Family: Caulimoviridae
Genus: Soymovirus
Distinguishing features
Members of this genus have virions and cytoplasmic inclusions similar to those of viruses in the genera Caulimovirus, Cavemovirus and Petuvirus, from which they differ in genome organization and phylogenetic placement based on analysis of polymerase gene sequences. Soymoviruses have seven or eight ORFs and can be distinguished from their closest relatives in the genus Caulimovirus by the presence of three ORFs (ORF1b, ORF2 and ORF3) between the movement protein and coat protein-coding ORFs. The negative-sense strand primer-binding site is also located in ORF1b or in a small intergenic region immediately downstream of this ORF. Furthermore, there is no intergenic region between ORF5 and ORF6, a feature of members of the genus Caulimovirus.
Virion
Morphology
Virions are isometric and about 50 nm in diameter.
Physicochemical and physical properties
When virion preparations of blueberry red ringspot virus (BRRSV) are centrifuged in sucrose or CsCl gradients, two components are observed with buoyant densities of 1.30 and 1.40 g cm−3, respectively. Virions from the two fractions have no morphological differences.
Nucleic acid
Virions contain a single molecule of non-covalently closed circular dsDNA of 8.1–8.3 kbp. The negative-sense strand DNA has a single discontinuity and the positive-sense strand DNA, two discontinuities.
Genome organization and replication
The genome contains seven or eight ORFs (ORFs 1a, 1b, 2–7, Figure 1 Soymovirus). There is one large intergenic region between ORF6 and ORF7, in which the pregenomic RNA promoter and the polyadenylation signal are located. Additionally, for blueberry red ringspot virus (BRRSV) and Cestrum yellow leaf curling virus (CmYLCV), there is a small intergenic region downstream of ORF1b. The location of the negative-sense strand primer-binding site is either within ORF1b [soybean chlorotic mottle virus (SbCMV) and peanut chlorotic streak virus (PCSV)] or in the small intergenic region downstream of ORF1b (BRRSV and CmYLCV). ORF1b and ORF7 are non-essential for replication and systemic infection and whether they are expressed is unknown. Furthermore, ORF7 is absent in CmYLCV. Protein P1a is the movement protein, the function of P2 is unknown, P3 is the virion-associated protein, P4 is the coat protein precursor, P5 is the polymerase polyprotein (aspartic protease, reverse transcriptase and RNaseH enzymatic activities) and P6 is the translation transactivator.
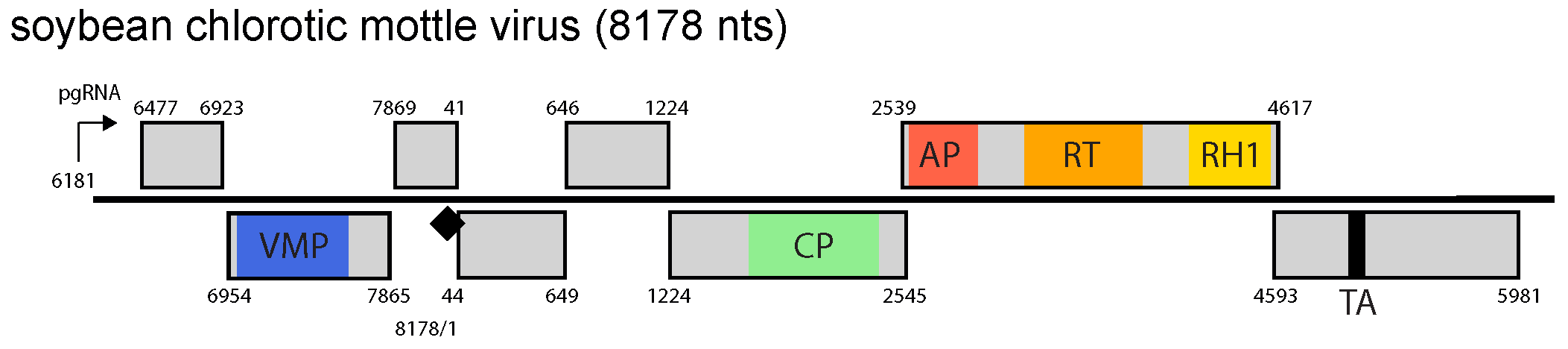 |
| Figure 1 Soymovirus. Soymovirus genome organization. The linearized map begins at the pgRNA transcription start site (black arrow, mapped or predicted ca. 32 nts downstream of TATA box; see (Pooggin et al., 1999)and references therein). The numbering begins from the first nucleotide of the Met-tRNA primer binding site (black diamond). Light grey boxes mark open reading frames (ORFs). Conserved protein domains as listed in the Pfam database (http://xfam.org) are colored: blue is the viral movement protein (VMP) (PF01107), red is the retropepsin (pepsin-like aspartic protease) (AP) (CD00303), orange is the reverse transcriptase (RT) (CD01647) and yellow is the RNase H1 (RH1) (CD06222). The conserved C-terminus of the coat protein (CP) is marked green. The conserved translation transactivator (TA) domain is shown in black. |
Two major RNA transcripts, the first representing the pregenomic RNA (about 8.2 kbp) and the second, a monocistronic RNA (1.8 kbp) containing ORF6, have been observed for SbCMV. No putative promoter sequences could be identified upstream of the 5′-end of the smaller RNA species. The pregenomic RNA serves as a polycistronic mRNA and protein P6 enables expression of the downstream ORFs. Electron-dense inclusion bodies (viroplasms) are visible in infected cells and are likely to fulfill a similar role to those observed in plants infected by members of the genus Caulimovirus. The 5′-leader sequence of the pregenomic RNA of SbCMV is not predicted to fold into a strong secondary structure as for other members of the family Caulimoviridae and this likely prevents ribosome shunt mechanism of initiation of translation of ORF1 (Pooggin and Ryabova 2018).
Biology
Host ranges are narrow (one to two plant families). All soymoviruses, but BRRSV, are mechanically transmissible. Spread of the viruses in the field is observed although the vectors are unknown. BRRSV is spread by clonal propagation of its host.
Antigenicity
Virions are moderately immunogenic if fixed in 0.5% (v/v) formaldehyde. Serological relationships between viruses belonging to distinct species in this genus are unknown, but no cross-reactivity with members of the genus Caulimovirus has been observed.
Species demarcation criteria
The criteria demarcating species in the genus are:
- Host range
- Differences in polymerase (RT + RNAse H) nucleotide sequences of more than 20%

