Subfamily: Alpharhabdovirinae
Genus: Betathriprhavirus
Distinguishing features
Viruses assigned to the genus Betathriprhavirus form a distinct monophyletic group based on well-supported Maximum Likelihood or Maximum Clade Credibility (MCC) trees inferred from complete L sequences. They have been detected in thrips (insects in the family Thripidae) and are most closely related to almendraviruses, which have been detected in mosquitoes. They are distant phylogenetically from viruses assigned to the genus Alphathriprhavirus.
Virion
Morphology
Viruses assigned to the genus have not yet been isolated or visualized by electron microscopy.
Nucleic acid
The genome consists of a single molecule of negative-sense, single-stranded RNA of approximately 10.8 kb.
Proteins
The N, P, M, G and L share sequence homology and/or structural characteristics with the cognate proteins of other rhabdoviruses. The small proteins encoded in the U1 ORFs range from 72 to 95 amino acids (8.3–10.7 kDa) and share clearly recognisable sequence homology.
Genome organisation and replication
Betathriprhavirus genomes include five genes (N, P, M, G and L) encoding the structural proteins and an additional gene between the G and L genes that encodes a protein (U1) of unknown function (Figure 1 Betathriprhavirus).
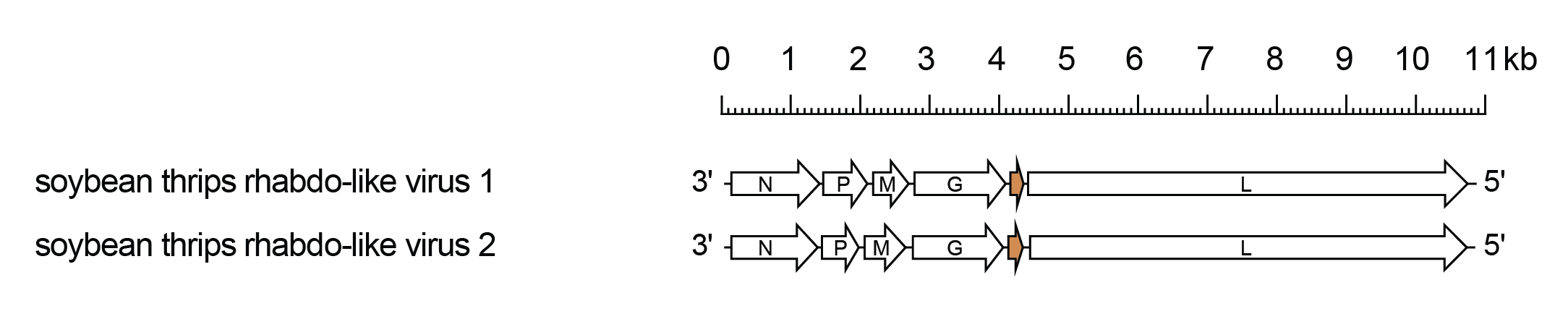 |
| Figure 1 Betathriprhavirus. Schematic representation of betathriprhavirus genomes shown in reverse (positive-sense) polarity. N, P, M, G and L represent ORFs encoding the structural proteins. The U1 ORF (orange) following the G gene encodes a protein of unknown function. |
Biology
Betathriprhaviruses have been detected only by high-throughput sequencing of thrips. Soybean trips rhabdo-like virus 1 (species Betathriprhavirus variabilis) and soybean trips rhabdo-like virus 2 (species Betathriprhavirus midwest) were each detected in a pool of soybean thrips (Neohydatothrips variabilis) samples collected at various locations, primarily in the midwestern states (Illinois, Iowa, Kansas, Michigan, Minnesota, Missouri and Wisconsin) of the USA, in 2018 (Thekke-Veetil et al., 2020).
Species demarcation criteria
Viruses assigned to different species within the genus Betathriprhavirus have several of the following characteristics: A) minimum amino acid sequence divergence of 12% in the G protein; B) minimum amino acid sequence divergence of 8% in the L protein; C) minimum amino acid sequence divergence of 4% in the N protein; D) can be distinguished in virus neutralization tests; E) exhibit significant differences in genome organization as evidenced by numbers and locations of ORFs; and F) occupy different ecological niches as evidenced by differences in hosts and or arthropod vectors.

