Family: Phenuiviridae
Genus: Laulavirus
Distinguishing features
Five viruses, grapevine associated cogu-like virus 2 (GaCLV2), grapevine associated cogu-like virus 3 (GaCLV3), grapevine associated cogu-like virus 4 (GaCLV4), Laurel Lake virus (LLV), and Wardell virus (WRDV) are assigned to the genus Laulavirus. Laulaviruses genomes comprise three negative-sense RNA segments encoding a large protein (L), a nucleocapsid protein (N) and a non-structural protein that likely functions as a movement protein (MP) that enables cell-to-cell movement in plant hosts. Based on well-supported Maximum Likelihood or Maximum Clade Credibility trees inferred from complete L protein sequences, viruses classified in the genus Laulavirus form a monophyletic cluster clearly distinguished from other phenuivirids (Tokarz et al., 2018, Bertazzon et al., 2020, Chiapello et al., 2020).
Virion
Virion morphology is unknown. Based on the putative proteins encoded by the virus genome, the virion is probably a filamentous virion without an envelope.
Nucleic acid and Protein
Laulavirus genomes encompass three segments of negative-sense RNA, namely RNA1 (7.3–7.4 kb), RNA2 (2.3–2.5 kb) and RNA3 (1.0–1.7 kb). RNA3 of GaCLV4 is ambisenese. The terminal nucleotides of each segment occur in a canonical, conserved sequence (in coding sense) 5′-ACACAAAGUC…GUCUUUGUGU-3′ and may form the panhandle structures typical of other members of the class Bunyaviricetes (Table 2 Phenuiviridae). All three segments contain untranslated regions flanking a single ORF which, based on comparisons with other negative-sense RNA viruses, is predicted to be contained in the virus-complementary strand. RNA1 encodes a protein with a predicted molecular mass of 256–275 kDa that includes a region homologous with the bunyaviral RNA-directed RNA polymerase (RdRP) domain. RNA2 encodes a protein of 77–79 kDa that shares sequence homology and/or structural characteristics with the MP of plant viruses. RNA3 encodes a protein of 28–30 kDa that is homologous with the tenuivirus/phlebovirus N domain (Table 3 Phenuiviridae) (Tokarz et al., 2018, Bertazzon et al., 2020, Chiapello et al., 2020).
Genome organization and replication
The genome of laulaviruses consists of three negative-sense RNA segments (Figure 1 Laulavirus). These segments putatively encode L, MP (rather than external glycoproteins) and N, respectively. The laulavirus RNA3 segment does not encode a non-structural protein, but the GaCLV4 RNA3 exhibits an ambisense coding strategy and encodes a small non-structural protein of unknown function. Details of virus replication are unknown (Bertazzon et al., 2020, Chiapello et al., 2020, Rodriguez Coy et al., 2022).
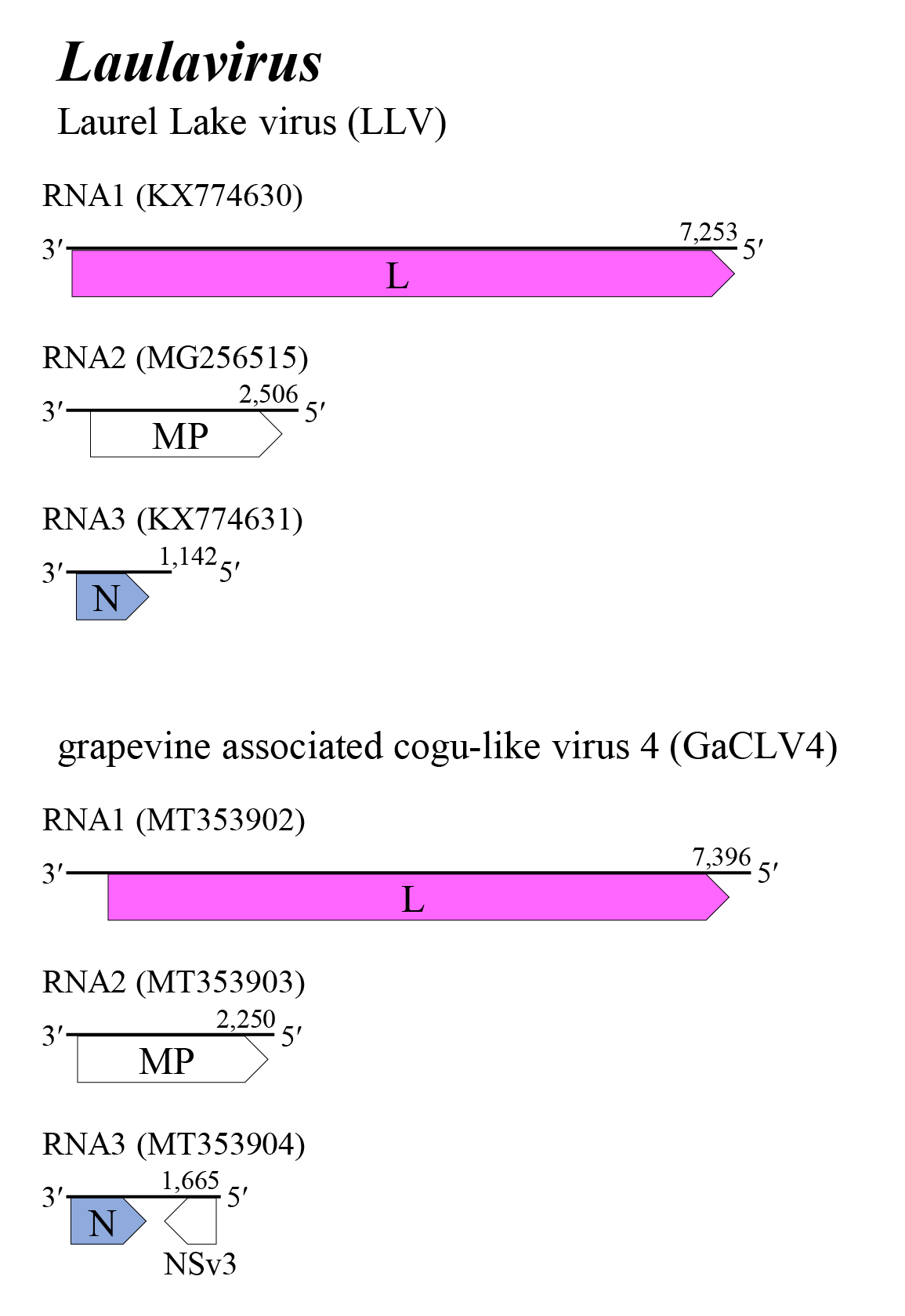 |
| Figure 1 Laulavirus. Genome organization of some laulaviruses. Coloured boxes depict ORFs that encode N, nucleocapsid protein and L, large protein. White boxes depict ORFs that encodes MP, non-structural cell-to-cell movement protein and NSv3, non-structural protein. |
Biology
Laulavirus RNA has been found by high-throughput sequencing of RNAs from downy mildew [Plasmopara viticola ((Berk. & M.A. Curtis) Berl. & De Toni, 1888)] or esca-associated fungi obtained from grapevines in Spain and Italy, and from the RNA of a pool of the ticks Ixodes scapularis (Say, 1821) and I. holocyclus (Neumann, 1899)] in Australia and USA. Wider host ranges are unknown but it is suggested that the viruses are either fungus-specific or tick-specific (Tokarz et al., 2018, Bertazzon et al., 2020, Chiapello et al., 2020).
Species demarcation criteria
The criteria demarcating species in the genus are:
• Less than 95% identity in the amino acid sequence of the RdRP.
Related, unclassified viruses
| Virus name | Accession number | Virus abbreviation |
| Colletotrichum associated negative-stranded RNA virus 1 | RNA1:OP471407 | CaNSRV1 |
| Hǎinán phenui-like virus 7 | RNA1: MW896888; RNA2: MW896889; RNA3: MW896890 | HanPLV7 |
Virus names and virus abbreviations are not official ICTV designations.

