Subfamily: Alpharhabdovirinae
Genus: Zarhavirus
Distinguishing features
Viruses assigned to the genus Zarhavirus form a distinct monophyletic group based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L sequences. Zahedan rhabdovirus (ZARV; species Zarhavirus zahedan) was isolated from hard ticks.
Virion
Morphology
Virion morphology is unknown.
Nucleic acid
The ZARV genome consist of a single molecule of negative-sense, single-stranded RNA of approximately 13.2 kb (Dilcher et al., 2015).
Proteins
The ZARV N, P, M, G and L proteins share sequence homology and/or structural characteristics with the cognate proteins of other rhabdoviruses.
Genome organisation and replication
The ZARV genome includes only the five genes (N, P, M, G and L) encoding the structural protein (Figure 1 Zarhavirus).
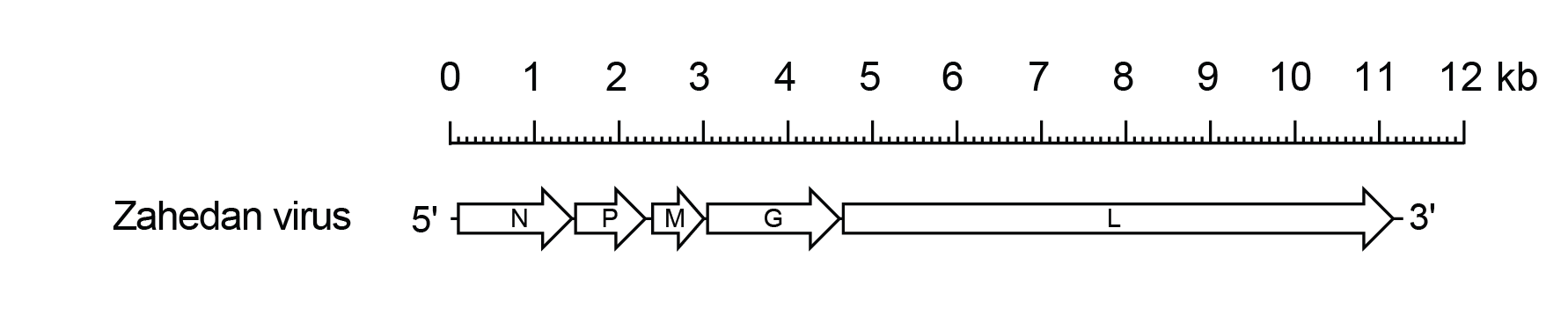 |
| Figure 1 Zarhavirus. Schematic representation of the Zahedan rhabdovirus genome shown in reverse (positive-sense) polarity. The genome contains only five long open reading frames (ORFs) in the N, P, M, G and L genes (open arrows). |
Biology
ZARV was isolated from hard ticks (Hyalomma anatolicum anatolicum) collected in 2001, in Zahedan, Iran (Dilcher et al., 2015). The virus has been shown to replicate in Vero cell cultures and newborn mice (Dilcher et al., 2015).
Species demarcation criteria
There is only one species in the genus. For viruses to be assigned additional species in the genus, several of the following characteristics would have to be observed: A) minimum amino acid sequence divergence of 10% in the N proteins; B) minimum amino acid sequence divergence of 10% in the L proteins; C) minimum amino acid sequence divergence of 15% in the G proteins; D) significant differences in genome organisation as evidenced by numbers and locations of ORFs; E) they can be distinguished in virus neutralisation tests; and F) they occupy different ecological niches as evidenced by differences in vertebrate hosts and or arthropod vectors.

