Family: Rhabdoviridae
Genus: Betahymrhavirus
Distinguishing features
Viruses assigned to the genus Betahymrhavirus form a distinct monophyletic group based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L sequences. Members of the genus have been detected in hymenopteran insects (Hymenoptera) including wasps. They are distant phylogenetically from viruses assigned to the genus Alphahymrhavirus.
Virion
Morphology
Viruses assigned to the genus have not yet been visualized by electron microscopy.
Nucleic acid
Betahymrhavirus genomes consist of a single molecule of negative-sense, single-stranded RNA of approximately 12.6–13.2 kb (Käfer et al., 2019).
Proteins
Betahymrhavirus N, P, M, G and L proteins share sequence homology and/or structural characteristics with the cognate proteins of other rhabdoviruses. Betahymrhavirus G proteins are class I transmembrane glycoproteins sharing 14 conserved cysteine residues which are likely to form seven disulphide bridges in the folded protein. However, alignment with the G protein of vesicular stomatitis Indiana virus (species Vesiculovirus indiana) indicates that few if any of these residues correspond to the conserved set of 12 cysteines that variously occur in other rhabdovirus G proteins (Walker and Kongsuwan 1999, Roche et al., 2006).
Genome organisation and replication
Betahymrhavirus genomes include the five genes (N, P, M, G and L) encoding the structural proteins (Figure 1.Betahymrhavirus). They also each have an additional gene (U1) between the M gene and G gene in with two overlapping reading frames with the major ORF (U1) commencing 160-170 nt downstream of the initiation codon of the minor ORF (U1x).
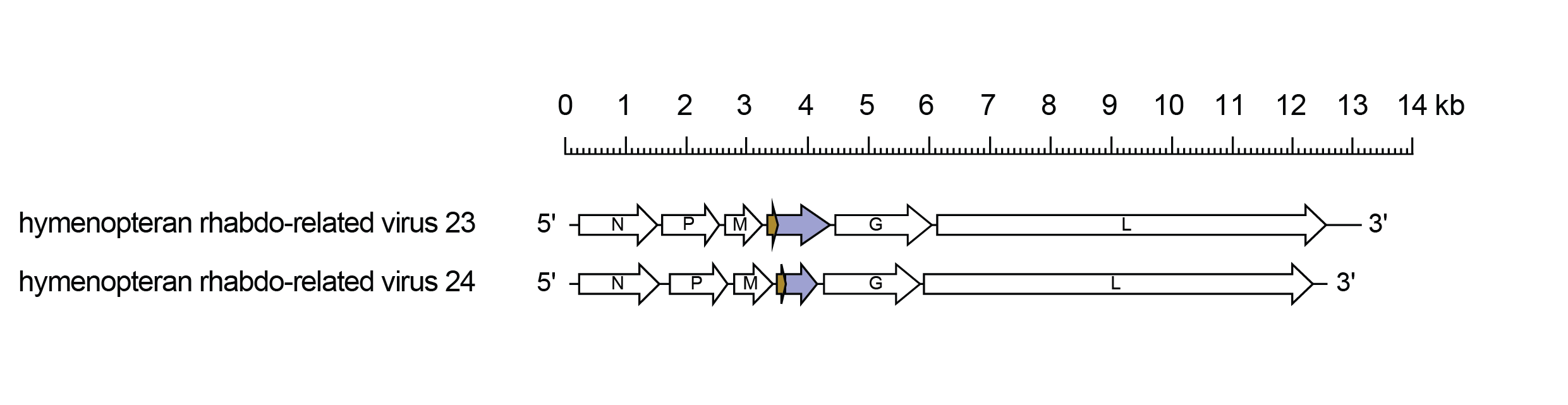 |
| Figure 1.Betahymrhavirus. Schematic representation of betahymrhavirus genomes shown in reverse (positive-sense) polarity. The five long open reading frames (ORFs) in the N, P, M, G and L genes are shown (open arrows). Overlapping ORFs U1 (light purple) and U1x (brown) that are located in an additional gene (U1) are also shown. |
Biology
Hymenopteran rhabdo-related virus 23 (species Betahymrhavirus austriaca) was discovered in the transcriptome shotgun assembly (TSA) of a cuckoo wasp (Chrysura austriaca) collected in Germany, in 2010 (Käfer et al., 2019). It was also detected in the TSA of a cuckoo wasp (Chrysis sp.) collected in Israel, in 2012 and named hymenopteran rhabdo-related virus 22 (Käfer et al., 2019). Hymenopteran rhabdo-related virus 24 (species Betahymrhavirus heterodontonyx) was discovered in the TSA of a leaden spider wasp (Heterodontonyx sp.) collected in Western Australia, in 2011 (Käfer et al., 2019). No isolates are currently available for either of these viruses.
Species demarcation criteria
Viruses assigned to different species within the genus have several of the following characteristics: A) minimum amino acid sequence divergence of 10% in N proteins; B) minimum amino acid sequence divergence of 10% in the L proteins; C) minimum amino acid sequence divergence of 15% in G proteins; D) significant differences in genome organisation as evidenced by numbers and locations of ORFs; E) can be distinguished in virus neutralisation tests; and F) occupy different ecological niches as evidenced by differences in invertebrate hosts.
Related, unclassified viruses
| Virus name | Accession number | Virus abbreviation |
| Xiangshan rhabdo-like virus 4 | OK491502 | XsRLV4 |
Virus names and virus abbreviations are not official ICTV designations.

