Family: Geminiviridae
Genus: Topilevirus
Distinguishing features
This genus includes two species, Tomato apical leaf curl virus (Vaghi Medina et al., 2018) and Tomato geminivirus 1 (Fontenele et al., 2017). Members of these species have been identified in South America (Argentina and Brazil) and contain monopartite genomes with the conserved geminivirus nonanucleotide (5′-TAATATTAC-3′). Both viruses infect the same host plant, tomato, and tomato geminivirus 1 have been also isolated from Cleome sp. plants.
Virion
See discussion under family description.
Genome organization and replication
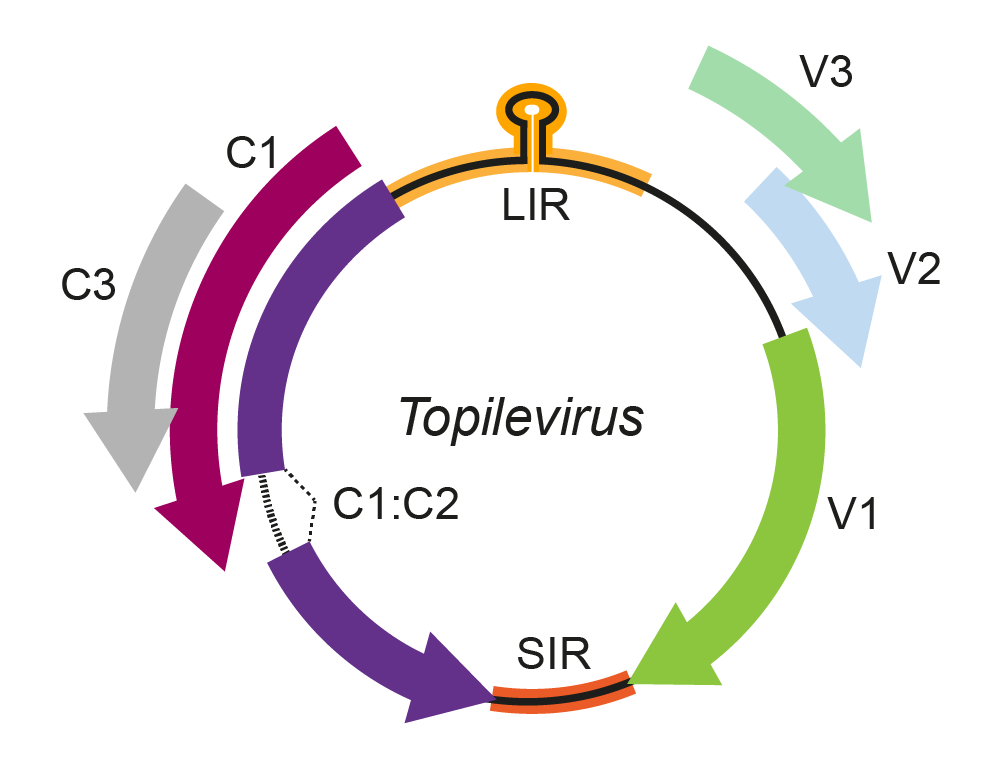
The topilevirus genomic arrangement contains an intergenic region that encloses the stem-loop structure which includes the nonanucleotide 5′-TAATATTAC-3′, highly conserved at the origin of virion strand replication in geminivirus genomes. Isolates of the type species, Tomato apical leaf curl virus, have three ORFs in the virion sense strand (V1, V2, and V3) and three in the complementary sense strand (C1, C2, and C3) (Figure 1. Topilevirus). By analogy with mastreviruses, the C1 and C2 ORFs likely encode the replication-associated protein (Rep) from a spliced transcript, and RepA is encoded from the unspliced C1 transcript. C3 is entirely embedded in the C1 ORF. Based on its position in the genome and sequence identity with other geminiviruses, the V1 ORF is predicted to encode the topilevirus coat protein and V2 ORF may encode a movement protein. C3 and V3 ORFs encode for proteins with an unknown function (Vaghi Medina et al., 2018).
|
|
|
Figure 1. Topilevirus. Genomic organization of topileviruses. ORFs are denoted as being encoded on the virion-sense (V) or complementary-sense (C) strand. The position of the stem-loop motif containing the conserved 5′-TAATATTAC-3′ sequence in the intergenic region (LIR) is shown. An intron is predicted to occur between ORFs C1 and C2. SIR, short intergenic region. |
Like mastreviruses, capulaviruses, grabloviruses and citlodaviruses, the genome of topileviruses contains two intergenic regions, the long intergenic region (LIR) where the v-ori and transcription start sites are located, and the short intergenic region (SIR) that would contain the complementary-strand replication origin and transcription termination sites.
Biology
Host range
Members of the two species included in this genus, tomato apical leaf curl virus (Vaghi Medina et al., 2018) and tomato associated geminivirus 1 (TGV1) (Fontenele et al., 2017) infect the same host plant, tomato, and TGV1 has been also isolated from Cleome sp. plants.
Species demarcation criteria
Isolates of species in the genus shares less than 65% nucleotide sequence identity with all other known geminiviruses. The tentative species demarcation criterion of 78% has been defined, but this value could change when additional topilevirus sequences become available.