Subfamily: Densovirinae
Genus: Hepandensovirus
Distinguishing features
The hepandensoviruses were known previously as hepatopancreatic parvoviruses [HPV] of prawns and shrimp. This genus shares a deep branch of the parvovirus phylogenetic tree with members of genera Penstyldensovirus and Brevidensovirus (Figures 5. Parvoviridae and 6A. Parvoviridae). Hepandensoviruses have relatively large (~6.3 kb) heterotelomeric genomes that lack discernable PLA2 motifs, and use a monosense transcription strategy. Genomic sequences of viruses from many different parts of the world are available, and all fall within the demarcation criteria for a single species, Decapod hepandensovirus 1.
Virion
See discussion under family description.
Genome organization and replication
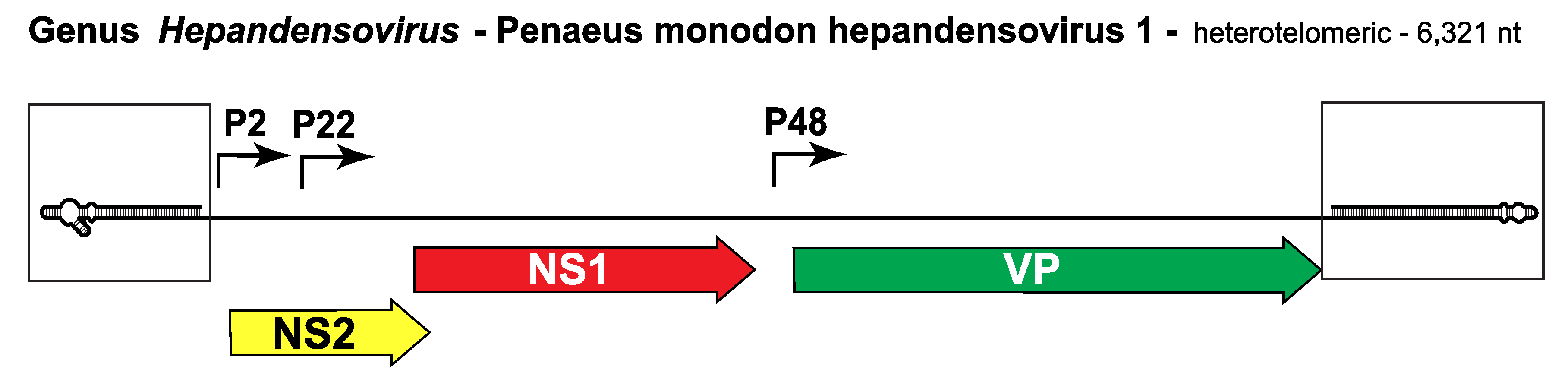
Genome organization is shown in Figure 1. Hepandensovirus.
|
|
| Figure 1. Hepandensovirus. Genetic strategy of Penaeus monodon hepandensovirus 1 (PmoHDV1), genus Hepandensovirus. The genome of PmoHDV1 is shown as a single line terminating in boxed hairpin structures that are magnified relative to the rest of the genome: the hairpin on the left is 136 nt, that on the right is 170 nt (Sukhumsirichart et al., 2006). Major open reading frames (ORFs) encoding proteins are shown as arrowed boxes. ORFs encoding the replication initiator protein, NS1, are shaded red, those encoding VP proteins are green, and those encoding NS2 are yellow. Solid arrows represent transcriptional promoters. |
Biology
These viruses are highly pathogenic in the wild, causing hepatopancreatic disease, and can constitute an economic threat in cultured shrimp populations on the rare occasions when larvae from wild-caught shrimp are introduced (Dhar et al., 2014).