Family: Roniviridae
Peter J. Walker, Jeff A. Cowley, Xuan Dong, Jie Huang, Nick Moody and John Ziebuhr
The citation for this ICTV Report chapter is the summary published as Walker et al., (2021):
ICTV Virus Taxonomy Profile: Roniviridae, Journal of General Virology, 102 (1): 001514
Corresponding author: Peter J. Walker ([email protected])
Edited by: Peter Simmonds
Posted: October 2020, November 2022, May 2023
PDF: ICTV_Roniviridae.pdf (2020 version)
Summary
The family Roniviridae includes two genera (Okavirus and Nimanivirus) for viruses infecting penaeid or palaemonid shrimp (Table 1.Roniviridae). Ronivirus genomes are non-segmented, positive-sense, single-stranded RNA of 26–29 kb and contain 5 canonical long open reading frames. ORF1a encodes a polyprotein (pp1a) containing chymotrypsin-like cysteine proteinase and papain-like proteinase domains. ORF1b overlaps ORF1a where a −1 ribosomal frameshift enables the expression of the pp1ab polyprotein. ORF1b encodes enzymes involved in RNA synthesis and modification. ORF2 encodes the virion nucleoprotein (p20). ORF3 encodes a precursor polyprotein (pp3) from which the gp116 and gp64 virion envelope glycoproteins are derived. Natural infections are common without apparent clinical signs. However, yellow head virus is a very virulent pathogen that causes mass mortalities in farmed shrimp. Some other members of the family have also been associated with high mortalities in larval or adult shrimp.
Table 1.Roniviridae. Characteristics of members of the family Roniviridae.
| Characteristic | Description |
| Typical member | gill-associated virus (AF227196), species Okavirus branchiae, genus Okavirus |
| Virion | Enveloped, bacilliform (rod-shaped) particles 150–200 nm in length and 40–60 nm in diameter with a helical nucleocapsid composed of the nucleocapsid protein (p20) surrounded by a lipid envelope containing two transmembrane glycoproteins (gp64 and gp116) |
| Genome | Non-segmented, positive-sense, single-stranded RNA of 26–29 kb containing five or six long open reading frames |
| Replication | Cytoplasmic; nucleocapsids bud at membranes of the endoplasmic reticulum/Golgi complex to form mature virions |
| Translation | From a nested set of 5′-capped and 3′-co-terminal polyadenylated mRNAs |
| Host range | Penaeid shrimp are natural hosts; experimental infection reported in penaeid and palaemonid shrimp of various species |
| Taxonomy | Realm Riboviria, kingdom Orthornavirae, phylum Pisuviricota, class Pisoniviricetes, order Nidovirales; suborder Ronidovirineae: one sub-family (Okanivirinae) including two genera (Okavirus and Nimanivirus) with the subgenus Tipravirus (3 species) in the genus Okavirus and the subgenus Marovirus (one species) in the genus Nimanivirus |
Virion
Morphology
Virions are enveloped, rod-shaped (bacilliform) particles with rounded ends, 40–60 nm in diameter and 150–200 nm in length (Figure 1.Roniviridae) (Boonyaratpalin et al., 1993, Chantanachookin et al., 1993, Spann et al., 1995, Spann et al., 1997, Wang and Chang 2000). The envelope is studded with prominent peplomers projecting approximately 11 nm from the surface. Nucleocapsids are 20–30 nm in diameter and have a helical symmetry with coil periodicity of 5–7 nm (Chantanachookin et al., 1993, Spann et al., 1995). Long, filamentous nucleocapsid precursors (approximately 15 nm in diameter and 80–450 nm in length) occur in the cytoplasm of infected cells and acquire envelopes by budding at membranes of the endoplasmic reticulum or other cellular membranes (Boonyaratpalin et al., 1993, Chantanachookin et al., 1993). Newly budded mature enveloped virions often appear as multimers arranged end-to-end, and paracrystalline arrays of virions can occur within vesicles.
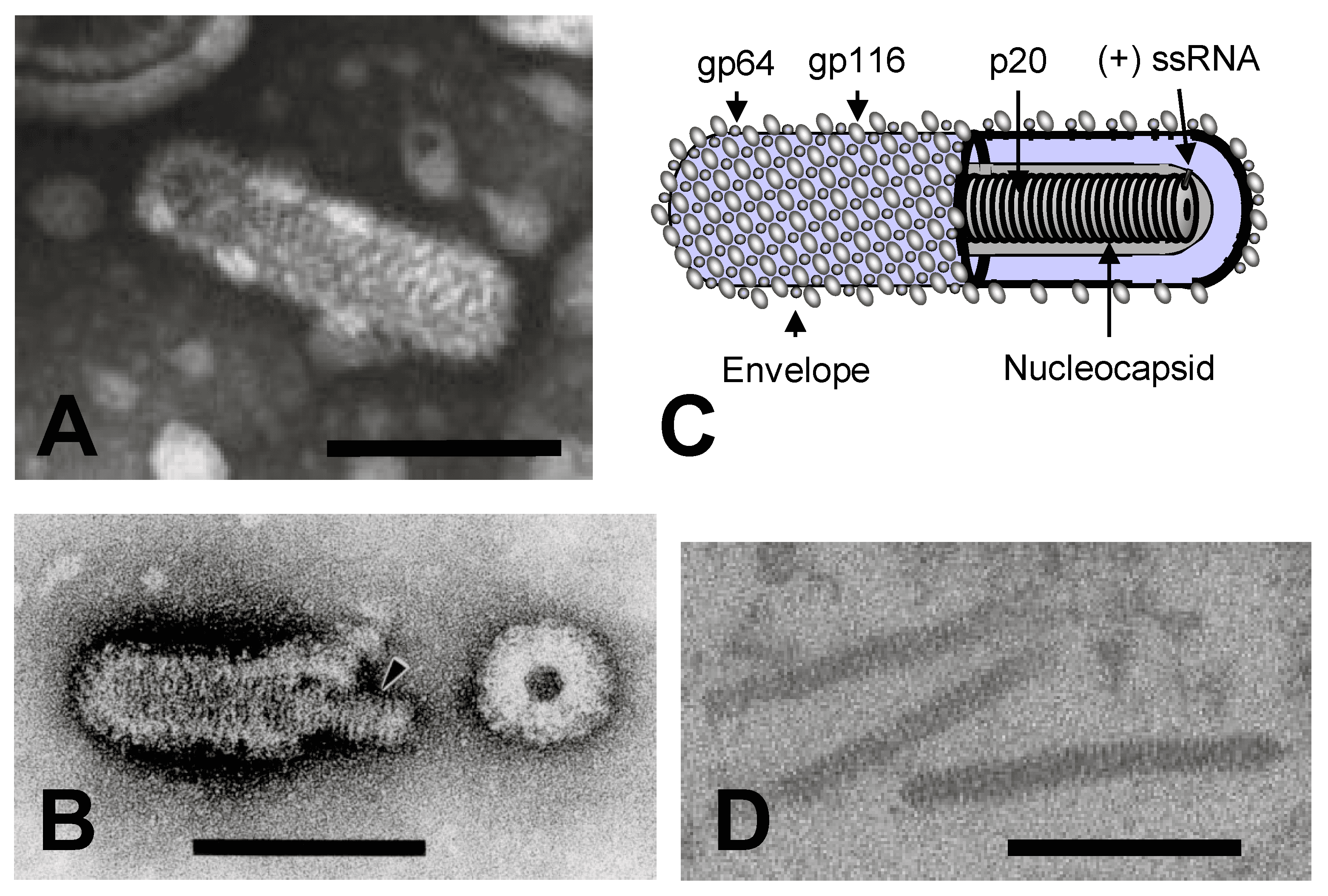 |
| Figure 1.Roniviridae. (A) Transmission electron micrograph of a negative-stained virion of gill-associated virus. (B) Transmission electron micrograph of a partially disrupted yellow head virus virion displaying the internal nucleocapsid and a ring-like structure which appears to be a disrupted virion in cross-section. (C) Schematic illustration of a ronivirus virion. (D) Transmission electron micrograph of unenveloped cytoplasmic nucleocapsids in a thin section of gill-associated virus-infected shrimp cells. The bars represent 100 nm. (Courtesy of K.M. Spann, P. Loh, J.A. Cowley and R.J. McCulloch, Panels A, B and C reproduced from (Dhar et al., 2004) with permission) |
Physicochemical and physical properties
Yellow head virus (YHV, species Okavirus flavicapitis) virions have a buoyant density in sucrose of 1.18–1.20 g ml-1 (Nadala et al., 1997). YHV is inactivated by heating at 60°C for 15–30 min but has been reported to survive at 25–28°C in seawater for at least three days (Flegel et al., 1995). Virions are sensitive to calcium hypochlorite and sodium dodecyl sulphate, but the sensitivity to other treatments is not known (Flegel et al., 1997).
Nucleic acid
Virions contain a single linear molecule of positive-sense (+) single-stranded (ss) RNA (26–29 kb) with a 5′-terminal cap (7-methylguanosine triphosphate) and a 3′-polyadenylated tail (Wongteerasupaya et al., 1995, Cowley and Walker 2002, Sittidilokratna et al., 2008, Hooper et al., 2020, Meng et al., 2021).
Proteins
Virions comprise a nucleoprotein of 20–22 kDa (p20) which complexes with the RNA genome to form the nucleocapsid, and two envelope glycoproteins of 110–135 kDa (gp116) and 63–67 kDa (gp64) which form the prominent peplomers on the virion surface (Jitrapakdee et al., 2003, Soowannayan et al., 2010). The mature gp116 and gp64 glycoproteins originate by post-translational cleavage of a precursor polyprotein (pp3) encoded in ORF3 (Cowley and Walker 2002, Jitrapakdee et al., 2003). In YHV, cleavage occurs at the C-terminal side of the third and fifth of six hydrophobic transmembrane (TM) domains, where the sequences have characteristics of signal peptidase type 1 cleavage sites. Glycoproteins gp116 and gp64 are anchored in the virion envelope by either two (TM4 and TM5) or one (TM6) transmembrane domains, respectively. There is no evidence that gp116 and gp64 are linked by intermolecular disulphide bonds (Jitrapakdee et al., 2003).
Lipids
The virion lipid envelope is derived from endoplasmic membranes of host cells during budding (Boonyaratpalin et al., 1993, Chantanachookin et al., 1993).
Carbohydrates
N-linked glycosylation sites are present in the amino acid sequences of gp116 and gp64. In YHV, all but one of the seven sites in gp116 are glycosylated and three of four sites in gp64 are glycosylated (Jitrapakdee et al., 2003). Carbohydrates attached to gp64 and gp116 comprise primarily mannose-type glycans; those on gp116 also comprise N-acetyl-galactosamine and N-acetyl-glucosamine in lesser abundance (Soowannayan et al., 2010). Few if any of the multiple O-linked glycosylation sites in each glycoprotein appear to be utilized.
Genome organisation and replication
The positive-sense (+) ssRNA genome contains five or six long ORFs (Figure 2.Roniviridae). In order from the 5′-terminus, they include: ORF1a encoding papain-like and 3C-like proteases; overlapping ORF1b encoding nidovirus RdRP-associated nucleotidyltransferase (NiRAN), RNA-directed RNA polymerase (RdRP), helicase-associated zinc-binding domain, superfamily 1 helicase, exoribonuclease, guanosine N7-methyltransferase, and ribose 2′-O-methyltransferase domains; ORF2 encoding the nucleoprotein (p20); and ORF3 encoding the precursor polyprotein (pp3) from which the envelope glycoproteins gp116 and gp64 are derived (Cowley and Walker 2002, Ziebuhr et al., 2003, Sittidilokratna et al., 2008, Lehmann et al., 2015, Zeng et al., 2016, Dong et al., 2017, Hooper et al., 2020, Meng et al., 2021). In okaviruses, cleavage of pp3 also generates a polypeptide (p25) at the N-terminus containing three predicted TM domains and N-linked glycosylation sites in the ectodomain. Although p25 is similar in size to the M proteins of some vertebrate nidoviruses, there is no clear evidence of its association with virions. In Macrobrachium rosenbergii Golda virus (MrGV; species Nimanivirus lahi) ORF3 encodes a polyprotein with a predicted N-terminal signal peptide and only three predicted TM domains, and encodes homologues of gp116 and gp64; the N-terminal region corresponding to the okavirus triple-membrane-spanning p25 protein is absent. An alternative reading frame (ORFX) commences 3 nucleotides downstream of the pp3 initiation codon in all okaviruses but is absent from MrGV. ORFX encodes a small double-membrane-spanning protein (px) of 86 amino acids (Firth and Atkins 2009) which has the structural characteristics of a class IIb viroporin (Nieva et al., 2012). In gill-associated virus (GAV, species Okavirus branchiae), ORF3 is followed by a sixth open reading frame (ORF4) encoding a small putative protein of 98 amino acids (Cowley and Walker 2002). However, ORF4 is severely truncated in other roniviruses and evidence for its expression in GAV is poor (Sittidilokratna et al., 2008, Wijegoonawardane et al., 2008, Dong et al., 2017).
Ronivirus genome organisation differs from those of vertebrate nidoviruses in several aspects: i) the nucleoprotein gene (ORF2) is located upstream of the glycoprotein gene (ORF3); ii) there is no discrete gene encoding a structural membrane (M) protein; and iii) the virion envelope glycoproteins gp116 and gp64 are generated by proteolysis of a pp3 precursor polyprotein encoded by the ORF3 gene (Cowley and Walker 2002, Cowley et al., 2004, Sittidilokratna et al., 2008, Dong et al., 2017). These characteristics more closely resemble those of invertebrate nidoviruses infecting insects (genus Alphamesonivirus, family Mesoniviridae) (Nga et al., 2011, Lauber et al., 2012, Vasilakis et al., 2014).
As in most other nidoviruses, the overlap between ronivirus ORF1a and ORF1b contains a −1 ribosomal frameshift site allowing translation of the pp1ab polyprotein encoded by ORF1a and ORF1b; expression of ORF1b occurs with an efficiency of 23–25% (Cowley et al., 2000). In roniviruses, however, both the ribosomal slippage site and downstream RNA secondary structural element that stimulates slippage differs from those used in either vertebrate nidoviruses or mesoniviruses. In roniviruses, the slippage site (5′-AAAUUUU-3′ or 5′-GGGUUUU-3′) is followed by a large, complex pseudoknot structure (Cowley et al., 2000, Sittidilokratna et al., 2002, Hooper et al., 2020, Meng et al., 2021) whilst in vertebrate nidoviruses the slippage site (5′-[G/U]UUAAAC-3′) is followed by a more compact hairpin (H)-type pseudoknot. In mesoniviruses, the slippage site (5′-GGAUUUU-3′) is followed by a stem-loop structure rather than a pseudoknot (Brierley 1995, Nga et al., 2011). Interestingly, in the Euroniviridae (a closely related family of crustacean nidoviruses), viruses assigned to the genus Paguronivirus appear to employ the same slippage site as roniviruses (5′-AAAUUUU-3′) and viruses assigned to the genus Charybnivirus appear to employ a similar site (5′-GCUUUUU-3′) (Shi et al., 2016).
In roniviruses, the ORF1a coding sequence contains three or four hydrophobic domains (HD1–HD4) and a chymotrypsin-like cysteine (3C-like) proteinase (3CLpro) or “main” protease (Mpro) which is flanked by HD3 and HD4 and involved in autolytic processing of the pp1a and pp1ab polyprotein (Cowley et al., 2000, Sittidilokratna et al., 2002, Hooper et al., 2020, Meng et al., 2021). The 3CLpro of GAV has a predicted consensus cleavage site specificity VxHE↓(L/V) that distinguishes it from the corresponding Mpro enzymes in vertebrate nidoviruses. Taken together with sequence differences, the ronivirus 3CLpro appears to combine the Cys-His catalytic dyad of the coronavirus Mpro with a potyvirus-like substrate binding pocket (Ziebuhr et al., 2003). ORF1a also contains two putative papain-like proteinase domains (PLP1 and PLPx) that share remote structural similarity, but very little direct sequence identity, with the PLP domains of coronaviruses (Sittidilokratna et al., 2008). Like the corresponding papain-like proteases (PLpro) of toroviruses, both PLP1 and PLPx lack a Zn2+ binding finger. In vertebrate nidoviruses, the 3CLpro and PLpros process the pp1a/ pp1ab polyproteins into functional nonstructural proteins numbered from the N-terminus as nsp1 to nsp12 in arteriviruses and nsp1 to nsp16 in coronaviruses (Gorbalenya et al., 2006). Whilst the 3CLpro of GAV has been shown to cleave at an immediate upstream site, and at a site toward the C-terminus of pp1ab, the number and composition of nsps generated from pp1ab by the 3CLpro in combination with the putative PLpros is not yet known (Ziebuhr et al., 2003). ORF1b also contains multiple sequence motifs with homologues in other nidoviruses. These include nidovirus RdRP-associated nucleotidyltransferase (NiRAN); RNA-directed RNA polymerase (RdRP) with an “SDD” active site motif; cysteine- and histidine-rich zinc binding domain (ZBD); superfamily 1 helicase (Hel1); exoribonuclease (ExoN); N7-methyltransferase (N-MT); and 2′-O-methyl transferase (O-MT) (Cowley et al., 2000, Sittidilokratna et al., 2002, Zeng et al., 2016). Like other invertebrate nidoviruses, roniviruses lack the nidoviral endoribonuclease specific for uridylate (NendoU) domain that immediately precedes the O-MT domain (Nga et al., 2011, Saberi et al., 2018).
In ronivirus-infected cells, two subgenomic (sg) mRNAs are transcribed, sg mRNA2 and sg mRNA3, from which ORF2 and ORF3 are translated, respectively (Cowley et al., 2002a, Sittidilokratna et al., 2008). The genome length RNA (g RNA1) and the two sg mRNAs are 3′-coterminal and each possesses a 5′-cap (7-methylguanosine triphosphate) structure and a 3′-poly(A) tail (Figure 2.Roniviridae). It is likely that ronivirus sg mRNAs are transcribed from complementary sg negative-sense strand templates as is the case in other nidoviruses. In support of this, double stranded (ds) RNAs equivalent in size to the genomic and two sg mRNAs have been detected in shrimp cells infected with GAV, and likely represent replicative-intermediate RNAs (Cowley et al., 2000). However, unlike the sg mRNAs of coronaviruses and arteriviruses, the ronivirus sg mRNAs do not possess a common leader sequence identical to the genome 5′-terminal sequence. In contrast, g RNA1, sg mRNA2 and sg mRNA3 each initiate with a 5′-AC dinucleotide (Cowley et al., 2002a). For the sg mRNAs, this 5′-terminal AC-dinucleotide maps precisely to a central position of a sequence (5′-GGUCAAUUACAACCUA-3′) that is conserved in the intergenic regions (IGRs) preceding ORF2 and ORF3. Presumably, this conserved element serves a dual function; in the genome as a terminator of negative-sense strand RNA synthesis and in the resulting negative-sense strand sg RNA templates, as a transcriptional promoter for sg mRNA synthesis. Thus, roniviruses appear to differ from coronaviruses and arteriviruses by not using a discontinuous transcription process to produce their mRNAs, but rather a “continuous” transcription strategy similar to that utilized by members of the genus Torovirus (family Tobaniviridae) (van Vliet et al., 2002, Pasternak et al., 2006, Cowley and Walker 2008).
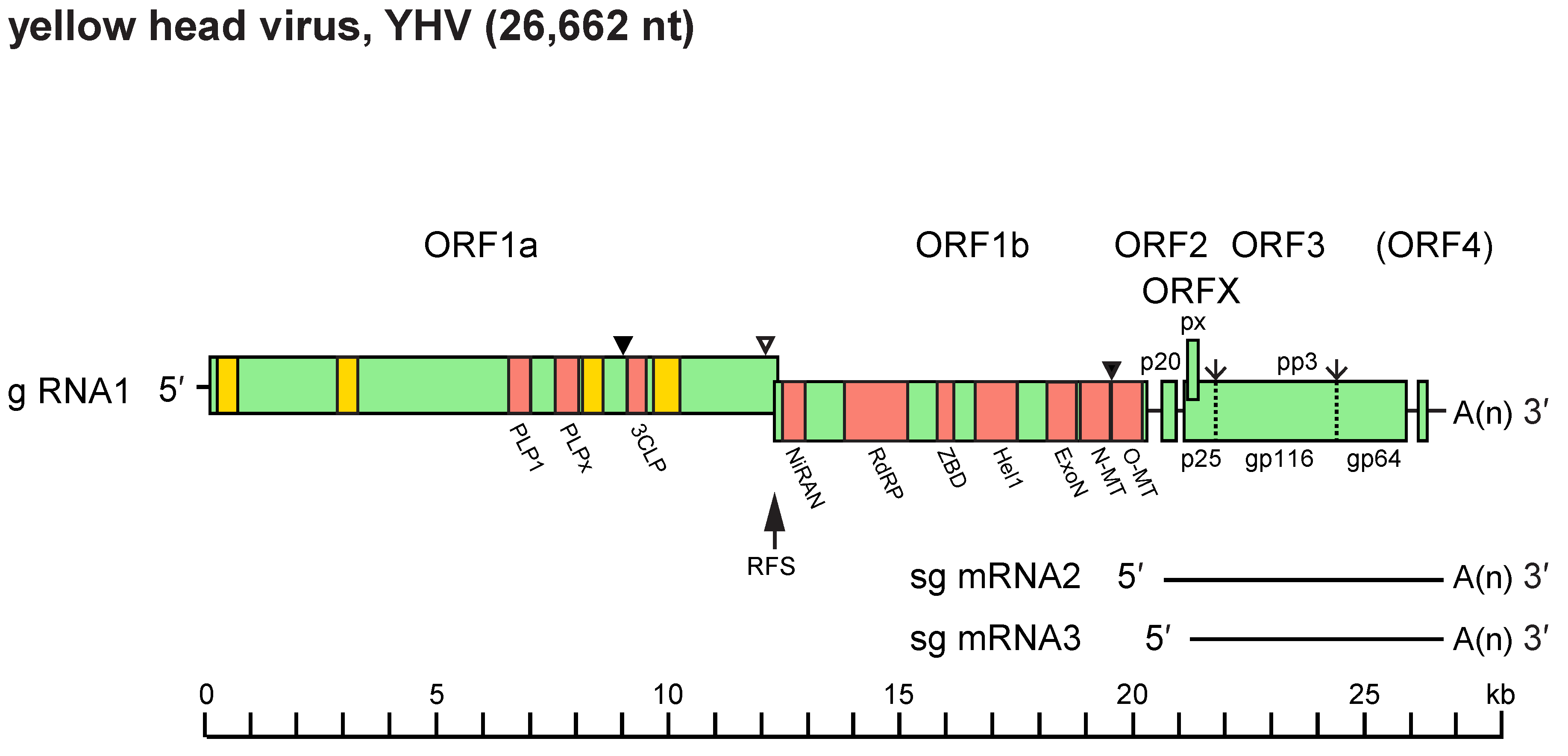 |
| Figure 2.Roniviridae. Schematic representation of the 26 662 nt polyadenylated (+) single-stranded RNA genome of yellow head virus (g RNA1) and the two 3′-coterminal sub-genomic RNAs, sg mRNA2 and sg mRNA3. Functional domains in ORF1a: hydrophobic regions (HD1–HD4, yellow boxes), 3C-like protease (3CLP), papain-like protease (PLP1) and a domain with homology to PLP1 but lacking the canonical α + β fold of papain-like proteases (PLPx). Functional domains in ORF1b: nidovirus RdRP-associated nucleotidyltransferase (NiRAN); RNA-directed RNA polymerase (RdRP); cysteine- and histidine-rich zinc binding domain (ZBD); superfamily 1 helicase (Hel1); exoribonuclease (ExoN); N7-methyltransferase (N-MT); and 2′-O-methyl transferase (O-MT). ORF2 encodes the nucleoprotein (p20). ORF3 encodes a precursor polyprotein (pp3) which undergoes post-translational processing to generate envelope glycoproteins (gp116 and gp64) and an N-terminal triple-membrane-spanning fragment of unknown function (p25). ORFX is an alternative reading frame in ORF3 encoding a putative small double-membrane-spanning protein (px). The ribosomal frameshift site (RFS) mediates a programmed (−1) frameshift upstream of the ORF1a stop codon and thus allows translation to be continued into ORF1b to produce polyprotein pp1ab. Known (▼) and likely (∇) sites of proteolytic cleavage of expressed polyproteins pp1a and pp1ab are indicated together with the two signal peptidase type 1 cleavage sites in pp3 (↓). |
Biology
Roniviruses have been detected only in crustaceans (Walker and Mohan 2009, Hooper et al., 2020, Meng et al., 2021). The giant tiger shrimp (Penaeus monodon) appears to be the primary natural host of GAV and YHV; yellow head virus-8 (YHV-8, species Okavirus orientale) has been reported to infect Chinese white shrimp (Penaeus chinensis) (Dong et al., 2017); MrGV infects giant freshwater prawns (Macrobrachium rosenbergii). Other penaeid and palaemonid shrimp species are susceptible to natural and experimental infection with members of the family (Spann et al., 2000, Longyant et al., 2005, Longyant et al., 2006, Liu et al., 2014, Zhu et al., 2016). The geographic range of roniviruses mirrors that of their primary penaeid hosts, extending from eastern Africa across the Indo-Pacific to South Asia, Australia and some islands in the South Pacific, and northwards throughout Southeast to East Asia (Flegel et al., 1995, Wang and Chang 2000, Wijegoonawardane et al., 2008, Mohr et al., 2015, Dong et al., 2017). YHV has also been detected in cultured blue shrimp (Penaeus stylirostris) from Mexico (Castro-Longoria et al., 2008). Infections may be chronic or acute and transmitted horizontally or vertically (Cowley et al., 2002b, Spann et al., 2003). The prevalence of subclinical infection in giant tiger shrimp can be high in some regions, with pathology limited to the lymphoid organ. In acute infections associated with disease and mortalities, the virus invades most tissues of ectodermal and mesodermal origin. YHV is a highly virulent pathogen of cultured shrimp (Flegel et al., 1995). Other roniviruses have also been associated with pathology and mortalities in shrimp (Spann et al., 1997, Oanh et al., 2011, Liu et al., 2014, Hooper et al., 2020, Meng et al., 2021).
Antigenicity
Monoclonal antibodies have been generated against YHV, targeting linear or discontinuous epitopes in the nucleoprotein (p20) or the envelope glycoproteins (gp116 and gp64) (Sithigorngul et al., 2000, Sithigorngul et al., 2002). One monoclonal antibody against a linear epitope in gp116 (V32B) has been shown to cross-react with GAV; two other monoclonal antibodies against linear epitopes in gp64 (Y18) and p20 (Y19) failed to cross-react with GAV (Soowannayan et al., 2003). Monoclonal antibody Y19 and two other monoclonal antibodies against YHV p20 target linear epitopes in the acidic C-terminal domain (I116–E137) (Sittidilokratna et al., 2006). Polyclonal rabbit antibody against purified YHV reacts with linear epitopes in all regions of p20 except the central domain (V37–Q74) (Sittidilokratna et al., 2006).
Derivation of names
Nimanivirus: from nidovirus, remaining derivation obscure
Okanivirinae, Okavirus: refers to the lymphoid or so named “oka” organ, an anatomical structure common to penaeid shrimp, and which is a site of virus replication and pathology in both subclinical and acute infections.
Roniviridae: from rod-shaped nidovirus referring to the virion morphology of viruses in the family.
Tipravirus: from tiger prawn virus
Subfamily and genus demarcation criteria
Genera are defined on the basis of DEmARC analysis of portions of their complete genome sequences. Genus demarcation is on the basis of analyses using DEmARC (DivErsity pArtitioning by hieRarchical Clustering) - a computational framework applied to all nidoviruses (Lauber and Gorbalenya 2012). DEmARC uses profiles of multiple sequence alignments of the 3CLpro, NiRAN, RdRP, ZBD and HEL1 domains, Bayesian and Maximum-likelihood phylogenetic trees, and profiles of clustering cost (CC) function that are produced for weighted hierarchical clustering of pairwise patristic distances (PPD). The species demarcation threshold is set as a range of PUD values (0.029%–0.043 %) which are deduced from PPD values for which the number of clusters (taxa) remained constant and the CC = 0. PUD values represent the % of different residues in compared proteins.
This demarcation derived from DEmARC analysis should also reflect significant differences in ecology and/or genome organisation between members assigned to different genera.
Relationships within the family
Okaviruses have been assigned unofficially to eight genotypes based on phylogenetic analyses of the nucleotide sequence of a region ORF1b (Figure 3.Roniviridae) (Wijegoonawardane et al., 2008, Mohr et al., 2015, Dong et al., 2017). Yellow head virus (YHV; species Okavirus flavicapitis) is assigned to genotype 1 and includes two sub-types (1a and 1b) which cluster separately. The subtypes are also distinguishable by the presence (subtype 1a) or absence (subtype 1b) of a 162 nt indel in ORF3, resulting in a 54 amino acid sequence variation near the N-terminus of gp116 (Sittidilokratna et al., 2009). Gill-associated virus (GAV; species Okavirus branchiae) is assigned to genotype 2 and other viruses (YHV-3 to YHV-8) are assigned to genotypes 3 to 8, respectively. Viruses assigned to genotype 8 are classified to the species Okavirus orientale; viruses assigned to genotypes 3-7 have not yet been classified taxonomically as complete genome sequences are not yet available. The genotypes vary in geographic distribution. Viruses assigned to genotypes 1, 2, 3 and 5 have been detected in shrimp from various locations in Southeast Asia and China (Wijegoonawardane et al., 2008, Chen et al., 2018). Genotype 4 has been detected only in India (Wijegoonawardane et al., 2008), genotype 6 in East Africa and Australia (Wijegoonawardane et al., 2008, Cowley et al., 2019), genotype 7 only in Australia (Mohr et al., 2015) and genotype 8 only in China (Dong et al., 2017). There is evidence of naturally occurring genetic recombination involving genotypes 1, 2, 3 and 5 (Gangnonngiw et al., 2009, Wijegoonawardane et al., 2009).
Nimaniviruses are presently represented by only two very closely related variants of MrGV (Hooper et al., 2020, Meng et al., 2021).
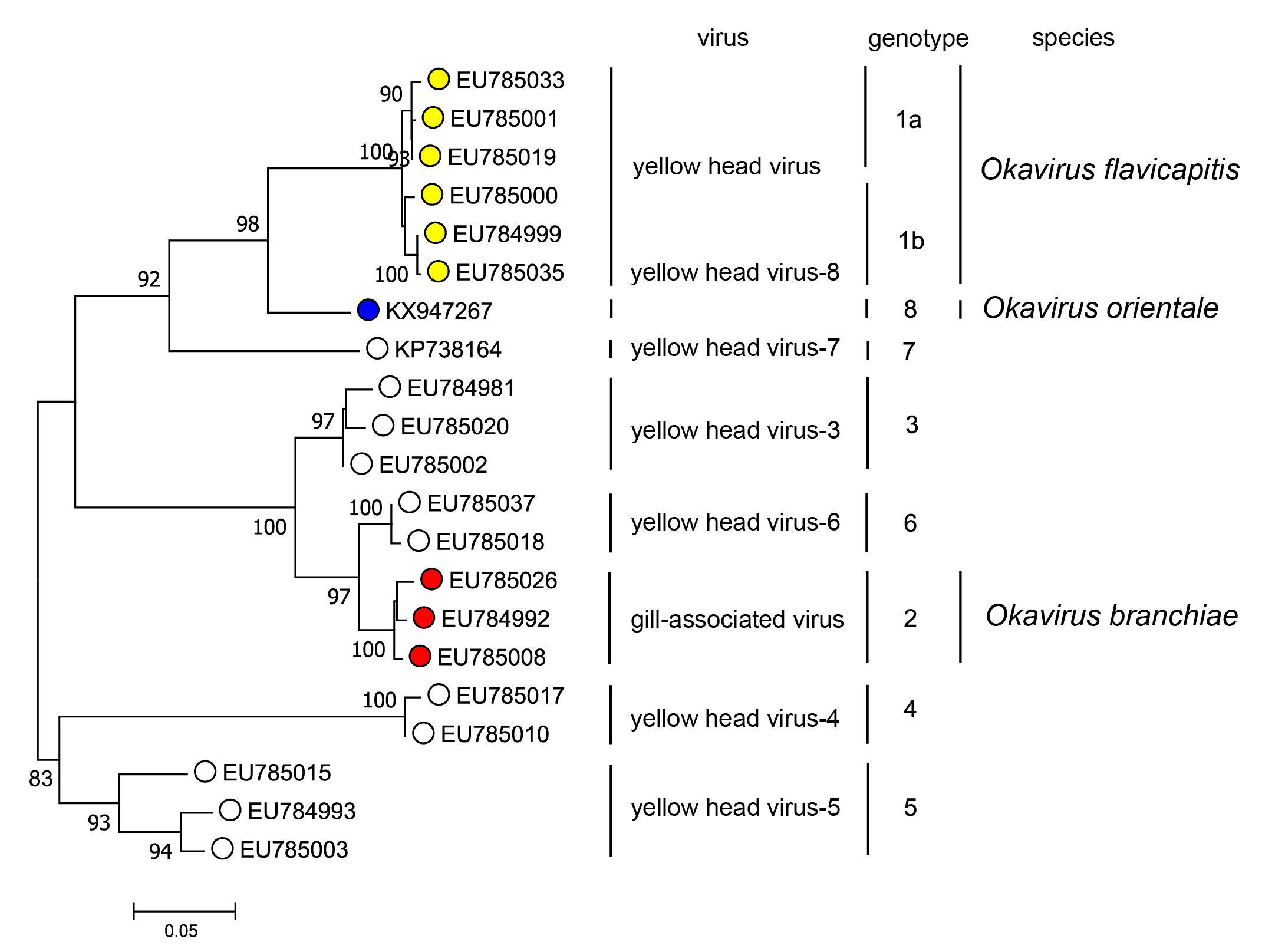 |
| Figure 3.Roniviridae. Maximum Likelihood phylogenetic tree inferred in MEGA 7.0 (Kumar et al., 2016) from a ClustalW nucleotide sequence alignment of a 661 nucleotide region of ORF1b for 21 viruses representing eight identified genotypes in the yellow head virus complex. The aligned sequence extends from within the HEL1 helicase domain (encompassing motifs V, Va and VI) and into the variable region immediately preceding the ExoN exonuclease domain. The evolutionary relationships were inferred by using the Kimura 2-parameter model of nucleotide substitution (Kimura 1980). The initial tree for the heuristic search was obtained automatically by applying Neighbour-Join and BIONJ algorithms to a matrix of pairwise distances estimated using Maximum Composite Likelihood (MCL) method (Lele and Taper 2002), and then selecting the topology with superior log likelihood value. The tree with the highest log likelihood (-3065.67) is shown. Numbers at nodes indicate bootstrap support where these are > 80%. The scale bar indicates 0.05 substitutions per site. This phylogenetic tree and corresponding sequence alignment are available to download from the Resources page. |
Relationships with other taxa
Similarities in the genome size and organization, sequences of core replicase proteins and the sg mRNA transcription strategy used by roniviruses clearly link them to other members of the order Nidovirales (Cowley et al., 2001). Based on conserved functional domains in replicase regions of ORF1b, roniviruses are most closely related phylogenetically to other nidoviruses infecting arthropods including members of the families Mesoniviridae (from mosquitoes) and Euroniviridae (from crustaceans) (Figure 4.Roniviridae). The RNA pseudoknot component of the −1 ribosomal frameshift in the ronivirus ORF1a/1b gene overlap resembles the gag/pol pseudoknots of some retroviruses (Cowley et al., 2000). The structure and substrate specificity of the pp1ab chymotrypsin-like cysteine proteinase (3CLpro) bridges phylogenetic distinctions between the corresponding proteinases of coronaviruses and plant potyviruses (Ziebuhr et al., 2003).
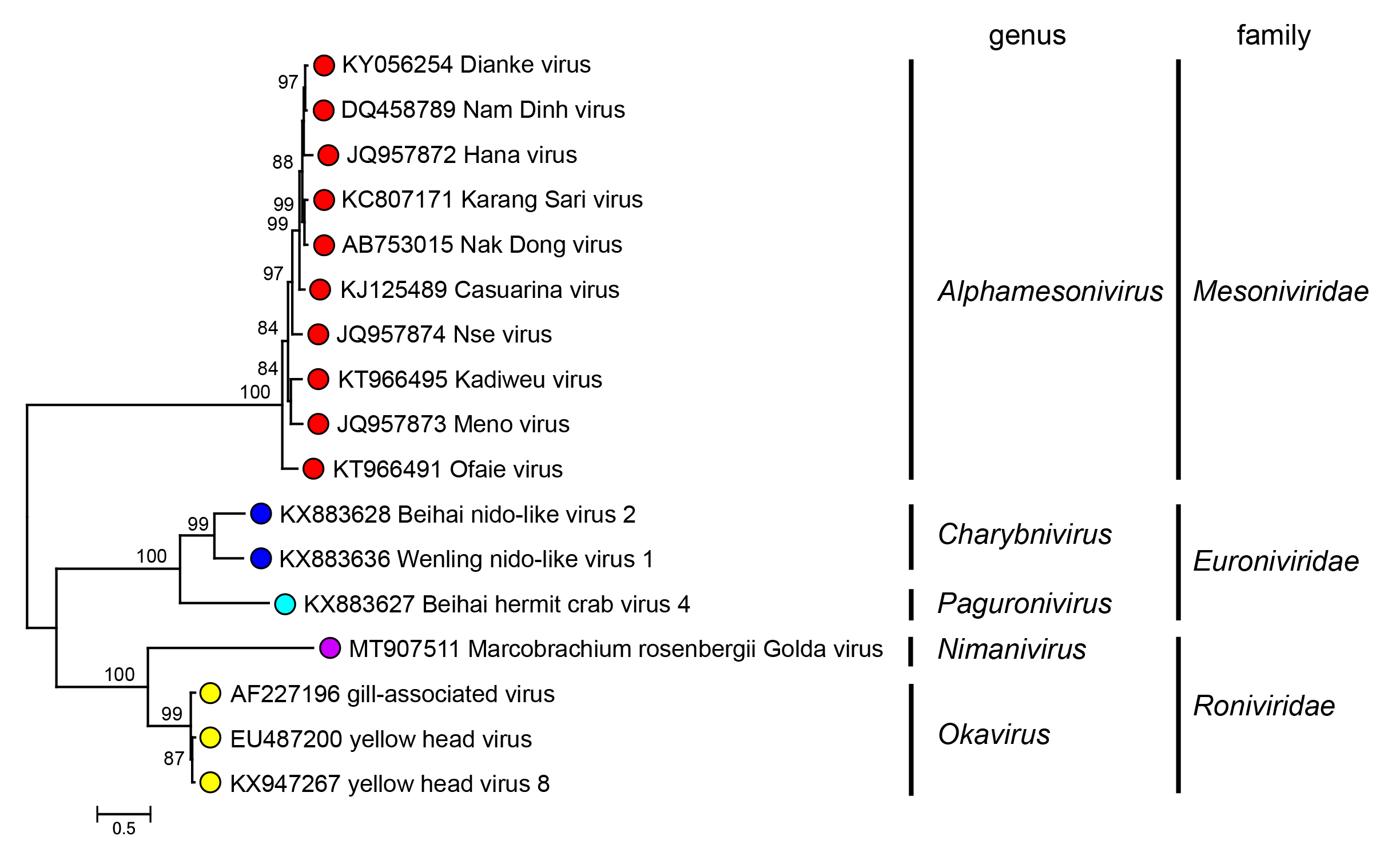 |
| Figure 4.Roniviridae. Maximum Likelihood phylogenetic tree inferred in MEGA 7.0 (Kumar et al., 2016) from an amino acid sequence alignment of the concatenated replicase domains in ORF1b of 17 viruses assigned to the three assigned families of arthropod nidoviruses (Mesoniviridae, Euroniviridae and Roniviridae). The amino acid sequences were aligned in ClustalW and then adjusted manually to reflect predicted conserved motifs in seven identified structural domains (NiRAN, RdRP, ZBD, HEL1, ExoN, N-MT and O-MT) (Saberi et al., 2018). The evolutionary relationships were inferred by using the WAG + frequency model of amino acid substitution (Whelan and Goldman 2001) with sub-tree pruning and re-grafting (SPR) branch-swapping. The initial tree for the heuristic search was obtained automatically by applying the neighbour-joining algorithm to a matrix of pairwise distances estimated using a JTT model (Jones et al., 1992), and then selecting the topology with superior log likelihood value. The tree with the highest log likelihood (−15561.24) is shown. Numbers at nodes indicate the bootstrap support values (1000 iterations) > 80%. The scale bar indicates 0.5 substitutions per site. This phylogenetic tree and corresponding sequence alignment are available to download from the Resources page. |

