Family: Geminiviridae
Genus: Mulcrilevirus
Distinguishing features
Mulcrileviruses have been described from symptomatic plants (Morus alba or Broussonetia papyrifera) from China. This genus includes two species, Mulberry crinkle leaf virus (Lu et al., 2015, Ma et al., 2015) and Paper mulberry leaf curl virus 1 (Qiu et al., 2020). Members of the genus contain monopartite genomes with the conserved origin of replication (5′-TAATATTAC-3′).
Virion
See discussion under family description.
Genome organization and replication
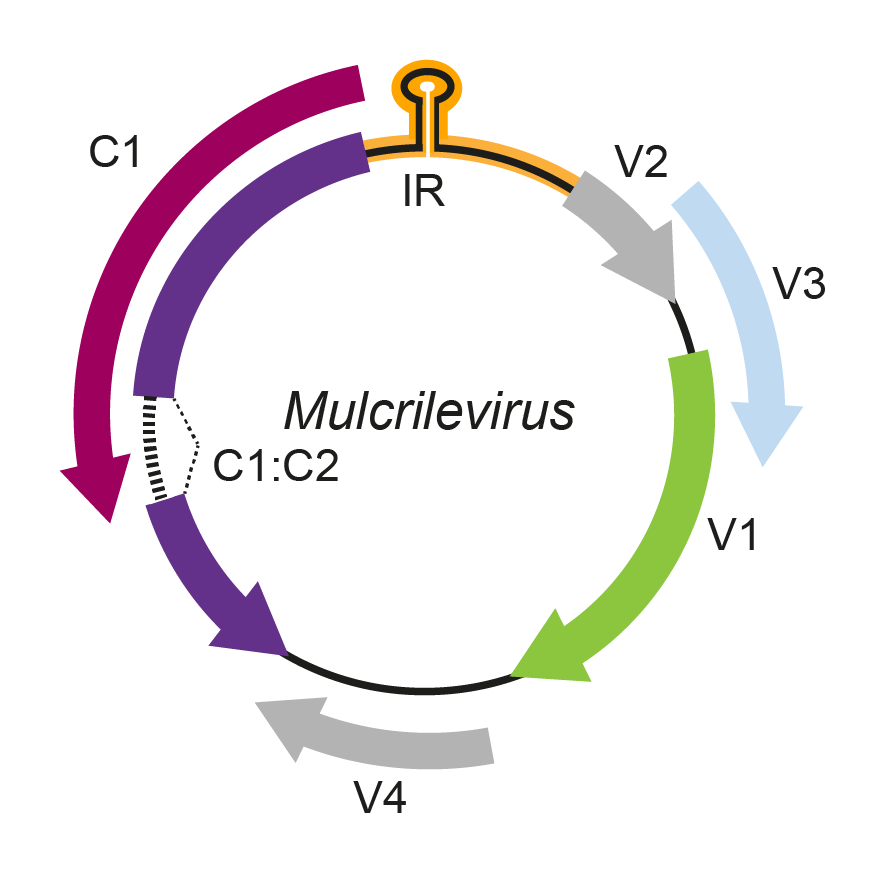
The mulcrilevirus genomic arrangement contains a putative stem-loop structure that encloses the nonanucleotide 5′-TAATATTAC-3′, highly conserved at the origin of virion strand replication in geminivirus genomes. The genome contains six open reading frames. The complementary strand of the genome potentially encodes geminivirus-like RepA and/or Rep proteins. Rep and RepA proteins are expressed from an alternatively spliced complementary strand transcript, as it was identified by small RNA (sRNA)-seq and RNA-seq reads mapping of paper mulberry leaf curl virus 1 antisense transcripts (Qiu et al., 2020). The virion sense genome strand potentially encodes a coat protein, movement protein, and two other hypothetical proteins (Figure 1. Mulcrilevirus).
|
|
|
Figure 1. Mulcrilevirus. Genomic organization of mulcrileviruses.ORFs are denoted as being encoded on the virion-sense (V) or complementary-sense (C) strand. The position of the stem-loop motif containing the conserved 5′-TAATATTAC-3′ sequence in the intergenic region (IR) is shown. |
Biology
Host range
Isolates of mulberry crinkle leaf virus have been isolated from symptomatic mulberry (Morus alba) trees (Lu et al., 2015, Ma et al., 2015) and those from paper mulberry leaf curl virus 1 from symptomatic paper mulberry (Broussonetia papyrifera) trees (Qiu et al., 2020).
Transmission
Recently, the leafhopper Tautoneura mori has been identified as the vector of mulberry crinkle leaf virus. The infected mulberry plants did not show any virus-like symptoms, as happen in mulberry plants experimentally infected with an infectious clone (Lu et al., 2021).
Species demarcation criteria
Isolates of species in the genus shares less than 61% nucleotide sequence identity with all other known geminiviruses. The tentative species demarcation criterion of 78% has been defined, but this value could change when additional mulcrilevirus sequences become available.