Family: Caulimoviridae
Genus: Petuvirus
Distinguishing features
Petunia vein clearing virus (PVCV), a member of the sole species in the genus, Petuvirus venapetuniae, is distinguishable from all other members of the Caulimoviridae except for vacciniviruses by its simple genome organization (one ORF) and phylogenetic placement based upon analysis of polymerase gene sequences.
Virion
Morphology
Virions consist of isometric, non-enveloped particles with an average diameter of 43–46 nm. Virions are isometric in shape (Richert-Pöggeler and Lesemann 2007).
Nucleic acid
Virions contain a single molecule of non-covalently closed circular dsDNA of about 7.2 kbp.
Proteins
The presence of multiple consensus sequences for proteases in the viral polyprotein indicates post-translational processing. (Stavolone et al., 2001) identified functional coiled-coil domains which could be part of movement- and virion-associated proteins. Such domains are required for homo- and hetero-oligomerization. They are conserved among members of the family Caulimoviridae. A putative viral suppressor of gene silencing has been identified at the C-terminal end of the PVCV polyprotein (Kuriyama et al., 2020).
Genome organization and replication
The genome contains a single ORF encoding a large polyprotein and an intergenic region containing the pregenomic RNA promoter, a polyadenylation signal and the negative-sense strand primer-binding site (Figure 1 Petuvirus). Protein domains characteristic of the movement protein, capsid protein, aspartic protease, reverse transcriptase and RNase H1 are present in the ORF1 polyprotein in that order. There is also a putative virion-associated protein homolog between the movement protein and capsid protein domains of the polyprotein (Richert-Pöggeler and Shepherd 1997).
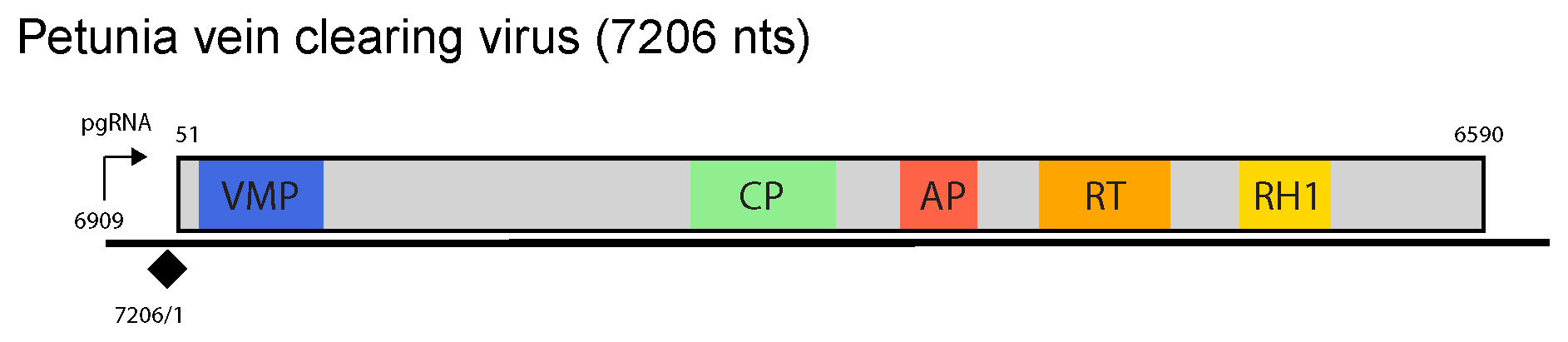 |
| Figure 1 Petuvirus. Petuvirus genome organization. The linearized map begins at the pgRNA transcription start site (black arrow, mapped or predicted ca. 32 nts downstream of TATA box; see (Pooggin et al., 1999) and references therein). The numbering begins from the first nucleotide of the Met-tRNA primer binding site (black diamond). Light grey boxes mark open reading frames (ORFs). Conserved protein domains as listed in the Pfam database (http://xfam.org) are colored: blue is the viral movement protein (VMP) (PF01107), red is the retropepsin (pepsin-like aspartic protease) (AP) (CD00303), orange is the reverse transcriptase (RT) (CD01647) and yellow is the RNase H1 (RH1) (CD06222). The conserved C-terminus of the coat protein (CP) is marked green. |
The 5′-leader sequence of the pregenomic RNA folds into a stable hairpin structure. A ribosome shunt mechanism of initiation of translation is assumed to be utilized (Pooggin et al., 1999). The polyprotein is processed by the aspartic protease to generate mature proteins. How the timing and level of expression of the different proteins is coordinated during the replication cycle is unknown. Infected cells contain electron-dense inclusion bodies similar to those produced by infections of members of the genus Caulimovirus.
Biology
Infection of Petunia hybrida is due to activation of replication-competent endogenous PVCV (ePVCV) elements in the nuclear genome of the plant (Richert-Pöggeler et al., 2003). Wounding (e.g. plant pruning), vegetative propagation, water stress as well as aging can induce infection. The virus is vertically transmitted to all progeny as part of the host’s genome and is also graft-transmissible to Petunia parodii and Nicotiana glutinosa. PVCV has no known insect vector and it is not mechanically transmissible.
Antigenicity
Purified PVCV preparations seem to be weakly immunogenic. No serological relationships have been detected in decoration tests with antisera to cauliflower mosaic virus (CaMV), Dahlia mosaic virus (DaMV), carnation etched ring virus (CERV) and soybean chlorotic mottle virus (SbCMV) (Richert-Pöggeler and Lesemann 2007).
Species demarcation criteria
PVCV is the sole member of the genus.

