabdovirus 1, Pelargonium alphacytorhabdovirus 1, persimmon virus A, Pinellia alphacytorhabdovirus 1, Plumbago necrotic spot associated virus, Pogostemom alphacytorhabdovirus 2, Pogostemom alphacytorhabdovirus 3, raspberry vein chlorosis virus, rose alphacytorhabdovirus 1, Rubus alphacytorhabdovirus 1, Scutellaria alphacytorhabdovirus 1, strawberry crinkle virus, Taraxacum cytorhabdovirus 1, tomato yellow mottle associated virus, Trifolium pratense virus A, Triticum alphacytorhabdovirus 1, wetland metagenome associated alphacytorhabdovirus 1, Wuhan insect virus 4, Wurfbainia alphacytorhabdovirus 1 and Zea alphacytorhabdovirus 1 are similar to alfalfa dwarf virus; |
Biology
A wide variety of monocot and dicot plants are susceptible to alphacytorhabdoviruses, although each virus usually has a restricted host range. Alphacytorhabdoviruses are transmitted by aphids. Some viruses are also transmitted during vegetative propagation, and some can also be transmitted mechanically from infected sap. Seed transmission has not been reported. In all carefully examined cases, viruses replicate in cells of the insect vector as well as in the plant host (Jackson et al., 2005a, Redinbaugh & Hogenhout, 2005).
Species demarcation criteria
Viruses assigned to different species within the genus Alphacytorhabdovirus have several of the following characteristics: A) nucleotide sequence identity less than 75% for the coding-complete genome sequence; B) amino acid sequence identity less than 86% in proteins encoded by all the cognate open reading frames; and C) they occupy different ecological niches as evidenced by differences in hosts and/or arthropod vectors.
Related, unclassified viruses
| Virus name | Accession number | Virus abbreviation |
| Achillea alphacytorhabdovirus 1 | PX121434 | AchACRV1 |
| Artemisia alphacytorhabdovirus 4 | PX121437 | ArtACRV4 |
| buckwheat alphacytorhabdovirus 1 | PX121447 | BucACRV1 |
| Capsicum alphacytorhabdovirus 1 | PX121449 | CapACRV1 |
| creeping Charlie alphacytorhabdovirus 1 | PX121454 | CrChACRV1 |
| Ficus alphacytorhabdovirus 2 | PX121455 | FicACRV2 |
| hogweed alphacytorhabdovirus 1 | PX121457 | HogACRV1 |
| Mentha alphacytorhabdovirus 2 | PX121461 | MenACRV2 |
| parsley alphacytorhabdovirus 1 | PX051447 | ParACRV1 |
| parsley latent virus | PX051446 | ParLV |
| Pelargonium alphacytorhabdovirus 2 | PX121464 | PelACRV2 |
| Sedum alphacytorhabdovirus 1 | PX121466 | SedACRV1 |
| Spiraea cytorhabdovirus 1 | PV231214 | SpiCRV1 |
Virus names and virus abbreviations are not official ICTV designations.

