Family: Potyviridae
Genus: Brambyvirus
Distinguishing features
The genus includes a single species, members of which are distinguished from all other members of the family in that they encode a very large P1 protein (83.6 kDa) containing an AlkB domain, and are also phylogenetically distinct (Susaimuthu et al., 2008).
Virion
Morphology
Virions are flexuous filaments 800×11–15 nm in size.
Nucleic acid
Virions contain a single molecule of linear, positive-sense ssRNA of about 11 kb.
Proteins
There is a single coat (capsid) protein of 40.9 kDa.
Genome organization and replication
Apart from the size of the P1 coding region, the genome organization (Figure 1.Brambyvirus) is identical to that of most monopartite viruses in the family Potyviridae (Figure 2.Potyviridae).
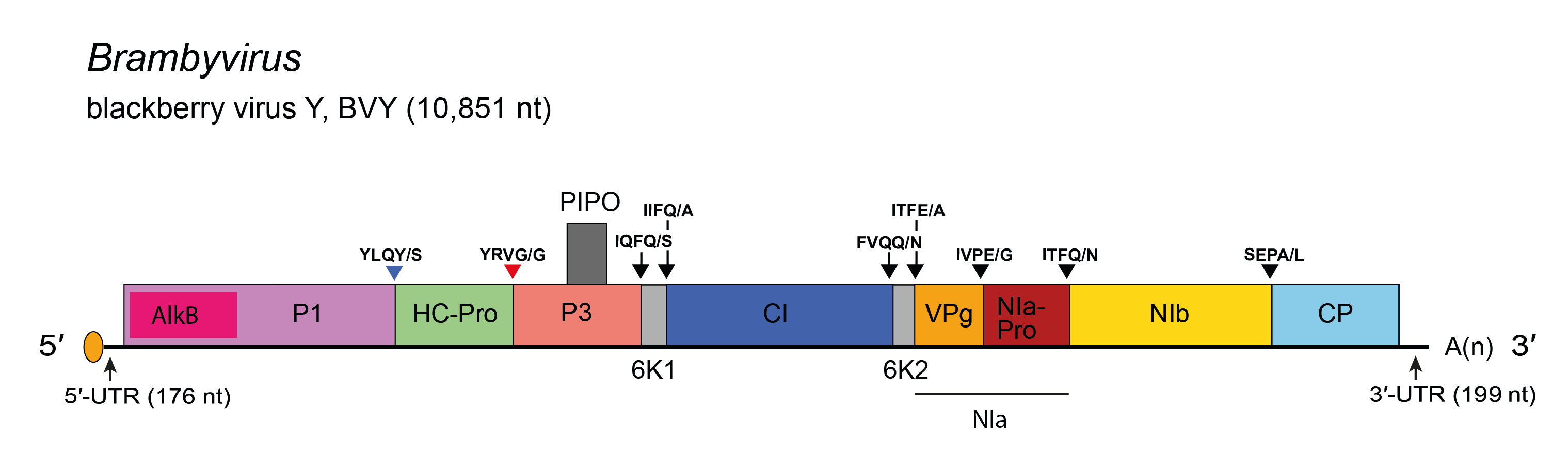 |
| Figure 1.Brambyvirus. Schematic diagram of blackberry virus Y (BVY) genome. The polyprotein ORF is indicated by the large open box divided into putative mature proteins. The pretty interesting Potyviridae protein (PIPO) is represented by a small box. The untranslated regions (UTR) are represented by lines on each end of the large ORF. Activities of mature proteins are postulated by analogy with genus Potyvirus. Conventions are as for the potyvirus genome organization map (Figure 2.Potyviridae). The P1 cistron contains an embedded AlkB domain. Not to scale. |
Biology
Host range
The virus has been reported only from wild and cultivated blackberry (Rubus sp.) where it is often symptomless but is also a component of a complex of viruses. It is not known to cause symptoms in any herbaceous test host.
Transmission
The virus is presumed to be transmitted by an aerial vector that has not yet been identified.
Antigenicity
The virus could not be detected by a universal potyvirus monoclonal antibody but there are no additional data.
Species demarcation criteria
See discussion under family description.

