Family: Natareviridae
Jens H. Kuhn, Nolwenn M. Dheilly, Sandra Junglen, Sofia Paraskevopoulou (Σοφία Παρασκευοπούλου), Mang Shi (施莽) and Nicholas Di Paola
The citation for this ICTV Report chapter is the summary published as:
Corresponding author: Nicholas Di Paola (nicholas.dipaola.civ@health.mil)
Edited by: Jens H. Kuhn and Stuart G. Siddell
Posted: November 2023
Summary
Natareviridae is a family for a negative-sense RNA virus with a genome of about 12.9 kb (Table 1.Natareviridae). This virus was found in crustaceans in China. The family includes a single genus with one species for this virus. The natarevirid genome contains four open reading frames (ORFs) that encode a nucleoprotein (NP), two glycoproteins (GPs), and a large (L) protein containing an RNA-directed RNA polymerase (RdRP) domain.
Table 1.Natareviridae. Characteristics of members of the family Natareviridae
| Characteristic | Description |
| Example | Wēnzhōu crab virus 3 (KM817603), species Charybdivirus charybdis, genus Charybdivirus |
| Virion | Unknown |
| Genome | About 12.9 kb of nonsegmented negative-sense RNA |
| Replication | Unknown |
| Translation | Unknown |
| Host range | Portunid crustaceans |
| Taxonomy | Realm Riboviria, kingdom Orthornavirae, phylum Negarnaviricota, class Monjiviricetes, order Jingchuvirales: the family includes one genus and one species. |
Virion
Morphology
Unknown.
Nucleic acid
Natarevirids have nonsegmented linear negative-sense RNA genomes with total lengths of about 12.9 kb (Li et al., 2015).
Genome organization and replication
Natarevirid genomes have four ORFs that encode two glycoproteins (GPs), a nucleoprotein (NP), and a large (L) protein (Li et al., 2015) (Figure 1.Natareviridae). The replication cycle of natarevirids remains to be elucidated.
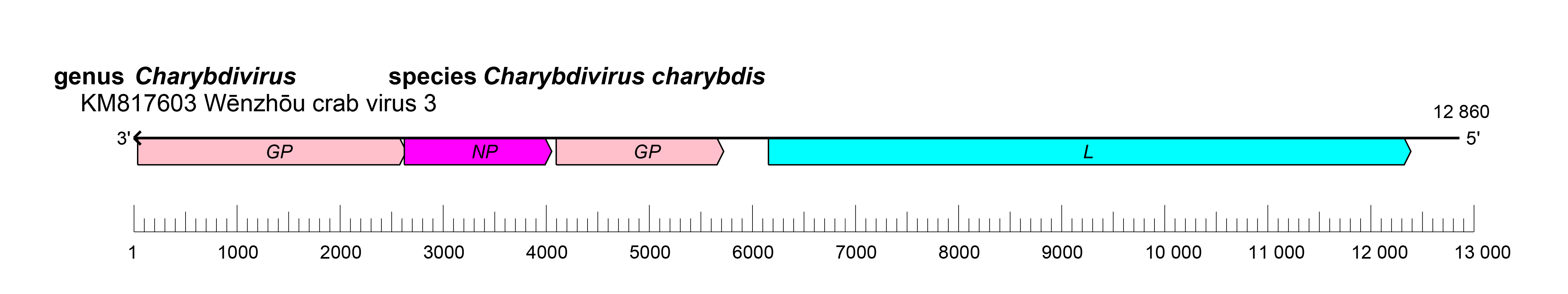 |
| Figure 1.Natareviridae. Genome organization of Wēnzhōu crab virus 3. ORFs are colored according to the predicted protein function. GP, glycoprotein gene; NP; nucleoprotein gene; L, large protein gene. |
Biology
The only classified natarevirid, Wēnzhōu crab virus 3 (WzCVV3), has been associated with portunid crustaceans (Charybdis japonica (A. Milne-Edwards, 1861)) sampled in China (Li et al., 2015). The presence of endogenized natarevirid elements (in particular, GP-gene-derived nucleic acids) in arthropod genomes indicates that natarevirids are millions of years old (Dezordi et al., 2023).
Derivation of names
Charybdivirus, charybdis: from Charybdis, a sea monster in Greek mythology
Natareviridae: from nata, an inflection of the Latin nato, meaning “swim, float”
Genus demarcation criteria
Not applicable (the family includes only a single genus).
Species demarcation criteria
Not applicable (the only genus includes one species).
Relationships within the family
Phylogenetic relationships of members of the family Natareviridae are shown in Figure 2.Natareviridae.
 |
| Figure 2.Natareviridae. Phylogenetic relationships of natarevirids. Maximum-likelihood tree (midpoint-rooted) inferred by using large protein gene (L) sequences. Sequences were initially aligned by MAFFT version 7 (https://mafft.cbrc.jp/alignment/software/) in Geneious version R9 (http://www.geneious.com) and realigned using ClustalW (https://www.genome.jp/tools-bin/clustalw). The tree was estimated using PhyML 3.0 (http://www.atgc-montpellier.fr/phyml/ , a subtree pruning and regrafting (SPR) topology searching algorithm, and a Bayes branch support algorithm. Numbers near nodes on the trees indicate bootstrap values as percentages. Tree branches are scaled to nucleotide substitutions per site (scale bar). Two mononegavirals were included as an outgroup. |
Relationships with other taxa
Viruses in the family Natareviridae are most closely related to jingchuviral aliusvirids, chuvirids, crepuscuvirids, and myriavirids (Li et al., 2015, Di Paola et al., 2022).

