Family: Geminiviridae
Genus: Begomovirus
Distinguishing features
Begomoviruses constitute a group of plant viruses which have emerged in the recent decades as serious threats to the production of many vegetable, root and fiber crops in the tropical, subtropical and temperate regions of the world (Navas-Castillo et al., 2011). Begomoviruses have monopartite or bipartite genomes, are whitefly-transmitted and are found in both the Old (both genome types) and New World (mostly bipartite genomes, with few monopartite genome viruses) (Brown et al., 2015). The whitefly vector Bemisia tabaci (Genn.) is currently considered to be a complex of cryptic species (De Barro et al., 2011). Whitefly vector specificity is associated with specific amino acid sequences in the viral coat protein (Briddon et al., 1990). Begomoviruses infect only dicot hosts. Bipartite begomoviruses encode between seven and eight proteins, and monopartite begomoviruses encode five or six proteins.
Three classes of circular ssDNA satellites have been described associated with begomoviruses: betasatellites, alphasatellites and deltasatellites (Zhou 2013, Lozano et al., 2016). Alphasatellites are related to the replication-associated protein (Rep)-encoding components of nanovirids. Betasatellites and deltasatellites contain sequences non-homologous to viruses except for a 5′-TAATATTAC-3′ sequence located in the loop of a potential stem-loop structure within the intergenic region (IR). Alphasatellites need the helper begomovirus for movement in plants and transmission by B. tabaci and betasatellites and deltasatellites also rely on these viruses for replication. Betasatellites in many cases modulate virulence by suppression of host gene silencing. In contrast, the contribution of alphasatellites to the begomovirus infection cycle is not clear. Deltasatellites do not encode any proteins but some of them affect viral DNA accumulation and symptomatology (Ferro et al., 2021).
Virion
See discussion under family description.
Genome organization and replication
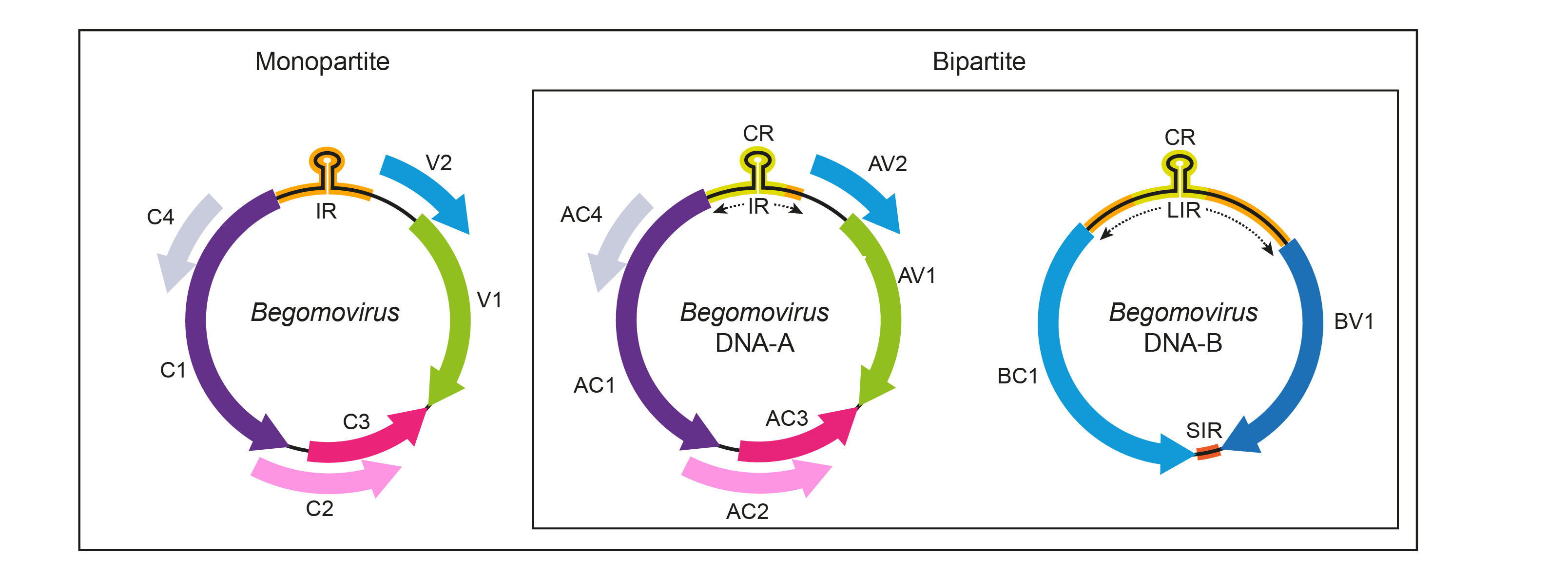
The genomes of bipartite begomoviruses consist of two components, referred to as DNA-A and DNA-B, each of 2.5–2.6 kb. The DNA-A component of the bipartite begomoviruses can replicate autonomously and produce virions but requires the DNA-B component for systemic infection. Cognate DNA-A and DNA- B components share approximately 200 b of sequence within the intergenic region (termed the "common region", CR), encompassing the conserved stem-loop with the 5′-TAATATTAC-3′ sequence at the v-ori (Figure 1. Begomovirus). The organization of ORFs in the genomes of monopartite begomoviruses resembles that seen in the bipartite viral DNA-A component (Hanley-Bowdoin et al., 2000, Navas-Castillo and Fiallo-Olivé 2021).
|
|
|
Figure 1. Begomovirus. Genomic organization of begomoviruses. The ORFs are denoted as being encoded on the virion-sense (V) or complementary-sense (C) strand, preceded by component designation (A or B) if bipartite. Corresponding protein products are indicated. ORF AV2/V2 is not present in begomoviruses from the New World. The "common region" that is shared between the two genomic components of bipartite viruses is shown as light green boxes within the intergenic region (IR) / long intergenic region (LIR). The position of the stem-loop containing the conserved 5′-TAATATTAC-3′ sequence located in the intergenic region (IR) or LIR is shown. CP, coat protein; Rep, replication-associated protein; TrAP, transcriptional activator protein; REn, replication enhancer protein; MP, movement protein; NSP, nuclear shuttle protein; SIR, short intergenic region. |
The DNA-A (and the monopartite genome) virion-sense strand encodes both the coat protein (ORF AV1/V1) that encapsidates the virion-sense ssDNA and may be involved in virus movement, and the AV2/V2 protein (ORF AV2/V2), which has also been implicated in virus movement. New World begomoviruses lack the AV2/V2 ORF. The DNA-A complementary-sense strand encodes the replication-associated protein (ORF AC1/C1), a transcriptional activator protein (ORF AC2/C2), a replication enhancer protein (ORF AC3/C3) and the AC4/C4 protein (ORF AC4/C4) (Hanley-Bowdoin et al., 2000). Rep initiates viral DNA replication by binding to iterated motifs (iterons) within the intergenic region and introducing a nick into the conserved 5′-TAATATT↓AC-3′ sequence (Laufs et al., 1995, Fontes et al., 1994). Rep also binds to the plant homologue of retinoblastoma protein (Rb) to regulate cell-cycle progression, altering the environment of terminally differentiated cells to provide host factors that support viral DNA replication (Arguello-Astorga et al., 2004). TrAP transactivates expression of virion-sense gene expression from both DNA-A and DNA-B, and also functions in the suppression of transcriptional and post-transcriptional gene silencing (Sunter and Bisaro 1992, Bisaro 2006). REn is required for efficient viral DNA replication (Sunter et al., 1990). AC4 is an important symptom determinant implicated in cell-cycle control, and C4 may counter a host response to Rep expression (Hanley-Bowdoin et al., 2013). The DNA-B encodes a nuclear shuttle protein (ORF BV1) on the virion-sense strand and a movement protein (ORF BC1) on the complementary-sense strand (Noueiry et al., 1994, Sanderfoot and Lazarowitz 1996). It has been demonstrated for a begomovirus, tomato yellow leaf curl virus, that in addition to the host plants its replication also occurs in the salivary glands of B. tabaci (He et al., 2020).
The begomoviruses further display a clear subdivision into four phylogenetic groups: Old World, New World, and the so-called "legumoviruses" and "sweepoviruses" (Ilyas et al., 2009, Trenado et al., 2011). Old World begomoviruses can be either mono- or bipartite, contain an AV2/V2 ORF. New World begomoviruses are mostly bipartite and do not contain an AV2/V2 ORF. Legumoviruses and sweepoviruses are associated, respectively, with legumes and with sweet potato.
Satellites
Circular ssDNA satellites of approximately half (betasatellites and alphasatellites) or quarter (deltasatellites) begomovirus size are associated with some begomoviruses. Betasatellites (previously known as DNA-β) are associated with many Old World monopartite begomoviruses. They are approximately 1.3 kb and contain a single ORF termed βC1, the product of which has been shown to function as a suppressor of host gene silencing (Cui et al., 2005). This has the effect of dramatically increasing the virulence of the helper begomovirus. The only homologues of begomoviral sequences found in betasatellites are the stem-loop and 5′-TAATATTAC-3′ sequences. The remainder of the satellite sequence is otherwise not related to the helper begomovirus (Briddon and Stanley 2006). Betasatellites depend on the helper begomoviruses for their replication and encapsidation, and are, in some cases, essential for the maintenance of disease in the field (Mansoor et al., 2003, Saunders et al., 2000). They are thought to be promiscuous in that they may associate and function with more than a single helper begomovirus. Autonomously replicating nanovirus-like satellites are frequently associated with the begomovirus/betasatellite complexes, and are referred to as alphasatellites (previously known as DNA-1) (Saunders et al., 2000). At least three types of alphasatellites are known (Rosario et al., 2013). One type (type 1) has been shown to ameliorate symptoms when co-infecting plants with the helper virus and its associated betasatellite, suggesting that it down-regulates virulence to some degree, possibly by reducing the accumulation of betasatellite molecules (Idris et al., 2011). Alphasatellites of types 2 and 3 have been found in association with bipartite begomoviruses in the New World (Brazil, Cuba and Venezuela) (Paprotka et al., 2010, Romay et al., 2010).
Deltasatellites are approximately 0.7 kb in size. They have been found associated with monopartite Old World begomoviruses (Dry et al., 1997), bipartite New World begomoviruses (Fiallo-Olivé et al., 2012) and sweepoviruses (Lozano et al., 2016). Unlike betasatellites and alphasatellites, deltasatellites do not encode for any protein. They need the helper begomovirus for replication and movement in plants and transmission by B. tabaci. The presence of deltasatellites in most cases do not affect the symptoms caused by the helper begomoviruses, and its effect on viral DNA accumulation depends on the helper virus–host plant combination (Ferro et al., 2021, Fiallo-Olivé et al., 2016, Hassan et al., 2016).
Biology
Host range
Collectively, begomoviruses infect a wide range of dicotyledonous plants although, individually, most have limited host ranges (Rojas et al., 2005).
Transmission
Begomoviruses are transmitted by the whitefly Bemisia tabaci, a cryptic species complex. Many begomoviruses are differentially transmitted by different species of the B. tabaci complex (Fiallo-Olivé et al., 2020). Some begomoviruses are experimentally transmissible by mechanical inoculation, although most require either Agrobacterium-mediated transfer (agroinoculation) from partially or tandemly repeated cloned genomic DNA or biolistic delivery of cloned genomic DNA for their experimental transmission (Rojas et al., 2005).
Antigenicity
Serological tests show all begomoviruses to be antigenically closely related. The use of monoclonal antibodies has shown that begomoviruses may be grouped geographically based on shared epitopes (Harrison and Robinson 1999).
Species demarcation criteria
The following criteria should be used as a guideline to establish taxonomic status:
- Number of genomic components: presence or absence of a DNA-B component.
- Organization of the genome: presence or absence of ORF AV2.
- Nucleotide sequence identity: the complete genome nucleotide sequence (DNA-A sequence in the case of bipartite begomoviruses) is required. Members of different species have <91% shared nt identity (for full-length genome of monopartite or full-length DNA-A for bipartite begomoviruses), based on pairwise alignments with pairwise deletion of gaps (Brown et al., 2015). As long as sequence identities are calculated using true pairwise alignments with pairwise deletion of gaps, this criterion can be applied for the description of new species in the absence of additional data. However, when approaching the cut-off value, the decision should take into account the biological properties of the virus. The taxonomic status of a recombinant will depend on its relatedness to the parental viruses, the frequency and extent of recombination events, and its biological properties compared with the parental viruses. Information concerning the diversity of related recombinants may be helpful to determine status.
- Trans-replication of genomic components: the inability of a Rep protein of one virus to trans-replicate genomic component(s) of another virus suggests the two viruses should be considered members of distinct species. However, small changes in the Rep binding site of otherwise identical viruses might prevent functional interaction and, equally, recombination involving a small part of the genome may render even highly diverged species trans-replication competent.
- Production of viable pseudorecombinants: account should be taken of the fitness of the pseudorecombinant in the natural host(s) of the parental viruses. It should be ensured that pseudorecombinant viability is not simply the result of inter-component recombination.
- Natural host range and symptom phenotypes may be associated with a particular species but their most common use will be to distinguish strains.
The large number of completely sequenced begomovirus isolates allows for a useful and convenient subdivision of several species into well defined strains (Fauquet and Stanley 2005), and the Geminiviridae Study Group has proposed a strain demarcation boundary of 94% pairwise sequence identity.