Family: Xinmoviridae
Luis Hernandez-Pelegrin, Binit Lamichhane, Stephen R. Sharpe
The citation for this ICTV Report chapter is the summary published as Sharpe and Paraskevopoulou (2023):
ICTV Virus Taxonomy Profile: Xinmoviridae 2023, Journal of General Virology, 104: 001906
Corresponding author: Stephen Sharpe [email protected]
Edited by: Holly R. Hughes, Jens H. Kuhn and Stuart G. Siddell
Posted: September 2023, updated December 2024
Summary
Xinmoviridae is a family of viruses with negative-sense RNA genomes of 9–16 kb (Table 1 Xinmoviridae). The family includes 22 genera and 25 species. Xinmovirids typically infect beneficial and pest insects in Africa, Asia, Europe, and Oceania, but their host range and geographic distribution has not yet been investigated systematically and hence may be broader.
Table 1 Xinmoviridae. Characteristics of members of the family Xinmoviridae
| Characteristic | Description |
| Example | Drosophila unispina virus 1 (KR822819), species Drunivirus chambonense, genus Drunivirus |
| Virion | Not known |
| Genome | 10–16 kb of negative-sense RNA |
| Replication | Not known |
| Translation | Not known |
| Host range | Arthropoda |
| Taxonomy | Realm Riboviria, kingdom Orthornavirae, phylum Negarnaviricota, class Monjiviricetes, order Mononegavirales: 22 genera and 25 species |
Virion
Morphology
Xinmovirids are thus far only known from metagenomics studies. Virions have not yet been visualised and structural proteins have not been studied.
Genome organization and replication
Xinmovirids have negative-sense RNA genomes of 10–16 kb with three to six ORFs (Figure 1 Xinmoviridae) (Parry and Asgari 2018). These ORFs encode at least three proteins that have been identified via comparison with proteins encoded by other viruses in the order Mononegavirales: a glycoprotein, a nucleoprotein, and an RNA directed RNA polymerase (RdRP) (Scarpassa et al., 2019).
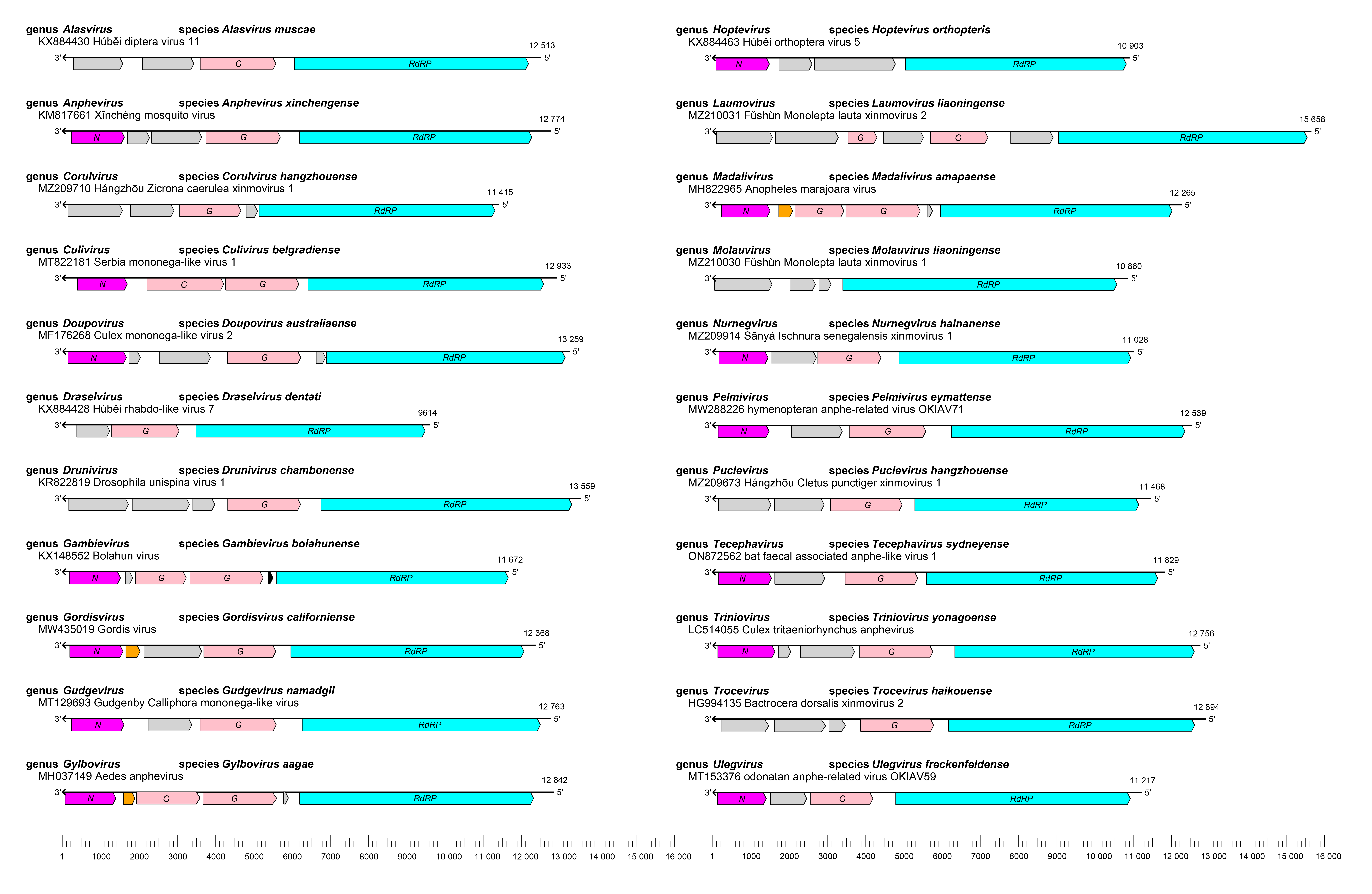 |
| Figure 1 Xinmoviridae. Genome organisation of representative viruses of each genus in the family Xinmoviridae. Boxes show the position of ORFs coloured according to their predicted protein function: N, nucleoprotein gene (magenta); G, glycoprotein gene (pink); RdRP, RNA-directed RNA polymerase gene (cyan); transmembrane protein (orange); putative zinc-finger protein (black). The GenBank accession number and length of each genome is included. |
Biology
Viruses in the family Xinmoviridae have been detected in insects from Africa (Tanzania), Asia (Japan), Europe (Croatia, Germany, Greece, Italy, Serbia, Switzerland), and Oceania (Australia), and may infect insects of more than 12 orders, including Blattodea, Coleoptera, Diptera (Shi et al., 2017, Scarpassa et al., 2019, Manni and Zdobnov 2020, Stanojevic et al., 2020, Zhang et al., 2020, Batson et al., 2021, Sharpe et al., 2021, Lamichhane et al., 2024), Hemiptera, Hymenoptera, and Lepidoptera, Mantodea, Neuroptera, Odonata, Orthoptera, Psocodea, Trichoptera, and Zygentoma (Käfer et al., 2019).
Derivation of names
Alasvirus: from the Latin alas meaning wings, referring to Húběi diptera virus 11, which was discovered by HTS in a pool of two-winged flies. The species epithet muscae derives from musca, the Latin word for fly.
Anphevirus: from the insect genus Anopheles, Xīnchéng anphevirus having been discovered by HTS in Anopheles sinensis mosquitoes. The species epithet xinchengense refers to Xīnchéng, China, the sample location.
Corulvirus: from Zircona caerulea, the species of shieldbug in which Hangzhou zircona caerulea xinmovirus 1 was discovered by HTS. The species epithet hangzhouense derives from its discovered geography location Hangzhou, China.
Culivirus: from the word Culicidae, the family of mosquito, Culex pipiens, in which Serbia mononega-like virus 1 was discovered by HTS. The species epithet belgradiense derives from Belgrade, Serbia, the sampling site.
Doupovirus: from Point Douro, Australia, where Culex mononega-like virus 2 was discovered by HTS in Culex australicus mosquitoes. The species epithet australiaense refers to Australia.
Draselvirus: from dragonflies and damselflies, odonates in which Húběi rhabdo-like virus 7 was discovered by HTS. The species epithet dentati derives from dentatum, the Latin word for toothed (odonate is derived from ὀδούς, the Greek word for tooth).
Drunivirus: from Drosophila unispina, the host in which Drosophila unispina virus 1 was discovered. The species epithet chambonense derives from Chambon, France, the sample location.
Gambievirus: from Gambie virus which was discovered by HTS in Anopheles gambiae mosquitoes sampled in Senegal (Gambie is the French name of Gambia). The species epithet bolahunense derives from Bolahun virus discovered by HTS in Anopheles gambiae mosquitoes sampled in Bolahun, Liberia. The species epithet senegalense derives from Senegal, the sampling location for Gambie virus.
Gordisvirus: from the word Gorgis. The species epithet californiense derives from its discovered geography location California.
Gudgevirus: from the word Gudgenby, the location it was discovered by HTS. The species epithet narnadgii derives from its discovered geography location Narnadgi.
Gylbovirus: from Aedes aegypti and Aedes albopictus, hosts or host cells associated with Aedes anphevirus infection. The species epithet aagae derives from the cell line Aag2.
Hoptevirus: from Orthoptera, the order to which insects in which Húběi orthoptera virus 5 was discovered by HTS are assigned. The species epithet orthopteris is derived from Orthoptera.
Laumovirus: from Monolepta lauta, the species of beetle in which Fushun monolepta lauta xinmovirus 2 was discovered by HTS. The species epithet liaoningense derives from Liaoning province, China, the sampling site.
Madalivirus: from Anopheles marajoara and Anopheles darling, the species of mosquitoes in which viruses in the genus were first discovered. The species epithet amapaense derives from Amapá state, Brazil, the sampling location of Anopheles marajoara virus. The species epithet amazonaense derives from Amazonas state, Brazil, one of two mosquito sampling sites in Brazil where Anopheles darlingi virus was discovered by HTS.
Molauvirus: from Monolepta lauta, the species of beetle in which Fushun monolepta lauta xinmovirus 1 was discovered by HTS. The species epithet liaoningense derives from Liaoning,province, China, the sampling site.
Nurnegvirus: from Ischnura senegalensis, the species of damselfly in which Sanya ischnura senegalensis xinmovirus 1 was discovered by HTS. The species epithet hainanense refers to the sample site of Hainan, China.
Pelmivirus: from Heteropelma amictum, the species of parasitoid wasps in which hymenopteran anphe-related virus OKIAV71 was discovered by HTS. The species epithet eymattense refers to the sample site of Eymatt, Bern, Switzerland.
Puclevirus: from Cletus punctiger, the species of stinkbug in which Hangzhou cletus punctiger xinmovirus 1 was discovered by HTS. The species epithet hangzhouense derives from Hangzhou, Zhejiang province, China, the site of sampling.
Tecephavirus: from Pteropus poliocephalus, the species of bat in which Bat faecal associated anphe-like virus 1 was discovered by HTS. The species epithet sydneyense derives from Sydney, Australia, the sampling site.
Triniovirus: from Culex tritaeniorhynchus, the species of mosquitoes in which Culex tritaeniorhynchus anphevirus was discovered by HTS. The species epithet yonagoense derives from Yonago, Japan, the site of sampling.
Trocevirus: from Bactrocera dorsalis, the species of tephritid fruit fly in which Bactrocera dorsalis xinmovirus 2 was discovered by HTS. The species epithet haikouense derives from Haikou, Hainan, China, the site of sampling.
Ulegvirus: from Libellula and Cordulegaster, the genera of dragonflies in which viruses of this genus were discovered by HTS. The species epithet freckenfeldense derives from Freckenfeld, Rhineland-Palatinate, Germany, the sampling site.
Xinmoviridae: from Xīnchéng mosquito virus, the first virus that was assigned to the family.
Genus demarcation criteria
Members of different genera have RdRP amino acid identities of 60% or less.
Species demarcation criteria
Members of different species within a genus have RdRP amino acid identities of 66% or less.
Relationships within the family
Phylogenetic relationships of Xinmoviridae are shown in Figure 2 Xinmoviridae.
 |
| Figure 2 Xinmoviridae. Phylogenetic tree of xinomovirid RdRP amino acid sequences. Sequences were aligned using MUSCLE v5.2 (Edgar 2022) and a neighbour-joining tree of amino acid distances was created using the Poisson correction within MEGA11 (Tamura et al., 2021). The virus Wǔchāng romanomermis nematode virus 2 (family Lispviridae) was used as an outgroup. Circles at tips are coloured according to genus. |
Relationships with other taxa
Viruses in the family Xinmoviridae are most closely related to members of the families Bornaviridae and Nyamiviridae, these families also being placed in the order Mononegavirales.
Related, unclassified viruses
The list of unclassified viruses which are likely members of the family Xinmoviridae mainly includes viruses associated to different species of Dipteran insects (Caldas-Garcia et al., 2023, de Santana et al., 2023, Litov et al., 2023, Divekar et al., 2024, Lamichhane et al., 2024, Maia et al., 2024). Moreover, a putative xinmovirus was described in Microplitis mediator (Hymenoptera) (Ji et al., 2024), and Oxya chinensis (Orthoptera). The table below includes only those viruses with complete or near-complete sequences, and only viruses with an accession number on repository database.
| Virus name | Accession number | Virus abbre-viation | Length | Host |
| Albipes mosquito Gordis-like virus | PP946246 | 11812 | Psorophora albipes | |
| Bactrocera dorsalis borna-like virus | MN745081 | 12450 | Bactrocera dorsalis | |
| Burswood mono-chu-like virus | PP066189 | 6094 | Culex globocoxitus | |
| draselvirus dasyheleae 1 | BK063243 | DVd1 | 3812 | Dasyhelea sp. |
| draselvirus dasyheleae 2 | BK063244 | DVd2 | 4803 | Dasyhelea sp. |
| Ferox mosquito mononega-like virus | PP946236 | 13366 | Psorophora ferox | |
| Guiyang xinmovirus 1 | MZ209642 | 11813 | Oxya chinensis | |
| Medvezhye Haematopota Xinmo-like virus | OR724669 | 11486 | Haematopota pluvialis | |
| Medvezhye Chrysops Xinmo-like virus | OR724668 | 10398 | Chrysops | |
| Microplitis mediator mononega-like virus strain CHB | BK064877 | 12918 | Microplitis mediator | |
| soldier fly-associated anphevirus | PP410010 | SFaAV | 12420 | Inopus flavus |
| unclassified forcipomyiae 1 | BK063245 | Un-f1 | 12795 | Forcipomyia taiwana |
| unclassified forcipomyiae 2 | BK063246 | Un-f2 | 6687 | Forcipomyia taiwana |
Virus names and virus abbreviations are not official ICTV designations.

