Family: Spinareoviridae
Jelle Matthijnssens, Houssam Attoui, Krisztián Bányai, Corina P. D. Brussaard, Pranav Danthi, Mariana del Vas, Terence S. Dermody, Roy Duncan, Qín Fāng (方勤), Reimar Johne, Peter P. C. Mertens, Fauziah Mohd Jaafar, John Patton, Takahide Sasaya (笹谷孝英), Nobuhiro Suzuki (鈴木信弘) and Taiyun Wei (魏太云)
The citation for this ICTV Report chapter is the summary published as Matthijnssens et al., (2022) ICTV Virus Taxonomy Profile: Spinareoviridae, Journal of General Virology (2022) 103:001781.
Corresponding author: Jelle Matthijnssens ([email protected])
Edited by: Jens H. Kuhn and Stuart G. Siddell
Posted: October 2021, June 2022, May 2025
Summary
The family Spinareoviridae includes viruses that have relatively large spikes or turrets situated at the 12 icosahedral vertices of either the virion or core particle (Table1 Spinareoviridae), whereas these are not found for viruses in the family Sedoreoviridae. The transcriptionally active core particle of spinareovirids appears to contain only a single complete capsid layer to which the projecting spikes or turrets are attached. In most cases, the core is surrounded (in the complete virion) by an incomplete protein layer (with T=13 symmetry) that forms the outer capsid, which is penetrated by the projections from the core surface (Dryden et al., 1993). Exceptions are the cypoviruses, which have transcriptionally active but fully intact virions composed of only a single capsid shell that is equivalent to the core particles of viruses from other genera. However, virions of most cypoviruses are characteristically occluded (either singly or multiply) within the matrix of proteinaceous crystals called polyhedra. These are composed primarily (>90%) of the viral polyhedrin protein. In most cases, phylogenetic analysis of RdRP sequences identifies spinareovirids and sedoreovirids as members of separate clades.
Table 1 Spinareoviridae. Characteristics of members of the family Spinareoviridae
| Characteristic | Description |
| Example | mammalian orthoreovirus-3 Dearing (T3D; L1: HM159613; L2: HM159614; L3: HM159615; M1: HM159616; M2: HM159617; M3: HM159618; S1: HM159619; S2: HM159620; S3: HM159621; S4: HM159622), species Orthoreovirus mammalis |
| Virion | Non-enveloped, icosahedral, 60–85 nm virions composed of 1–3 concentric capsid proteins layers |
| Genome | 23–29 kbp of segmented linear dsRNA, with each of the 9–12 segments ranging from 0.5–4.8 kbp. |
| Replication | Replication occurs in the cytoplasm in electron-dense structures called viroplasms, viral inclusions, or viral factories |
| Translation | From full-length transcribed mRNAs that possess a 5′-terminal cap but no poly(A)-tail |
| Host Range | Mammals, aquatic animals (fish, mammals, crustaceans, molluscs), birds, reptiles, arthropods, fungi, and plants |
| Taxonomy | Realm Riboviria, kingdom Orthornavirae, phylum Duplornaviricota, class Resentoviricetes, order Reovirales: 9 genera and 58 species |
Virion
Spinareovirid particles are icosahedral. The protein capsid is organized as 1–3 concentric layers of capsid proteins, with an overall diameter of 50–85 nm (Mohd Jaafar et al., 2005). Spinareoviridae members have spikes or turrets at the 12 icosahedral vertices of the subviral particle (Figure 1 Spinareoviridae) in contrast to members of the Sedoreoviridae, which have an almost spherical or “smooth” appearance. For more details see the Reovirales Report.
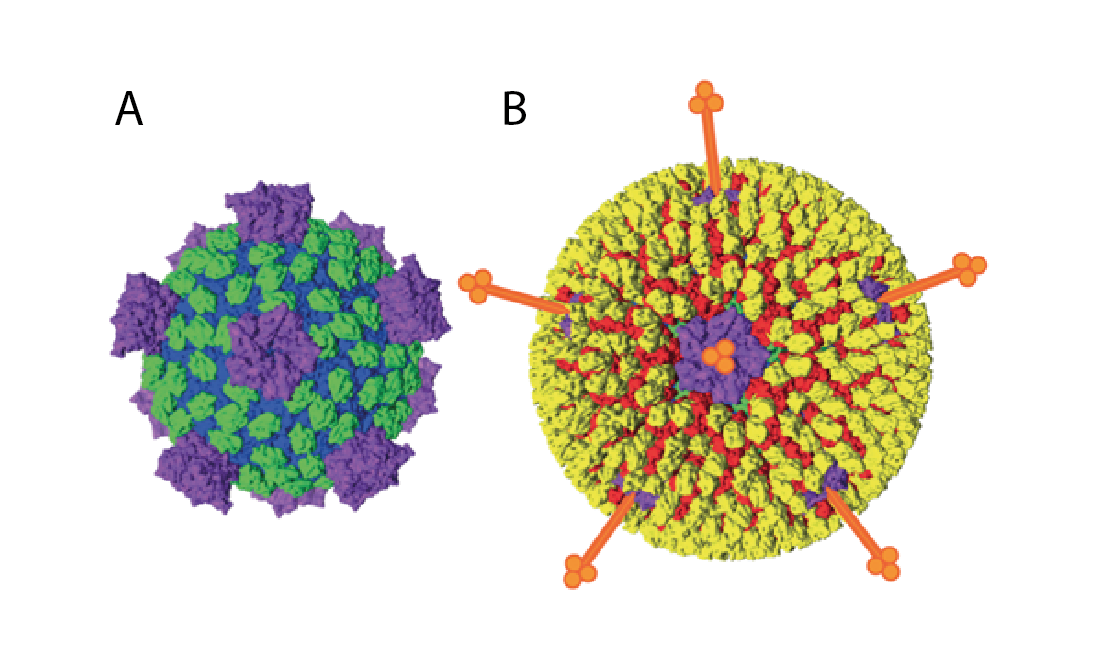 |
| Figure 1 Spinareoviridae Spinareovirid particle structure (e.g., mammalian orthoreovirus) particles are colored by functional similarity (blue – core shell, green – middle layer, yellow – outer capsid, purple – RNA capping, red – membrane penetration, orange – receptor binding) (shown schematically). A. Subviral particles. B. Virions. The diameter of the mature virion is approximately 80 nm (excluding spikes). Adapted from (Trask et al., 2012). |
Genome organisation and replication
Spinareovirids contain 9–12 segments of linear dsRNA comprising 23–29 kbp in total, with individual segments ranging from 0.5–4.8 kbp. The positive-sense strands of each duplex are modified with a 5′-terminal type 1 cap structure but no 3′-poly(A) tail. The viral RNAs are mostly monocistronic with relatively short 5′- and 3′-non-coding regions, although some segments have a second or third functional open-reading frame (Marzachi et al., 1995, Yang et al., 2020a).
Virus entry into cells varies between genera but usually results in loss of outer-capsid components. The resulting transcriptionally-active particles are released into the cytoplasm. The 5′-capped mRNAs are synthesized by structural enzymatic components of the viral particle and released into the cytoplasm through pores at the fivefold icosahedral vertices of the virion. Viral inclusions, also known as viral factories, are distributed throughout the cytoplasm. These neo-organelles are sites of viral mRNA synthesis, genome replication, and particle assembly (Boehme et al., 2011). Sets of a single copy of each capped mRNA are incorporated into progeny virus particles (Borodavka et al., 2018). These mRNAs serve as templates for negative-strand synthesis, thereby reconstituting genomic encapsidated dsRNAs. The steps involved in virion morphogenesis and virus egress from cells differ depending on the genus. Progeny virions are released from some cell types without compromising cell viability (e.g., budding) or from other types of cells following cell lysis (Roth et al., 2021).
Biology
Host are mammals, aquatic animals (fish, mammals, crustaceans, molluscs), birds, reptiles, arthropods, fungi, and plants (Mohd Jaafar et al., 2014, Suzuki et al., 2015). The biological properties of the viruses vary according to genus. See individual genus sections for details.
Derivation of names
Aquareovirus: from aqua, Latin for “water”
Coltivirus: from Colorado and tick
Cypovirus: from cytoplasmic polyhedrosis, in reference to the characteristic inclusions observed in infected cells.
Dinovernavirus: from double stranded insect novem (“nine” in Latin) segmented RNA viruses.
Fijivirus: from Fiji, the country where Fiji disease virus, a member of the genus, was first isolated.
Idnoreovirus: from insect-derived non-occluded (in contrast to the cypoviruses).
Mycoreovirus: from myco, Latin for “fungus”.
Orthoreovirus: from orthos, Greek for “straight”.
Oryzavirus: from the genus name of rice, the host of members of the type species.
Spinareoviridae: from spina, Latin for “spike”, denoting the presence of spikes or turrets on the surface of the core particles. The term 'spiked' is an alternative to 'turreted', that was used in early research to describe the structure of the particle, particularly with the cypoviruses (Hill et al., 1999).
Genus demarcation criteria
The number of genome segments (9, 10, 11, or 12) is in most cases characteristic of viruses within a single genus, although mycoreoviruses have either 11 or 12 genome segments (Kanematsu et al., 2004, Suzuki et al., 2004, Wei et al., 2004). Host (and vector) range and disease signs also are important indicators that help to identify viruses from different genera. Capsid structure (number of capsid layers, the presence of spiked or unspiked cores, and the symmetry and structure of the outer capsid) also can be significant. The level of sequence divergence, particularly in the more conserved genome segments and proteins (for example as detected by comparisons of RdRP or inner-capsid-shell proteins and the segments from which they are translated) can be used to distinguish members of different genera (Attoui et al., 2006). Available data suggest that isolates from different genera usually have <26% amino acid identity in comparisons between their RdRPs, while within a single genus, identities are usually >30% (Attoui et al., 2002a). However, Cryphonectria parasitica mycoreovirus 1 (CpMYRV1) has about 29% amino acid sequence identity with the RdRP of members of the genus Coltivirus (Hillman et al., 2004), and aquareoviruses and orthoreoviruses can have up to 42% aa identity.
Species demarcation criteria
The prime determinant for inclusion of virus isolates within a single virus species is their capacity to exchange genetic information during co-infection by genome segment reassortment, thereby generating viable progeny virus strains. However, data providing direct evidence of segment reassortment between isolates are available only for viruses of a few genera. The following methods are therefore commonly used (preferably in combination) to examine levels of similarity between isolates and to predict their possible compatibility as reassortants:
- Nucleotide and amino acid sequence analysis (viruses within different species should have low levels of sequence similarity between cognate genome segments).
- Serological comparisons of antigens or antibodies using either polyclonal antisera or monoclonal antibodies against conserved antigens. Methods used may include ELISA, complement fixation, and agar gel immunodiffusion. Closely related isolates and serotypes generally belong to the same species.
- Analysis of electropherotype by agarose gel electrophoresis (AGE) but not by polyacrylamide gel electrophoresis (PAGE). Virus isolates within the same species will show a relatively uniform electropherotype. However, a major deletion/insertion event may sometimes result in two distinct electropherotypes within a single species, and apparent similarities can exist between more closely related species.
- Identification of the conserved terminal regions of the genome segments. These are usually conserved across all segments within a species, although some closely related species also can have identical terminal sequences of at least some segments.
These criteria apply throughout the family. Additional or more specific criteria are provided in the section for each genus, where applicable.
Relationships within the family
Members of different genera segregate into different clades upon phylogenetic analysis of the RdRP protein (Figure 2 Spinareoviridae).
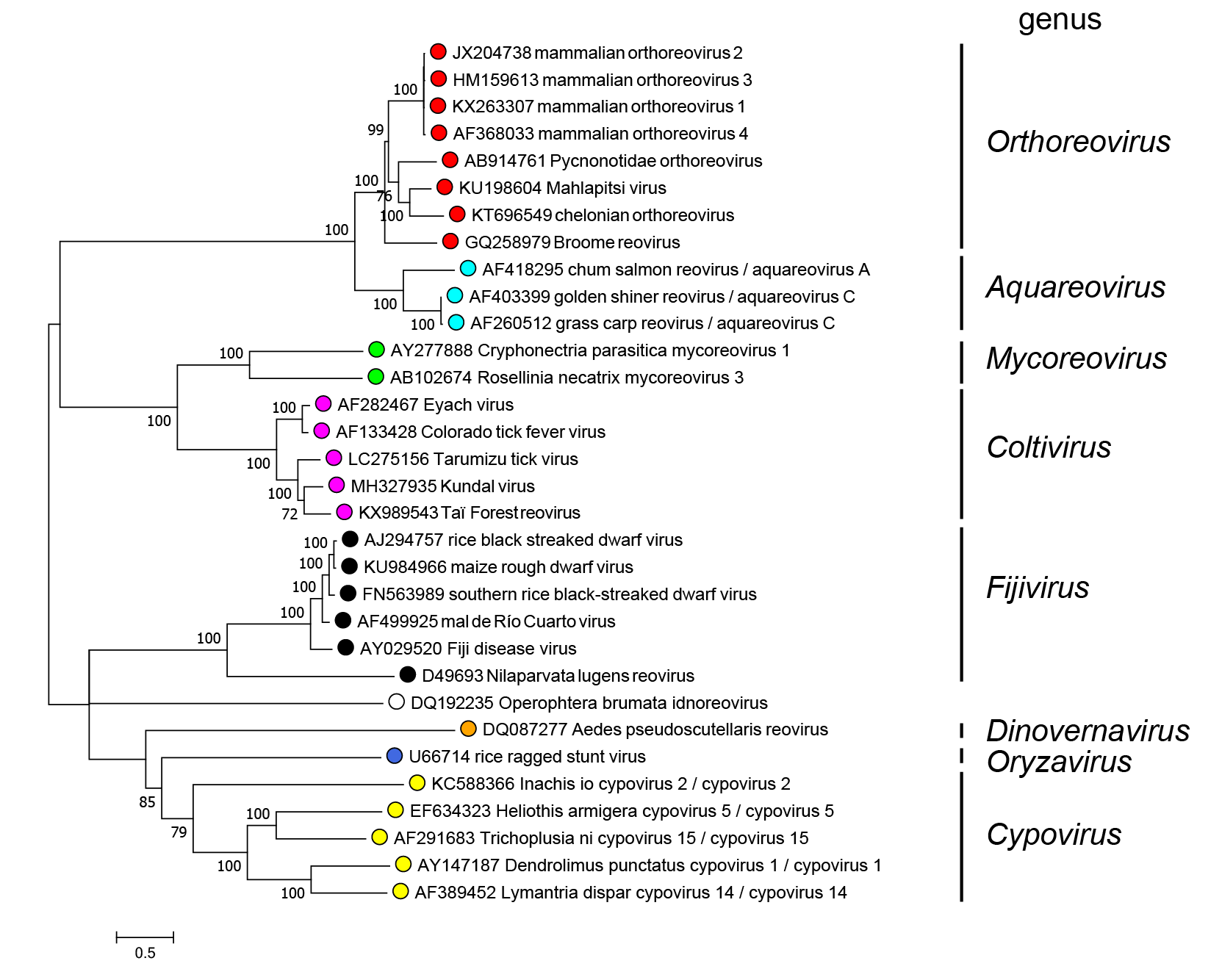 |
| Figure 2 Spinareoviridae. Phylogenetic tree of the RdRP protein of members of the family Spinareoviridae. Maximum likelihood tree constructed using the amino acid sequence of the putative RdRP of representative viruses in the family Spinareoviridae. The analysis used the Whelan And Goldman + Freq. model with a discrete gamma distribution of rates between sites, including some invariant sites amd was produced MEGA7 (Kumar et al., 2016). Values at the nodes represent bootstrap confidence levels (500 replications). This phylogenetic tree and corresponding sequence alignment are available to download from the Resources page. Diadromus pulchellus idnoreovirus 1 was not used as an isolate of the genus Idnoreovirus because its genome displays evidence of recombination. |
Relationships with other taxa
Spinareovirids are most closely related to the members of the family Sedoreoviridae, both families being placed in the order Reovirales. For a discussion of relationships with more distantly related viruses see the discussion in the Reovirales Report.
Related, unclassified viruses
| Virus name | Accession number | Abbreviation |
| Sclerotinia sclerotiorum reovirus 1 | Seg1: KU255424; Seg2: KU255425; Seg3: KU255426; Seg4: KU255427; Seg5: KU255428; Seg6: KU255429; Seg7: KU255430; Seg8: KU255431; Seg9: KU255432; Seg10: KU255433; Seg11: KU255434 | SsReV1 |
Virus names and virus abbreviations are not official ICTV designations.
Sclerotinia sclerotiorum reovirus 1 (SsReV1), was reported from Scletotinia sclerotiorum fungi and has an 11-segmented genome. SsReV1 is distantly related to mycoreoviruses and may represent a new genus in the family Spinareoviridae (Liu et al., 2017).

