Family: Filoviridae
Genus: Orthoebolavirus
Distinguishing features
Three orthoebolaviruses (Bundibugyo virus [BDBV], Ebola virus [EBOV], and Sudan virus [SUDV]) are highly lethal human pathogens. Taï Forest virus (TAFV) has caused a single recorded case of severe but nonlethal human disease. Reston virus (RESTV) has, as far as is known, only caused one inapparent human infection. Although the pathogenicity of Bombali virus (BOMV) for humans is unclear (Kuhn et al., 2020), infection studies using mice engrafted with human immune cells suggest that it might not be pathogenic in humans (Bodmer et al., 2023). Orthoebolaviruses are notable for encoding three distinct proteins from their glycoprotein (GP) genes, a strategy they share with cuevaviruses (Volchkov et al., 1995, Sanchez et al., 1996, Negredo et al., 2011).
Virion
Morphology
Virions are filamentous in shape, but can also be branched, circular, U-shaped, or 6 shaped. Spherical forms are rare to absent (Figure 1.Orthoebolavirus). Virions vary greatly in length (>20 μm) but have a diameter of 96–98 nm. Peak infectivity has been associated with particles of about 805 nm in length (Ellis et al., 1979a, Ellis et al., 1979b, Geisbert and Jahrling 1995, Ryabchikova and Price 2004, Beniac et al., 2012).
Virions are composed of a central core formed by a helical ribonucleoprotein (RNP) complex, surrounded by a matrix layer and a lipid envelope derived from the host cell plasma membrane. Polyploidy has been observed. Spikes about 7 nm in diameter and spaced at intervals of about 10 nm are seen as globular structures on the surface of virions (Geisbert and Jahrling 1995, Ryabchikova and Price 2004, Beniac et al., 2012, Sugita et al., 2018, Kirchdoerfer et al., 2019).
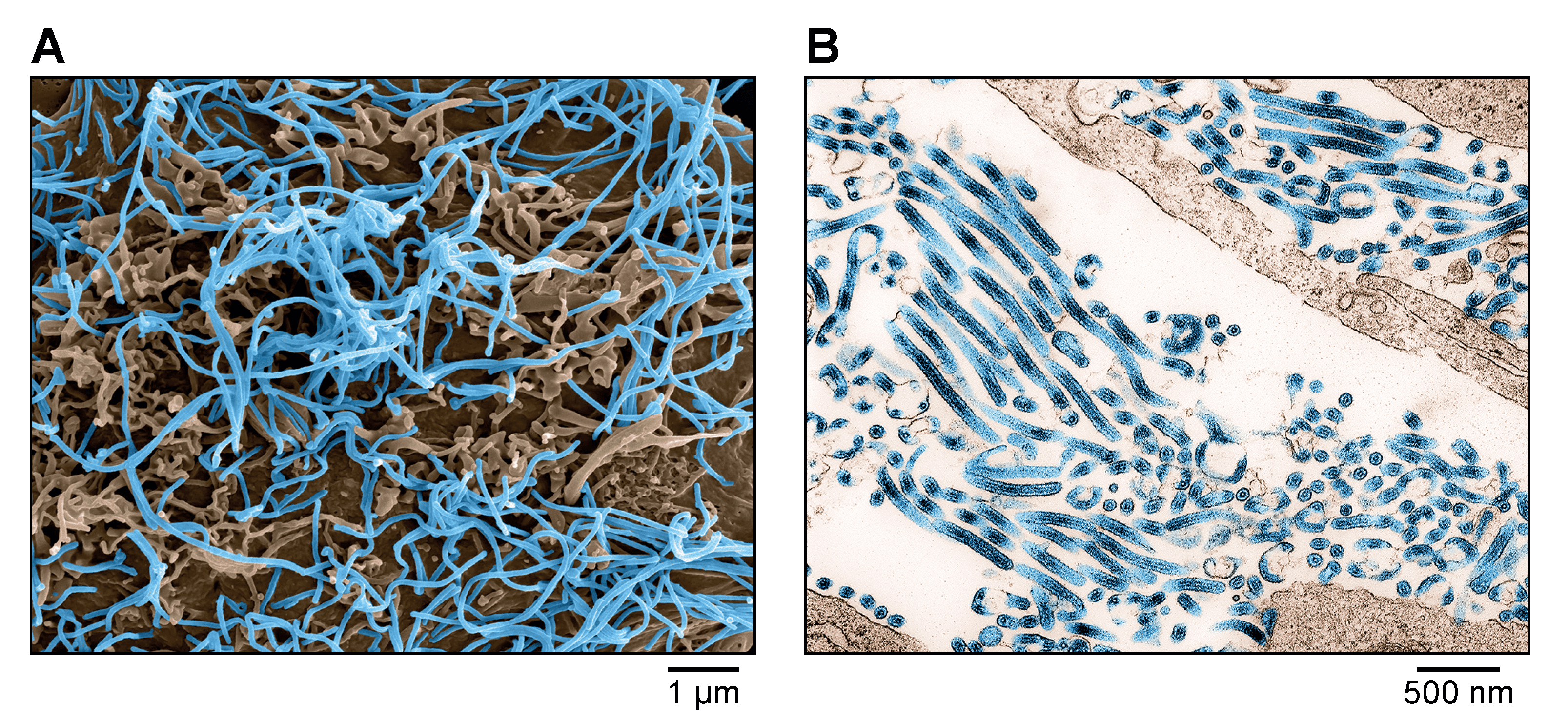 |
| Figure 1. Ebolavirus. A) Scanning electron micrograph of Ebola virus particles (blue) budding from an infected grivet (Chlorocebus aethiops (Linnaeus, 1758)) Vero E6 cell. B) Transmission electron micrograph of Ebola virus particles (blue) found both as extracellular particles and budding particles from infected Vero E6 cells. Images are colorized for clarity. Courtesy of John G. Bernbaum and Jiro Wada, NIH/NIAID/DCR/IRF-Frederick, Fort Detrick, MD, USA. |
Physicochemical and physical properties
EBOV particles have a molecular mass of about 3.82×105 kDa. The buoyant density of EBOV particles in potassium tartrate is about 1.14 g/ml. The sedimentation coefficient (S20,w) of uniform filamentous particles is 1,300–1,400 S, but higher for longer particles. The nucleocapsid has a buoyant density in cesium chloride (CsCl) of about 1.32 g cm−3 (Kiley et al., 1980).
Nucleic acid
Orthoebolavirus genomes are linear non-segmented RNA molecules of negative polarity. Genomic RNAs are about 18.9 kb and are not polyadenylated at their 3′-ends, and there is no evidence for 5′-end terminal cap structures or covalently linked proteins. The Mr of genomic RNA is about 4.2×106 and represents about 1.1% of the total virion mass (Kiley et al., 1980, Regnery et al., 1980, Elliott et al., 1985, Boehmann et al., 2005).
Proteins
Orthoebolaviruses express seven structural proteins, all of which are homologous to those of cuevaviruses, dianloviruses, and orthomarburgviruses (Table 1.Orthoebolavirus). After VP40, the second most abundant structural protein in virions is the nucleoprotein (NP), which encapsidates the orthoebolavirus genome. The least abundant protein is the large protein (L), which mediates orthoebolavirus genome replication and transcription (Kiley et al., 1980, Sanchez et al., 1993). The orthoebolavirus RNP complex consists of NP, RNP complex associated protein (VP24), polymerase cofactor (VP35), transcriptional activator (VP30), and L. The RNP complexes associate with the matrix protein (VP40), which line the inner side of the virion membrane, and the glycoprotein (GP1 and GP2), which forms globular spikes on the outside of the virion membrane (Mühlberger et al., 1999, Beniac et al., 2012, Banadyga et al., 2017, Kirchdoerfer et al., 2017, Sugita et al., 2018). Similar to cuevaviruses, orthoebolaviruses express their structural glycoprotein (GP1 and GP2) via co transcriptional editing and also express three soluble glycoproteins from the GP gene (sGP, ssGP, and Δ-peptide) (Volchkov et al., 1995, Sanchez et al., 1996, Volchkova et al., 1999, Mehedi et al., 2011).
Table 1.Orthoebolavirus. Location and functions of orthoebolavirus structural proteins.
| Protein | Encoding gene | Characteristics | Function | Reference |
|---|---|---|---|---|
| Nucleoprotein (NP) | 1 (NP) | RNP complex component; second most abundant protein in infected cells and in virions; consists of two distinct functional modules; homo oligomerizes to form helical polymers; binds to genomic and antigenomic RNA, VP35, VP40, VP30, and VP24; phosphorylated, O glycosylated, and possibly sialylated, interacts with PP1 and PP2A | Nucleocapsid and cellular inclusion body formation; encapsidation of orthoebolavirus genome and antigenome; genome replication and transcription.Activates unfolded protein response (UPR) | (Huang et al., 2002, Ebihara et al., 2006, Watanabe et al., 2006, Groseth et al., 2009, Hoenen et al., 2012, Peyrol et al., 2013, Kirchdoerfer et al., 2015, Su et al., 2018, Sugita et al., 2018,Mühlberger et al., 1999, Kruse et al., 2018, Rohde et al., 2022, Ahmad et al., 2023) |
| Polymerase cofactor (VP35) | 2 (VP35) | RNP complex component; homo oligomer; phosphorylated; ubiquitinylated; binds to double stranded RNA, NP, VP30, and L | Replicase-transcriptase cofactor; inhibits innate immune response by interfering with IRF3, IRF7, IFIH1, DDX58, and RNAi pathways; inhibits stress granule formation | (Mühlberger et al., 1999, Basler et al., 2000, Basler et al., 2003, Reid et al., 2005, Cárdenas et al., 2006, Haasnoot et al., 2007, Kimberlin et al., 2010, Leung et al., 2010, Biedenkopf et al., 2013, Kirchdoerfer et al., 2015, Nelson et al., 2016, Ivanov et al., 2020, van Tol et al., 2022) |
| Matrix protein (VP40) | 3 (VP40) | Most abundant protein in infected cells and in virions; consists of two distinct functional modules; homo oligomerizes to form dimers and circular hexamers and octamers; SUMOylated; binds single stranded RNA, tubulin alpha, VP35; hydrophobic; membrane associated; contains three late budding motifs; binds to NEDD4 and TSG101; binds to tubulin alpha and is ubiquitinylated | Matrix component; regulation of genome transcription and replication; regulation of virion morphogenesis and egress and virion entry/disassembly in late endosomes | (Dessen et al., 2000, Harty et al., 2000, Noda et al., 2002, Licata et al., 2003, Panchal et al., 2003, Timmins et al., 2003, Hoenen et al., 2010, Bornholdt et al., 2013, Clifton et al., 2015, Baz-Martínez et al., 2016, Pavadai et al., 2018, Winter et al., 2023) |
| Secreted glycoprotein (sGP) | 4 (GP) | Mostly nonstructural; secreted as a parallel homo dimer in high amounts from infected cells; N glycosylated, C mannosylated | Unknown. Hypothesized to be an antibody-decoy and an anti inflammatory agent | (Volchkova et al., 1998, Barrientos et al., 2004, Falzarano et al., 2006, Falzarano et al., 2007, de La Vega et al., 2015) |
| Glycoprotein subunits (GP1 and GP2) | 4 (GP) | Type I transmembrane and class I fusion protein; cleaved to GP1 and GP2 subunits that heterodimerize; mature protein is a trimer of GP1 and GP2 heterodimers; inserts into membranes; heavily N- and O glycosylated, sialylated, acylated, phosphorylated. ADAM17 converts GP1 and GP2 into a soluble form (GP1,2Δ) | Virion adsorption to orthoebolavirus susceptible cells via cellular attachment factors; determines orthoebolavirus cell and tissue tropism; induction of virus cell membrane fusion subsequent to endolysosomal binding to NPC1; inhibits innate immune response by interfering with BST2. GP1,2Δ triggers immune activation and increased vascular permeability | (Ito et al., 2001, Chandran et al., 2005, Schornberg et al., 2006, Lee et al., 2008, Ritchie et al., 2010, Carette et al., 2011, Côté et al., 2011, Collar et al., 2017, Takada et al., 1997, Volchkov et al., 1998a, Volchkov et al., 1998b, Malashkevich et al., 1999, Jeffers et al., 2002, Dolnik et al., 2004, Jouvenet et al., 2009, Kaletsky et al., 2009, Escudero-Pérez et al., 2014, Tran et al., 2014, Wang et al., 2016, Beniac and Booth 2017) |
| Secondary secreted glycoprotein (ssGP) | 4 (GP) | Predominantly nonstructural; secreted as a N glycosylated monomer | Unknown | (Volchkova et al., 1998, Mehedi et al., 2011) |
| Δ-peptide | 4 (GP) | Nonstructural; secreted; largely unstructured; O glycosylated and sialylated | Hypothesized to suppress filovirid superinfection and/or act as a viroporin; might act as an enterotoxin | (Volchkova et al., 1999, Radoshitzky et al., 2011, He et al., 2017, Pokhrel et al., 2019, Melnik et al., 2022) |
| Transcriptional activator (VP30) | 5 (VP30) | RNP complex component; hexameric zinc finger protein; binds single-stranded RNA, NP, VP35 and L; phosphorylated; interacts with SRPK1 and SRPK2 | Transcription initiation at the NP gene start site; potentially reinitiation and antitermination; regulation of genome transcription and replication | (Mühlberger et al., 1999, Modrof et al., 2002, Weik et al., 2002, Modrof et al., 2003, John et al., 2007, Groseth et al., 2009, Biedenkopf et al., 2013, Biedenkopf et al., 2016a, Biedenkopf et al., 2016b, Xu et al., 2017, Takamatsu et al., 2020) |
| RNP complex-associated protein (VP24) | 6 (VP24) | RNP complex component; homo tetramerizes; hydrophobic and membrane associated | Regulation of genome transcription and replication; regulation of virion morphogenesis and egress; inhibits phosphorylation of MAPK and prevents karyopherin shuttling from cytoplasm into the nucleus; inhibits host cell signaling downstream of IFNA1/IFNB1/IFNG; with the exception of BOMV, inhibits both RIGI- and IFIH1-stimulated IFNB and IFNL1 promoter activation, and IFNA-induced MX1 and IFITM3 promoter activation | (Han et al., 2003, Hoenen et al., 2006, Reid et al., 2007, Halfmann et al., 2011, Mateo et al., 2011, Schwarz et al., 2017, Ebihara et al., 2006, Watanabe et al., 2007, Zhang et al., 2012, Xu et al., 2014, Banadyga et al., 2017, Wan et al., 2017, He et al., 2021, Khan et al., 2023) |
| Large protein (L) | 7 (L) | RNP complex component; homo dimerizes; binds to genomic and antigenomic RNA, VP35, and VP30; mRNA capping enzyme; methyltransferase | Genome replication and mRNA transcription; co transcriptional editing | (Volchkov et al., 1995, Sanchez et al., 1996, Mühlberger et al., 1999, Ferron et al., 2002, Groseth et al., 2009, Shabman et al., 2013, Trunschke et al., 2013, Tchesnokov et al., 2018, Valle et al., 2021, Yuan et al., 2022, Peng et al., 2023) |
ADAM17, ADAM metallopeptidase domain 17; DDX58, DExD/H-box helicase 58; IFIH1, interferon induced with helicase C domain 1 (formerly MDA5); IFNA, interferon alpha; IFNB, interferon beta; IFNG, interferon gamma; IFNL, interferon lambda; IRF, interferon regulatory factor; MAPK, mitogen-activated protein kinase; MX1, MX dynamin like GTPAse 1 (formerly MxA); NPC1, NPC intracellular cholesterol transporter 1; NEDD4, E3 ubiquitin protein ligase; PP2A, protein phosphatase 2; RNP, ribonucleoprotein; SRPK, SRSF protein kinase; SUMO, small ubiquitin-like modifier; TSG101, tumor susceptibility 101; VP, virus protein
Lipids
The viral envelope is derived from host cell membranes and is considered to have a lipid composition similar to that of the host cell plasma membrane (Bavari et al., 2002, Feizpour et al., 2015). Orthoebolavirus glycoproteins are acylated (Ito et al., 2001).
Carbohydrates
The glycoproteins of orthoebolaviruses are highly glycosylated with N-linked high mannose, hybrid, and bi-, tri-, and tetra antennary complex glycans with and without fucose and sialic acid and with O linked glycans of the neutral mucin type. Glycans constitute >50% of the GP1 and GP2 total mass. In contrast to orthomarburgvirus GP1 and GP2, orthoebolavirus GP1 and GP2 may be strongly sialylated (Ritchie et al., 2010, Collar et al., 2017). Orthoebolavirus NP is O-glycosylated and possibly sialylated (Huang et al., 2002, Watanabe et al., 2006). sGP is N-glycosylated and C-mannosylated (Falzarano et al., 2006, Falzarano et al., 2007). ssGP is N-glycosylated (Mehedi et al., 2011). Δ-peptide is O glycosylated and sialylated (Volchkova et al., 1999).
Genome organization and replication
The orthoebolavirus genome has the gene order 3′-NP-VP35-VP40-GP-VP30-VP24-L-5′ (Figure 2.Orthoebolavirus). The extragenic sequences at the extreme 3′-end (leader) and 5′-end (trailer) of the genome are conserved and partially complementary. Genes are flanked by conserved transcriptional initiation and termination (polyadenylation) sites. Most genes are separated by non conserved intergenic sequences, but some genes overlap. Most of these overlaps are extremely short and limited to a highly conserved pentamer. In addition, most genes possess relatively long 3′- and 5′-noncoding regions (Sanchez et al., 1993, Ikegami et al., 2001, Sanchez and Rollin 2005, Towner et al., 2008). Similar to cuevaviruses, the GP genes of orthoebolaviruses possess three overlapping open reading frames (ORFs) that can be joined through co-transcriptional polymerase stuttering (Volchkov et al., 1995, Sanchez et al., 1996).
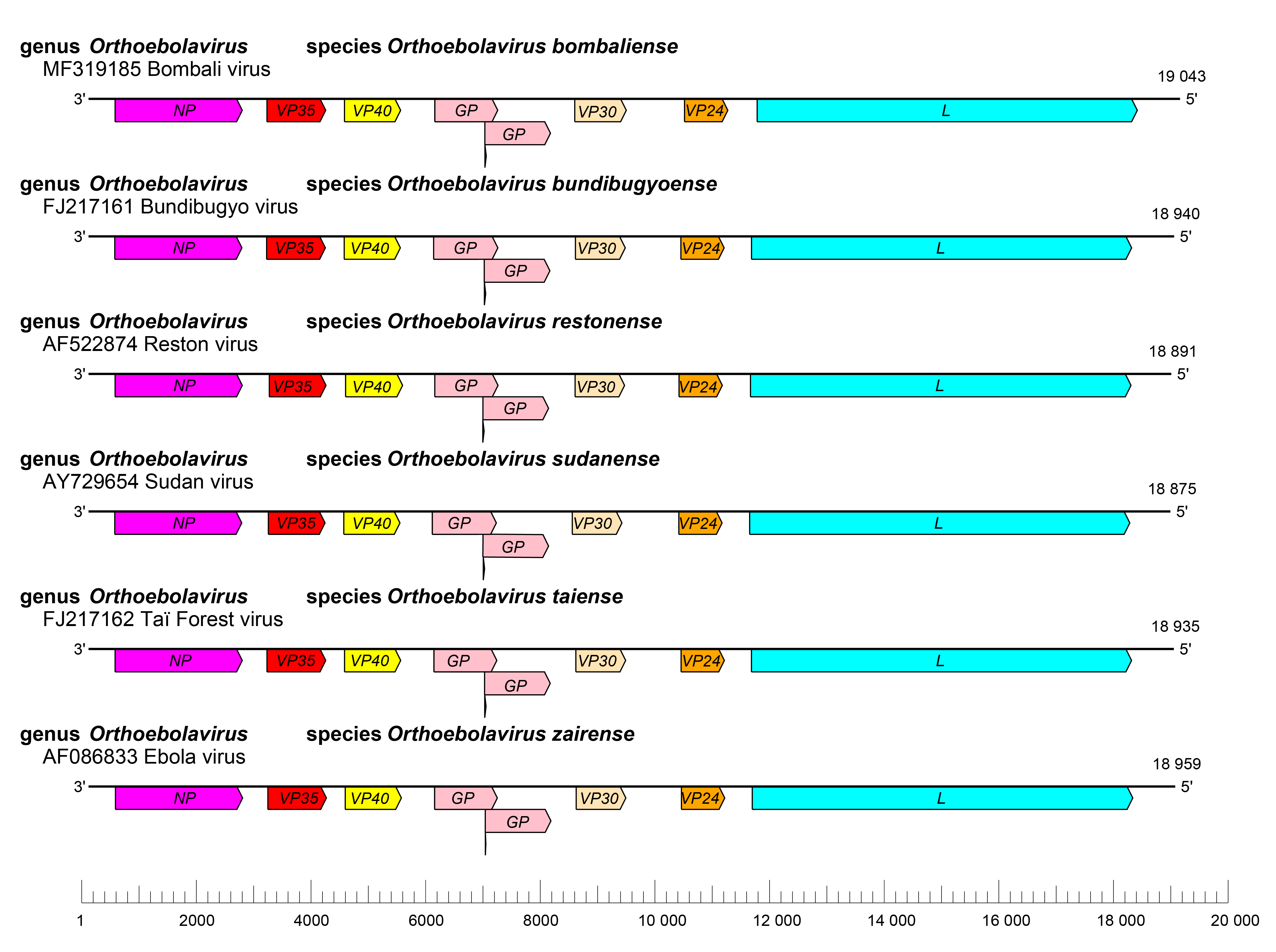 |
| Figure 2. Ebolavirus. Schematic representation of orthoebolavirus genome organization. Genomes are drawn to scale. |
The replication strategy of orthoebolaviruses is highly similar to that of cuevaviruses and reminiscent of that of dianloviruses and orthomarburgviruses (Brauburger et al., 2015, Manhart et al., 2018). Orthoebolavirions enter host cells mainly by macropinocytosis (Nanbo et al., 2010, Saeed et al., 2010, Aleksandrowicz et al., 2011). Orthoebolavirus GP1 and GP2 mediate cell surface attachment factor (e.g., C type lectins, integrins, hepatitis A virus cellular receptor 1 [HAVCR1]) binding and subsequent low-pH-dependent fusion into endosomes (Davey et al., 2017). Cathepsin L and/or B cleavage is required for GP1 and GP2 binding to the endosomal receptor NPC intracellular cholesterol transporter 1 (NPC1), which is also used by cuevaviruses, dianloviruses, and orthomarburgviruses (Chandran et al., 2005, Schornberg et al., 2006, Carette et al., 2011, Côté et al., 2011, Misasi et al., 2012, Ng et al., 2014). Uncoating is presumed to occur in a manner analogous to that of other mononegaviruses. Orthoebolavirus transcription and genome replication take place in the cytoplasm and, in general, follow the models for other members of the Filoviridae family. Transcription starts at the conserved transcriptional initiation site and is strongly dependent on transcriptional activator VP30. Polyadenylation occurs at a stretch of uridine residues within the transcriptional termination site. The 5′-terminal non-coding sequences likely form hairpin-like structures in all (capped and polyadenylated) mRNAs (Bach et al., 2021a, Bach et al., 2021b). Replication involves the synthesis of full-length positive-sense genome copies (antigenomes) with preceding synthesis of short leader RNAs (Sanchez and Kiley 1987, Bray et al., 2000, Weik et al., 2002, Brauburger et al., 2015, Bach et al., 2021c). The polymerase initiates internally at +2 nt and adds terminal nts (A or G) by a backpriming mechanism (Deflubé et al., 2019). During infection, massive amounts of nucleocapsids accumulate intracellularly and form intracytoplasmic inclusion bodies, which are the sites of orthoebolavirus transcription, replication, and nucleocapsid assembly (Hoenen et al., 2012). Mature nucleocapsids are transported based on actin polymerization for envelopment to the plasma membrane, where budding occurs in a VP40-mediated process (Harty et al., 2000, Noda et al., 2002, Licata et al., 2003, Bornholdt et al., 2013, Schudt et al., 2015, Pavadai et al., 2018, Takamatsu et al., 2018, Grikscheit et al., 2020) (Figure 3.Filoviridae).
Biology
Orthoebolaviruses are endemic in Eastern Africa (BDBV, SUDV), Middle Africa (BDBV, EBOV), and Western Africa (BOMV, EBOV, TAFV) and in Eastern and South-eastern Asia (RESTV). BOMV infects little free-tailed bats (molossid Chaerephon pumilus (Cretzschmar, 1826)) and Angolan free tailed bats (molossid Mops condylurus A. Smith, 1933), but the natural hosts of other orthoebolaviruses are unknown and spread of orthoebolaviruses is not associated with any vector (bats are suspected hosts for all orthoebolaviruses; RESTV also naturally infects domestic pigs (suid Sus scrofa domesticus Erxleben, 1777)). The route of initial human infection is unknown (Amman et al., 2017). The major route of human-to-human transmission of orthoebolaviruses requires direct contact with blood, bodily fluids, or injured skin. (Kuhn et al., 2020). In the laboratory, rodents (laboratory mice, domesticated guinea pigs, golden hamsters), nonhuman primates (common marmosets, crab-eating macaques, grivets, hamadryas baboons, rhesus monkeys), carnivores (domestic ferrets), and suids (domestic pigs) can be infected experimentally with various orthoebolaviruses, but lethal infection of rodents requires sequential adaptation (Marsh et al., 2011, St Claire et al., 2017, Siragam et al., 2018, Kuhn et al., 2020).
Antigenicity
The antigenicity of orthoebolaviruses is primarily due to their glycoproteins. Numerous anti GP1,2 monoclonal antibodies have been described that are specific for a particular orthoebolavirus or cross reactive among orthoebolaviruses. Importantly, the ability of a monoclonal antibody to neutralize orthoebolavirus infection in vitro is not necessarily predictive of protective efficacy in vivo; conversely, antibodies that are non-neutralizing in vitro have shown to be protective in vivo. Orthoebolaviruses and other filovirids have only limited antigenic relatedness, and individual orthoebolaviruses can be differentiated with certain antibodies (Lee et al., 2008, Dias et al., 2011, Wec et al., 2017, Bramble et al., 2018, Flyak et al., 2018, Saphire et al., 2018, West et al., 2018).
Species demarcation criteria
PAirwise Sequence Comparison (PASC) using coding-complete orthoebolavirus genomes is the primary tool for orthoebolavirus species demarcation. Genomic sequences of orthoebolaviruses of different species differ from each other by ≥23% (Bào et al., 2017). Genomic features, such as the number and location of gene overlaps, orthoebolavirus host and geographic distribution, and orthoebolavirus pathogenicity for different organisms are also taken into account for species assignment.
Phylogenetic relationships across the genus have been established from maximum likelihood trees generated using coding-complete or complete genome sequences (Figure 4.Filoviridae) or by phylogenetic analysis of RNA-directed RNA polymerase (RdRP) sequences (Wolf et al., 2018).

