Subfamily: Densovirinae
Genus: Iteradensovirus
Distinguishing features
These viruses are monophyletic and have similar genome sizes, genetic strategies, coding patterns, homotelomeric termini, and typical VP1-encoded PLA2 domains. Five species are recognized, which all contain viruses that infect insects from the order Lepidoptera.
Virion
See discussion under family description.
Genome organization and replication
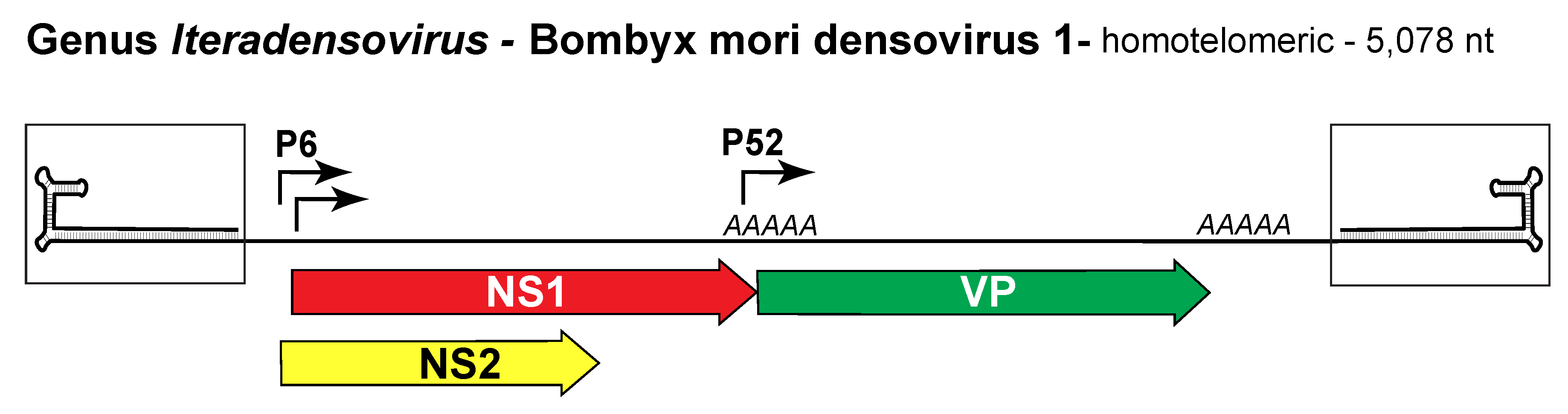
Transcription is monosense, with one mRNA promoter positioned upstream of each ORF (Figure 1. Iteradensovirus). The nonstructural proteins NS1 and NS2 are encoded in separate ORFs, while the VP ORF gives rise to 4 size variants of the capsid protein using a leaky scanning translation mechanism. In contrast to members of the other 4 species, Bombyx mori densovirus 1 (BmDV1, species Lepidopteran iteradensovirus 1) has a 45 nt direct repeat in the intergenic region.
|
|
|
Figure 1. Iteradensovirus. Genetic strategy of Bombyx mori densovirus 1 (BmDV1), genus Iteradensovirus. The homotelomeric genome of BmDV1 is shown as a single line terminating in boxed J-shaped hairpin structures that are magnified relative to the rest of the genome. At each terminus TRs of 230 nt end in 159 nt hairpin structures (Yu and Tijssen 2014). Major open reading frames encoding proteins are shown as arrowed boxes, and coloured as in previous legends. Solid arrows represent transcriptional promoters; AAAAA indicate polyadenylation sites. |
Biology
Natural infection appears confined to a single host species and replication is restricted to larval tissues. Since these viruses are pathogenic for pests of economically important plants, they have been studied as possible pest control agents. Infection typically prevents larvae from morphing into pupae, but monarch butterfly larvae remain active (although their pupae die and are a source of virus).
Species demarcation criteria
Viruses within a species are monophyletic and encode replication initiator proteins (called NS1 or Rep1, 68, or 78) that show >85% amino acid sequence identity.