Family: Geminiviridae
Genus: Opunvirus
Distinguishing features
The single member of this genus is Opuntia virus 1. Isolates of Opuntia virus 1 (OpV1) have been described from asymptomatic plants belonging to 20 cactus species and from cactus-feeding cochineal insects (Dactylopius sp.). OpV1 is clearly related to previously recognized geminiviruses, based on genome composition, similarities in the origin of replication (5′-TAATATTAC-3′), and the presence of homologous genes (Fontenele et al., 2020).
Virion
See discussion under family description.
Genome organization and replication
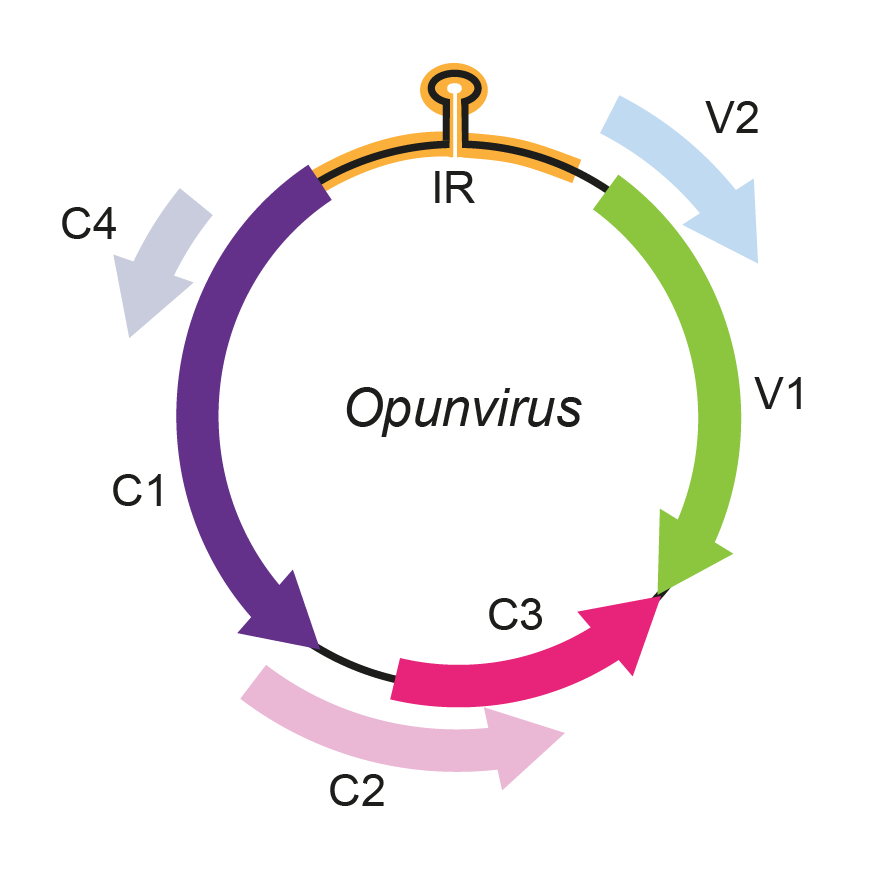
The opunvirus genomic arrangement contains an intergenic region that encloses the stem-loop structure which includes the nonanucleotide 5′-TAATATTAC-3′, highly conserved at the origin of virion strand replication in geminivirus genomes. The genome organization of the OpV1 sequences resembles that of monopartite Old World begomoviruses with six open reading frames (coat protein and movement protein in the viral sense and Rep, TrAP, Ren, C4 protein in the complementary sense) (Figure 1. Opunvirus).
|
|
|
Figure 1. Opunvirus. Genomic organization of opunviruses. ORFs are denoted as being encoded on the virion-sense (V) or complementary-sense (C) strand. The position of the stem-loop motif containing the conserved 5′-TAATATTAC-3′ sequence in the intergenic region (IR) is shown. |
Biology
Host range
Members of the the single species in this genus have been described from asymptomatic New World Cactaceae plants (belonging to 20 different species from both the Cactoideae and Opuntioideae subfamilies) and from cactus-feeding cochineal insects (Dactylopius sp.) (Fontenele et al., 2020).
Species demarcation criteria
Opuntia virus 1 genomes share >78.4% genome-wide pairwise identity with each other. The tentative species demarcation criteria of 78% have been defined, but this value could change when additional opunvirus sequences become available.