Family: Papillomaviridae
Koenraad Van Doorslaer, Zigui Chen, Hans-Ulrich Bernard, Paul K. S. Chan, Rob DeSalle, Joakim Dillner, Ola Forslund, Takeshi Haga, Alison A. McBride8, Luisa L. Villa and Robert D. Burk
The citation for this ICTV Report chapter is the summary published as Van Doorslaer et al., (2018):
ICTV Virus Taxonomy Profile: Papillomaviridae, Journal of General Virology, 99, 989–990.
Corresponding author: Koenraad Van Doorslaer ([email protected])
Edited by: Balázs Harrach and Stuart G. Siddell
Posted: June 2018
PDF: ICTV_Papillomaviridae.pdf
Summary
The Papillomaviridae is a family of small, non-enveloped viruses with double stranded DNA genomes of 5,748 bp to 8,607 bp (Table 1. Papillomaviridae). Their classification is based on pairwise nucleotide sequence identity across the L1 open reading frame. Members of the Papillomaviridae primarily infect mucosal and keratinized epithelia, and have been isolated from fish, reptiles, birds and mammals. Despite a long co-evolutionary history with their hosts, some papillomaviruses are pathogens of their natural host species.
Table 1. Papillomaviridae. Characteristics of members of the family Papillomaviridae.
| Characteristic | Description |
| Typical member | human papillomavirus 16 (K02718), species Alphapapillomavirus 9, genus Alphapapillomavirus, subfamily Firstpapillomavirinae |
| Virion | Non-enveloped, 55 nm, icosahedral |
| Genome | Circular dsDNA. Genome varies from 5,748 bp to 8,607 bp. |
| Replication | Bidirectional theta replication |
| Translation | Early and late transcripts, alternative splicing, alternative open reading frames |
| Host Range | Mammals, reptiles, birds, and fish |
| Taxonomy | Realm Monodnaviria, kingdom Shotokuvirae, phylum Cossaviricota, class Papovaviricetes, order Zurhausenvirales: two subfamilies include 53 genera and 133 species |
Virion
Morphology
Papillomavirus virions are non-enveloped. The viral capsid is ~600 Å in diameter and consists of 360 copies (arranged as 72 pentamers) of the major capsid protein, L1, and ~12 molecules of the L2 minor capsid protein (Figure 1. Papillomaviridae; (Finnen et al., 2003)). Expression of recombinant L1 with or without L2 allows for self-assembly of virus-like particles (VLPs). Each capsid packages a single copy of the viral circular dsDNA. The packaged viral DNA is associated with core histone proteins (Larsen et al., 1987).
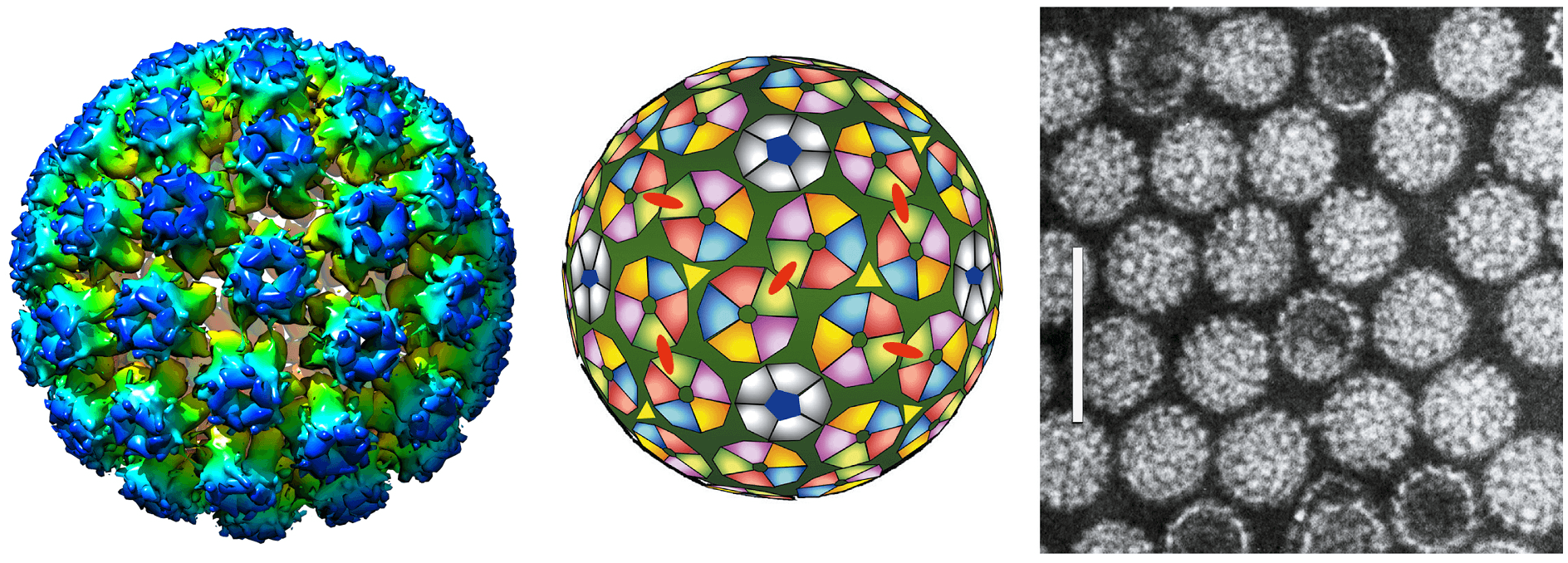 |
| Figure 1. Papillomaviridae. (Left) Atomic rendering of a papillomavirus capsid. Derived from an image reconstruction from Cryo-electron microscopy of human papillomavirus type 16 at 4.5 Å resolution and colored according to the radial coloring scheme shown (PDB: 5KEP; (Guan et al., 2017)). (Center) Schematic diagram representing the 72 capsomers in a T=7 arrangement of a papillomavirus capsid. The icosahedral structure is composed solely of pentameric capsomeres for a total of 360 capsid proteins. (Right) Negative-contrast electron micrograph of human papillomavirus 1 (HPV1) virions. The bar represents 100 nm. |
Physicochemical and physical properties
The virion Mr is 47×106. Buoyant density of virions in sucrose and CsCl gradients is 1.20 and 1.34–1.35 g cm−3, respectively. Virion S20,W is 300. Virions are resistant to a wide array of environmental and chemical treatments (Meyers et al., 2014).
Nucleic acid
The papillomavirus genome is approximately 7,500 bp. Viral genomes vary from 5,748 bp (Sparus aurata papillomavirus type 1; SaPV1) to 8,607 bp (canine papillomavirus type 1; CPV1). The genomes have an average GC content of about 42% (36–59%).
Proteins
Papillomavirus gene expression is tightly regulated at the level of transcription and RNA processing, including alternative mRNA polyadenylation and splicing (Schwartz 2013). A typical papillomavirus encodes six to nine proteins (Figure 2. Papillomaviridae). However, the ancestral papillomavirus may have only contained a core set of four proteins (E1, E2, L1, and L2). Temporal expression of the viral genome is associated with tissue differentiation (Figure 3. Papillomaviridae). The viral DNA helicase (E1) is the only viral enzyme, and is essential for replication and amplification of the viral chromosome in the nucleus of infected cells. (Bergvall et al., 2013). The viral E2 protein is the master regulator of the viral life cycle, and plays key roles in transcriptional regulation, initiation of DNA replication and partitioning the viral genome (McBride 2013). The E1^E4 gene product is typically translated from a spliced mRNA fusing approximately the first four E1 codons to the E4 ORF, present in an alternative reading frame to the E2 ORF (Doorbar 2013). A subset of viral mRNA encodes a short, hydrophobic, transmembrane protein, E5. Other than hydrophobicity, there is low sequence similarity among different E5 proteins (DiMaio and Petti 2013). The E5 proteins can be further divided into different classes based on phylogeny and hydrophobic profiles (Bravo and Alonso 2004). E5 proteins are typically encoded in the 3ʹ-end of the early coding region. However, hydrophobic proteins, located in other parts of the viral genome have also been described. Designated as E10, these non-E5 hydrophobic proteins either overprint the E6 ORF, or are located in this region in the absence of an E6 gene (Van Doorslaer and McBride 2016). The productive phase of the viral life cycle occurs in differentiated cells that have exited the cell cycle. In order to complete the viral life cycle, the virus needs to uncouple replication from differentiation (Figure 3). The E6 and E7 proteins have been shown to play key roles in usurping the cellular environment to allow for replication. The E6 protein contains two zinc-binding motifs essential for its function (Vande Pol and Klingelhutz 2013). The E7 amino terminus contains regions of similarity to conserved regions (CR) 1 and CR2 of the mastadenovirus E1A protein, and the polyomavirus large T antigen (Roman and Munger 2013). The E7 C-terminus contains a single zinc-binding motif homologous to the E6 motifs (Van Doorslaer 2013). The E6 and E7 proteins appear to be essential for members of the genus Alphapapillomavirus. Remarkably, the E6 and E7 proteins are not encoded by all papillomaviruses (Van Doorslaer 2013). The E8 exon is embedded within E1, and utilizes the same splice acceptor site as E1^E4 mRNA, generating mRNA for the E8^E2 protein. The E8^E2 viral repressor protein is present in essentially every known papillomavirus. E8^E2 inhibits viral replication and gene expression. Upon cellular differentiation, the viral capsid proteins L1 and L2 are expressed. The major capsid protein L1 is the structural component of the viral capsid (Buck et al., 2013). The minor capsid protein L2 plays an active role in viral assembly and throughout the infectious process (Wang and Roden 2013).
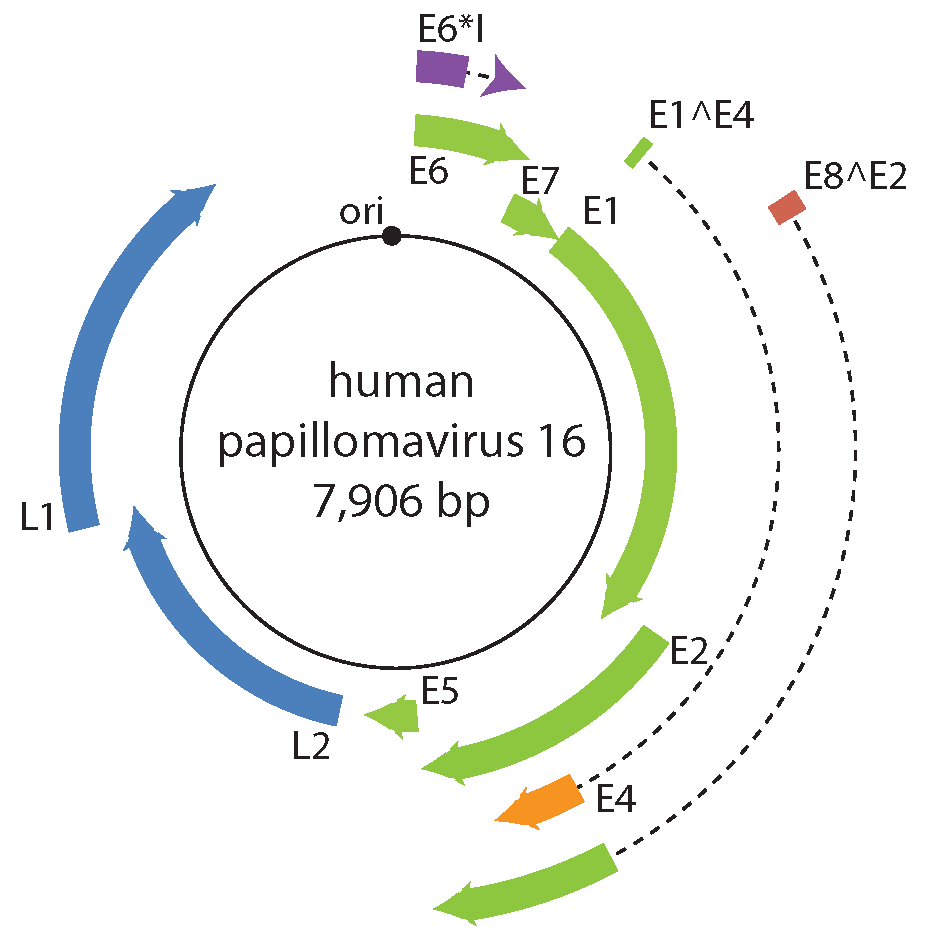 |
| Figure 2. . Papillomaviridae. (B) Diagram of the human papillomavirus 16 genome. The viral dsDNA is indicated. The outer boxes indicate the protein-coding open reading frames. Dotted lines represent intron sequences. The black circle represents the viral origin of replication (ori). |
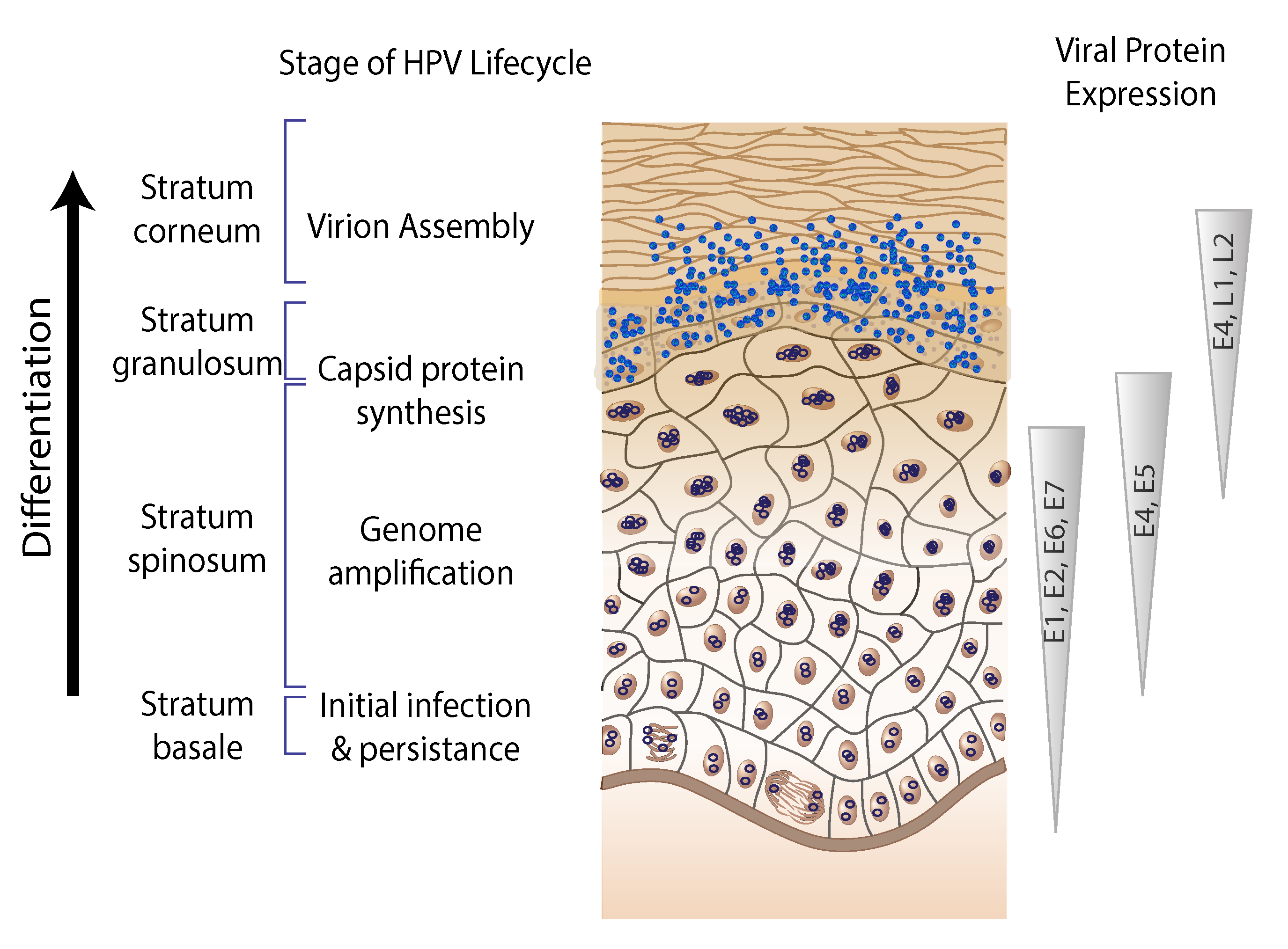 |
| Figure 3. . Papillomaviridae. Organization of the viral lifecycle. The different layers of the epithelium are shown on the left. The timing of expression and associated protein levels are summarized using triangles. Viral genome maintenance is facilitated by the expression of E6 and E7 together with E1 and E2. Increased levels of the viral replication proteins facilitate viral genome amplification. The expression of L1 and L2 allows for the formation of infectious virions (virus assembly). |
Genome organization and replication
Transcription of the circular virus genome occurs from only one DNA strand. The viral genome can be divided into three functional regions. The early region encodes viral proteins involved in transcription, replication, and manipulation of the cellular milieu. The late region encodes the capsid proteins L1, and L2. The upstream regulatory region (URR or LCR) is located between the L1 and E6 ORFs and contains the viral origin of replication as well as binding sites for viral and cellular transcription factors.
The viral replication cycle consists of three distinct phases of replication. Initial limited viral DNA amplification is supported by the viral E1 and E2 replication proteins. The viral E2 protein binds to its binding sites in the viral origin of replication, recruiting the viral E1 helicase allowing for replication. This initial burst of replication is followed by maintenance replication, during which the viral genome is maintained at a relatively low, but constant copy number in the proliferating cells of a clonally expanded population of infected cells. Finally, as an infected cell completes cellular differentiation, there is a switch towards differentiation-dependent genome amplification and eventual generation of progeny virions (McBride 2008). During maintenance replication, the virus needs to establish an S-phase like state in differentiated cells. Through a plethora of protein-protein interactions, the viral E6 and E7 proteins usurp the cellular environment, allowing for viral replication in differentiated cells. Remarkably, recent work has highlighted that different genera may induce this pseudo-S-phase through different mechanisms (White et al., 2012, Meyers et al., 2017, Brimer et al., 2017). The maintenance phase of the viral life cycle can last for months to years. In addition to regulating replication, the viral E2 protein plays a key role during maintenance by ensuring that the viral genomes are faithfully partitioned into the daughter cells. In the top layers of the differentiated epithelia, the viral DNA is amplified to a high copy number. The vegetative phase of the viral life-cycle requires the cellular DNA Damage Response (Bristol et al., 2017). The viral capsid proteins self-assemble into particles encapsidating the viral DNA. As the cells slough off into the environment, infectious virions are released, completing the viral life cycle.
Biology
Epidemiological and biological data is primarily available for the viral types belonging to the genus Alphapapillomavirus, and specifically those viruses associated with (cervical) cancer. An estimated 79 million Americans are infected, with an additional 14 million new HPV infections occurring every year (CDC 2017). HPV is spread by skin-to-skin contact, and infections with genital human papillomaviruses are the most common sexually transmitted infection (STI).
Papillomaviruses primarily infect epithelial cells. Following a micro-abrasion, the incoming virion complexes with extracellular heparin sulfate proteoglycans on the basement membrane. This interaction results in conformational changes in the L1 and L2 capsid proteins, in turn allowing for the transfer of the virion to an unknown entry receptor. Following furin cleavage of L2, the virion becomes internalized using a process that shares similarities with macropinocytosis (Campos 2017). Early trafficking events involve transporting virions from early endosomes into acidic late endosome and multivesicular bodies. This allows for capsid disassembly and uncoating. During this process, the viral DNA is believed to remain bound to L2. The L2-DNA complex traffics to the trans-Golgi network, remaining there until the onset of mitosis. During mitosis, the trans-Golgi network naturally vesiculates, and the vesicle-bound viral DNA finds its way into the nucleus. By metaphase, the viral DNA is associated with host chromosomes. Following mitosis, the viral DNA can be seen to be associated with nuclear ND10 bodies (Campos 2017).
As the life cycle is completed in cells already destined for cell death, papillomaviruses are not viremic and are hidden from the immune system. In addition, papillomaviruses have evolved a plethora of mechanisms actively limiting the interferon response, a key antiviral defense mechanism (Kanodia et al., 2007). Overall, papillomaviruses appear to effectively evade the innate immune response, thereby delaying the activation of adaptive immunity. In turn, this likely plays an important role in persistence of the virus for months or even years.
Pathogenicity
Healthy skin harbors a large spectrum of different papillomavirus types belonging to different genera. While the majority of viral infections are subclinical, certain viral types cause (cervical, anal and/or oral pharyngeal) cancers, and have been associated with an increasing number of squamous cell carcinomas at specific sites. Based on their tropism, papillomaviruses can be roughly divided into cutaneous or mucocutaneous. Epidemiologically, the mucocutaneous HPV types can be further subdivided based on whether they are associated with benign or malignant lesions (Cubie 2013). Importantly, even in the case of viral types associated with specific pathologies, the majority of infections still present as subclinical. In the cervical environment, approximately 90% of HPV infections are cleared within two years post infection. Where clearance depends on an effective cell-mediated immune response, it is not clear why some infections are able to persist. Importantly, in the case of the oncogenic alphapapillomavirus types, persistent infection, not an incident infection is the main risk factor for progression towards cancer (Burk et al., 2009). Indeed, cellular transformation and viral replication are mutually exclusive, suggesting that oncogenic progression is not a typical outcome of infection.
Antigenicity
Antigenicity is primarily determined by the major capsid protein, L1. Following vaccination, neutralizing epitopes typically map to a single variable loop (Ludmerer et al., 1997), or more commonly two non-contiguous loops (McClements et al., 2001, Christensen et al., 2001). Viral immunity appears to be highly species-specific, and there is only limited cross-protection, even to types within the same viral species. Furthermore, following natural infection, only approximately half of women seroconvert within 18 months following exposure (Carter et al., 2000). Prophylactic vaccines induce high-titer neutralizing antibodies restricted to a subset of (oncogenic) types within the genus Alphapapillomavirus. The current vaccines do not protect against types belonging to different genera. Vaccination with L2 (minor capsid protein) derived vaccines induces low-titer, yet broadly cross-neutralizing antibodies to heterologous PV types. These vaccines provide cross-protection in animal challenge models (Schellenbacher et al., 2017). Efforts to broaden the human papillomavirus (HPV) vaccines using L2 (poly)-peptides are currently underway (Schellenbacher et al., 2017).
Derivation of names
Papilloma: from Latin papilla, “nipple, pustule”, and Greek suffix -oma, used to form nouns denoting “tumors”. Viral genera belonging to the subfamily of the Firstpapillomavirinae are named according to the Greek alphabet (e.g., Alphapapillomavirus). The prefixes “Dyo-” and “Treis-” are used to accommodate the growing list of viral genera within this subfamily. The names Alphapapillomavirus, Betapapillomavirus, and Gammapapillomavirus have been used for the genera containing papillomaviruses that infect humans and are not used in combination with the Dyo- or Treis- prefixes. Genera within the Secondpapillomavirinae are named according to the Semitic abjads. Currently, the Secondpapillomavirinae contain a single genus, bearing the first letter of this alphabet (transcribed as “A” in Latin): Alefpapillomavirus.
Subfamily demarcation criteria
The current taxonomic classification of papillomaviruses is based on the nucleotide sequence of the L1 ORF. The L1 ORF of members of different subfamilies shares less than 45% sequence identity.
Genus demarcation criteria
The original criteria distinguishing genera stated: “Most types within a PV genus show less than 60% sequence identity to types of other genera based on global multiple sequence or pairwise alignments of the L1 genes. Nevertheless, the suggested percentage identities that define PV genera have to be taken as general, but not absolute criteria as curation is necessary” (de Villiers et al., 2004). Practically, papillomavirus genera are primarily delineated by visual inspection of phylogenetic trees derived from concatenated E1, E2, L1, and L2 nucleotide sequences. Efforts are underway to refine the papillomavirus classification scheme.
Relationships within the family
Phylogenetic analysis of papillomaviruses based on the concatenated alignment of four coding sequences (E1, E2, L2, and L1) from isolates from the type species of each of the 53 genera (Figure 4. Papillomaviridae) supports the existence of at least two distinct subfamilies (Firstpapillomavirinae and Secondpapillomavirinae). Likewise, this phylogeny, and that of an analysis of all 133 species (tree available in Resources), corroborates many genera and species within the Firstpapillomavirinae. However, not all genera or species are equally supported. There may be a need in the near future to base the taxonomy of the Papillomaviridae on the phylogenetic tree.
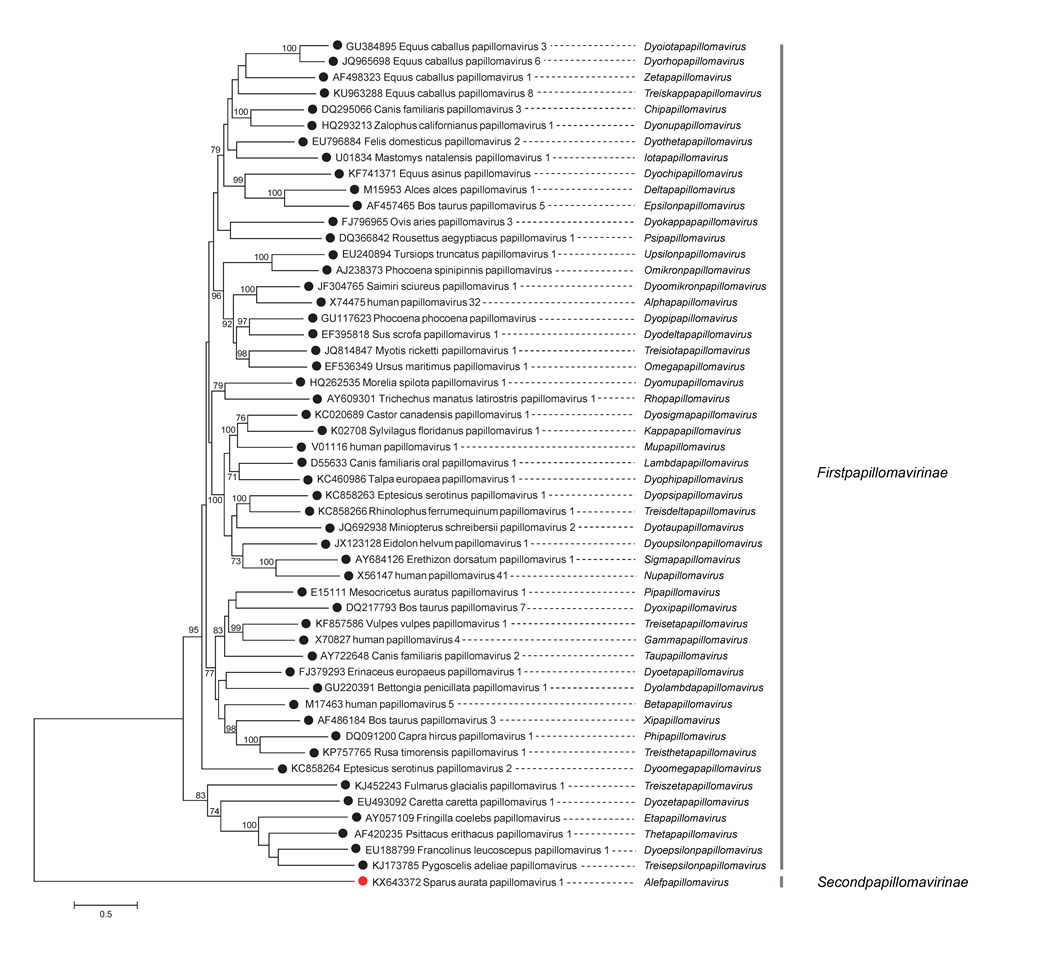 |
| Figure 4. . Papillomaviridae. Phylogenetic analysis of isolates from the type species of each papillomavirus genus. The E1, E2, L2, and L1 nucleotide sequences were translated into amino acids and aligned using MUSCLE v7.221 (Edgar 2004). A maximium likelihood tree of concatenated nucleotide sequences was produced using the optimal model of evolution (GTR + G) as determined within MEGA 7 (Kumar et al., 2016). Nodes supported by >70% of bootstrap replicates are indicated. This phylogenetic tree and corresponding sequence alignment are available to download from the Resources page, along with a complete tree including an additional 290 isolates. |
Relationships with other taxa
There is evidence of recombination between a polyomavirus and a papillomavirus. The unclassified bandicoot papillomatosis carcinomatosis virus types 1 and 2 (BPCV1 and BPCV2) have circular dsDNA genomes encoding large and small T antigens related to avian polyomaviruses and capsid proteins (L1 and L2) of a putative marsupial papillomavirus.

