Family: Partitiviridae
Eeva J. Vainio, Sotaro Chiba, Said A. Ghabrial, Edgar Maiss, Marilyn Roossinck, Sead Sabanadzovic, Nobuhiro Suzuki, Jiatao Xie and Max Nibert
The citation for this ICTV Report chapter is the summary published as Vainio et al., (2018):
ICTV Virus Taxonomy Profile: Partitiviridae, Journal of General Virology, 99:17–18.
Corresponding author: Eeva Vainio ([email protected])
Edited by: Peter Simmonds and Stuart G. Siddell
Posted: January 2018, updated May 2018, May 2024
PDF: ICTV_Partitiviridae.pdf (2018 version)
Summary
Partitiviridae is a family of small, isometric, non-enveloped viruses with bisegmented dsRNA genomes of 3–4.8 kbp (Table 1 Partitiviridae). The two genome segments are individually encapsidated. The family includes five genera with characteristic hosts for members of each genus: either plants or fungi for genera Alphapartitivirus and Betapartitivirus; fungi for genus Gammapartitivirus, plants for genus Deltapartitivirus and protozoa for genus Cryspovirus. Partitiviruses are transmitted intracellularly via seeds (plants), oocysts (protozoa) or hyphal anastomosis, cell division and sporogenesis (fungi); there are no known natural vectors. The five partitivirus genera include 45 species.
Table 1 Partitiviridae. Characteristics of members of the family Partitiviridae.
| Characteristic | Description |
| Typical member | Atkinsonella hypoxylon virus 2H (RNA1: L39125, RNA2: L39126), species Betapartitivirus atkinsonellae, genus Betapartitivirus |
| Genome | 3–4.8 kbp of bisegmented dsRNA |
| Virion | Isometric, non-enveloped, 25–43 nm in diameter; dsRNA1 and dsRNA2 are separately encapsidated |
| Replication | Cytoplasmic. Genomic RNA acts as a template for mRNA synthesis within the virus particle; transcription occurs by a semi-conservative mechanism |
| Translation | From monocistronic positive-sense transcripts of both genomic dsRNAs |
| Host Range | Plants, fungi, protozoa |
| Taxonomy | Realm Riboviria, kingdom Orthornavirae, phylum Pisuviricota, class Duplopiviricetes, order Durnavirales; 5 genera including 45 species |
Virion
Morphology
Virus particles are isometric, non-enveloped, 25–43 nm in diameter (Figure 1 Partitiviridae). The capsid is composed of 120 copies of a single coat protein (CP) arranged as 60 dimers with T=1 icosahedral symmetry (Ochoa et al., 2008, Pan et al., 2009, Tang et al., 2010, Nibert et al., 2013). Dimeric surface protrusions are frequently observed on viral capsids. One or two molecules of RNA-dependent RNA polymerase (RdRP) are packaged inside each particle (Nibert et al., 2014).
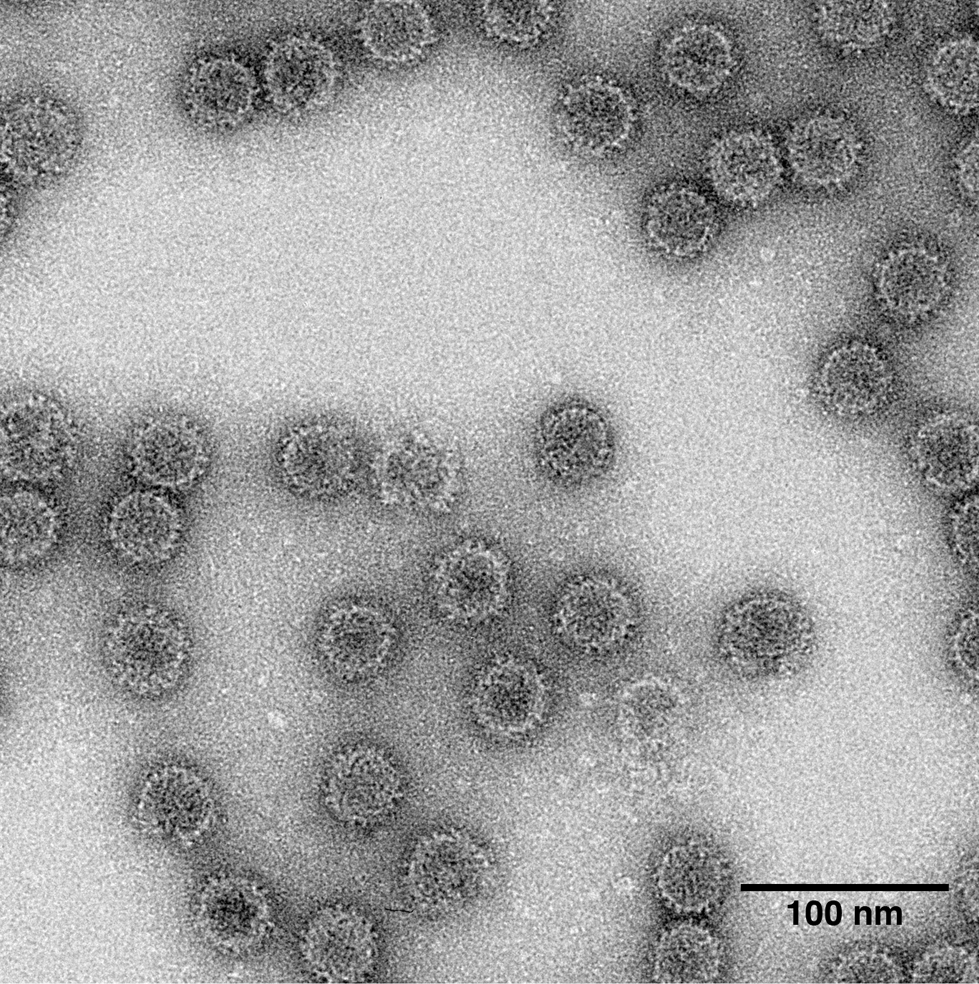 |
| Figure 1 Partitiviridae. Transmission electron micrograph of negatively-stained virions of an isolate of Gammapartitivirus essepenicillii. |
Physicochemical and physical properties
Virion buoyant densities on CsCl gradients range from 1.34 to 1.44 g cm−3 (see below).
Nucleic acid
Members of all five genera possess two essential genome segments, dsRNA1 and dsRNA2, each containing one large ORF on the positive-strand RNA molecule. One segment/ORF (dsRNA1) encodes the RdRP, and the other segment/ORF (dsRNA2) encodes the coat protein (CP) (Figure 2. Partitiviridae). The linear segments are 1.4–2.4 kbp. Each genome segment is separately encapsidated. Some partitiviruses have additional (satellite or defective) dsRNA elements (see below).
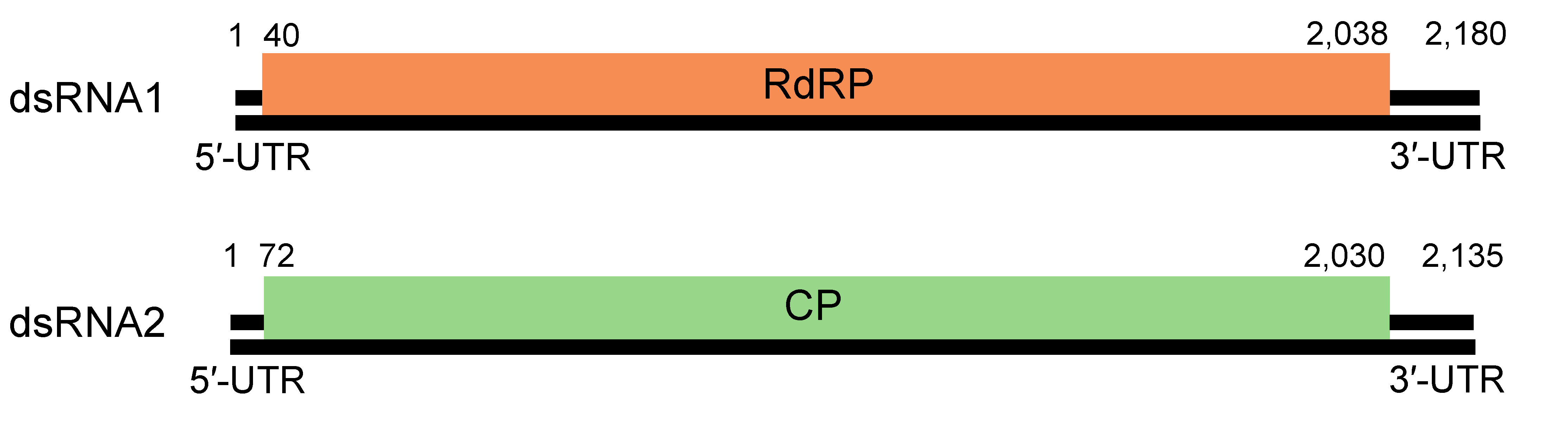 |
| Figure 2. Partitiviridae. Atkinsonella hypoxylon virus (AhV), an isolate of the species Betapartitivirus atkinsonellae, has a bipartite genome consisting of dsRNA1 and dsRNA2 (Oh and Hillman 1995). |
Proteins
There is a single major CP and a separately translated RdRP. Virion-associated RNA polymerase activity is present.
Genome organization and replication
Partitivirus genomes consist of two linear dsRNA segments; the smaller usually codes for the CP and the larger usually codes for the virion-associated RNA polymerase. Each dsRNA is monocistronic. The first 7–20 nucleotides in the 5′-non translated region of the coding RNA are usually highly conserved between the two genome segments of single viruses. The virion-associated RdRP catalyzes in vitro end-to-end transcription of each dsRNA to produce mRNA by a semi-conservative mechanism. Virions accumulate in the cytoplasm. A model for the replication strategy of an isolate of Gammapartitivirus essepenicillii is shown in Figure 3 Partitiviridae (see also (Ghabrial et al., 2008)).
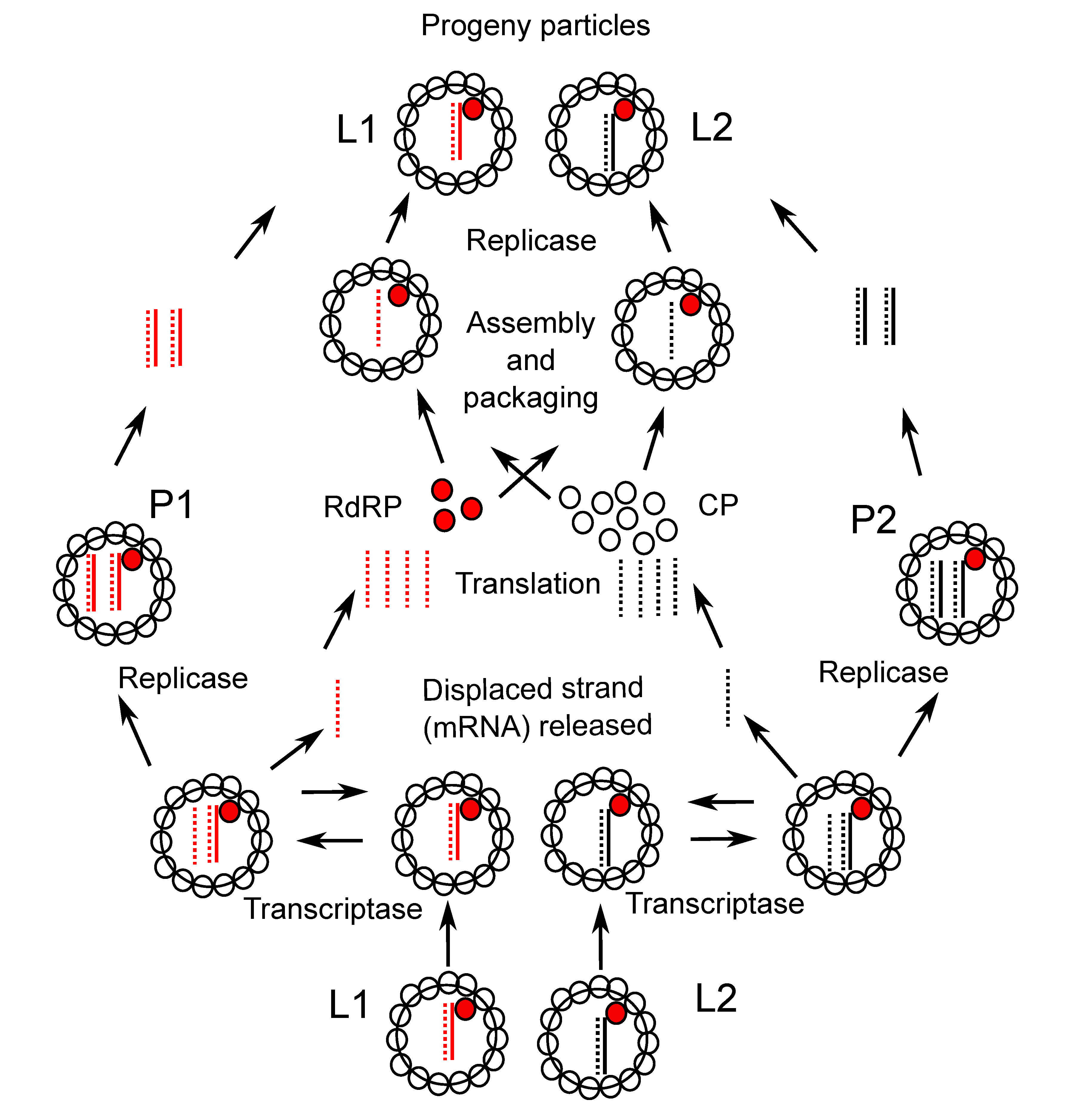 |
| Figure 3. Partitiviridae. Model for replication of Penicillium stoloniferum virus S (PsV-S). Virion particles contain one or two copies of RdRP (red circles), 120 copies of CP (open circles) and a dsRNA encoding either RdRP (L1, red lines) or CP (L2, black lines). Positive-strands are indicated by dashes and negative-strands by solid lines. Within each particle the RdRP mediates positive-strand RNA synthesis using the negative-strand of the genome dsRNA as a template. The newly synthesized positive-strand RNA is retained inside the particle as part of the dsRNA while the parental genomic positive-strand (encoding RdRP or CP) is released from the particle and used in translation by cellular ribosomes or in packaging into a new partitivirus particle. The RdRP is believed to function also as a replicase by catalyzing the synthesis of negative-strand RNA on the positive-strand template within the assembling virion, thereby reconstituting a genomic dsRNA segment. Also indicated is a proposed pathway leading to the production of L1 and L2 particles via P1 and P2 particles that contain two copies of a dsRNA segment (Buck 1978). |
Biology
The partitiviruses are typically associated with persistent infections of their fungal, protozoan and plant hosts. There are no known natural vectors. The fungal partitiviruses are transmitted intracellularly during cell division, hyphal anastomosis and sporogenesis. In some ascomycetes (e.g. Gaeumannomyces graminis), virus is usually eliminated during ascospore formation. Experimental transmission of purified fungal partitiviruses has been reported by transfection of virions into fungal protoplasts (Sasaki et al., 2006, Bhatti et al., 2011, Chiba et al., 2013, Xiao et al., 2014). Alpha- and betapartitiviruses infecting fungal hosts from the genus Heterobasidion are transmitted via hyphal contacts between somatically incompatible host strains (Ihrmark et al., 2002, Vainio et al., 2013, Vainio and Hantula 2016). The plant partitiviruses are transmitted by ovule and by pollen to the seed embryo (Boccardo et al., 1987). There is no graft transmission and apparently no cell-to-cell transport, except at cell division; seed transmission is the only known mode for the transmission of plant partitiviruses. Cryptosporidium partitiviruses disperse via oocysts.
The genera Alphapartitivirus and Betapartitivirus include viruses isolated from both fungi and plants. Therefore, a capacity for occassional successful transmission of these viruses between fungal and plant hosts appears likely. The genus Gammapartitivirus so far comprises only viruses isolated from fungal hosts, and genus Deltapartitivirus so far contains only viruses isolated from plants.
Use of the term “cryptovirus” (from Greek crypto, “hidden, covered, or secret”) in genus names of plant-infecting Partitiviridae members was recently discontinued as many of the fungal partitiviruses have no visible effects on their host and can also be considered “cryptic” viruses; thus, the term “cryptovirus” cannot be accurately used to differentiate plant from fungal partitiviruses. Moreover, some plant partitivirus genes appear to be associated with mutualistic host effects (Nakatsukasa-Akune et al., 2005, Roossinck 2010). The relationship between Cryptosporidium parvum virus 1 (genus Cryspovirus) and its protozoan host can be considered to be mutualistic (Jenkins et al., 2008), whereas some fungal partitiviruses mediate deleterious effects on their hosts, such as reduced growth or sporulation (Bhatti et al., 2011, Chiba et al., 2013, Xiao et al., 2014, Kanematsu et al., 2010, Vainio et al., 2010).
Antigenicity
Virions are effective immunogens. No serological relationships between studied fungal and plant viruses in the family Partitiviridae have been detected.
Derivation of names
Partitiviridae: from the Latin partitius, “divided”
Alphapartitivirus, Betapartitivirus, Gammapartitivirus, and Deltapartitivirus: the prefixes are derived from Greek numbers to designate four partitivirus genera
Cryspovirus: from the host genus name, Cryptosporidium
Genus demarcation criteria
The genus demarcation criteria within family Partitiviridae are:
- Characteristic hosts within each genus (either plants or fungi for genera Alphapartititivirus and Betapartitivirus, fungi for genus Gammapartitivirus, plants for genus Deltapartitivirus and protozoa only for genus Cryspovirus)
- Genome segment and protein lengths within a characteristic range for each genus (Table 2 Partitiviridae)
- < 24% RdRP amino acid sequence identity in pairwise comparisons of viruses from different genera
- Separate phylogenetic grouping of RdRP sequences from each genus
Table 2 Partitiviridae. Characteristic ranges of genome segment and protein lengths within the five genera of family Partitiviridae.
| Genus | dsRNA1 (bp) | dsRNA2 (bp) | RdRP (aa) | CP (aa) |
| Betapartitivirus | 2180–2444 | 2135–2354 | 663–746 | 636–686 |
| Alphapartitivirus | 1873–2027 | 1708–1866 | 581–621 | 463–521 |
| Gammapartitivirus | 1645–1787 | 1445–1611 | 519–539 | 413–443 |
| Deltapartitivirus | 1563–1696 | 1415–1575 | 472–479 | 337–430 |
| Cryspovirus* | 1786 (1836)‡ | 1374 (1510) ‡ | 524 | 319 |
* This genus includes a single species
‡ Genome segments in Cryptosporidium parvum virus 1 strain CSpV1-Iowa (Vong et al., 2017).
Relationships within the family
Phylogenetic analysis based on amino acid sequences of RdRPs of members of the family Partitiviridae led to the identification of five clusters corresponding to the five partitivirus genera (Figure 4 Partitiviridae). Within the genus Betapartitivirus, the plant viruses constitute a highly supported subclade distinct from fungal betapartitiviruses. Similarly, classified plant viruses in the genus Alphapartitivirus form a distinct subcluster among fungal alphapartitiviruses. (Nibert et al., 2014). The three remaining partitivirus genera form three distinct clades that appear to be specific to plants (genus Deltapartitivirus), fungi (genus Gammapartitivirus) and protozoa (genus Cryspovirus).
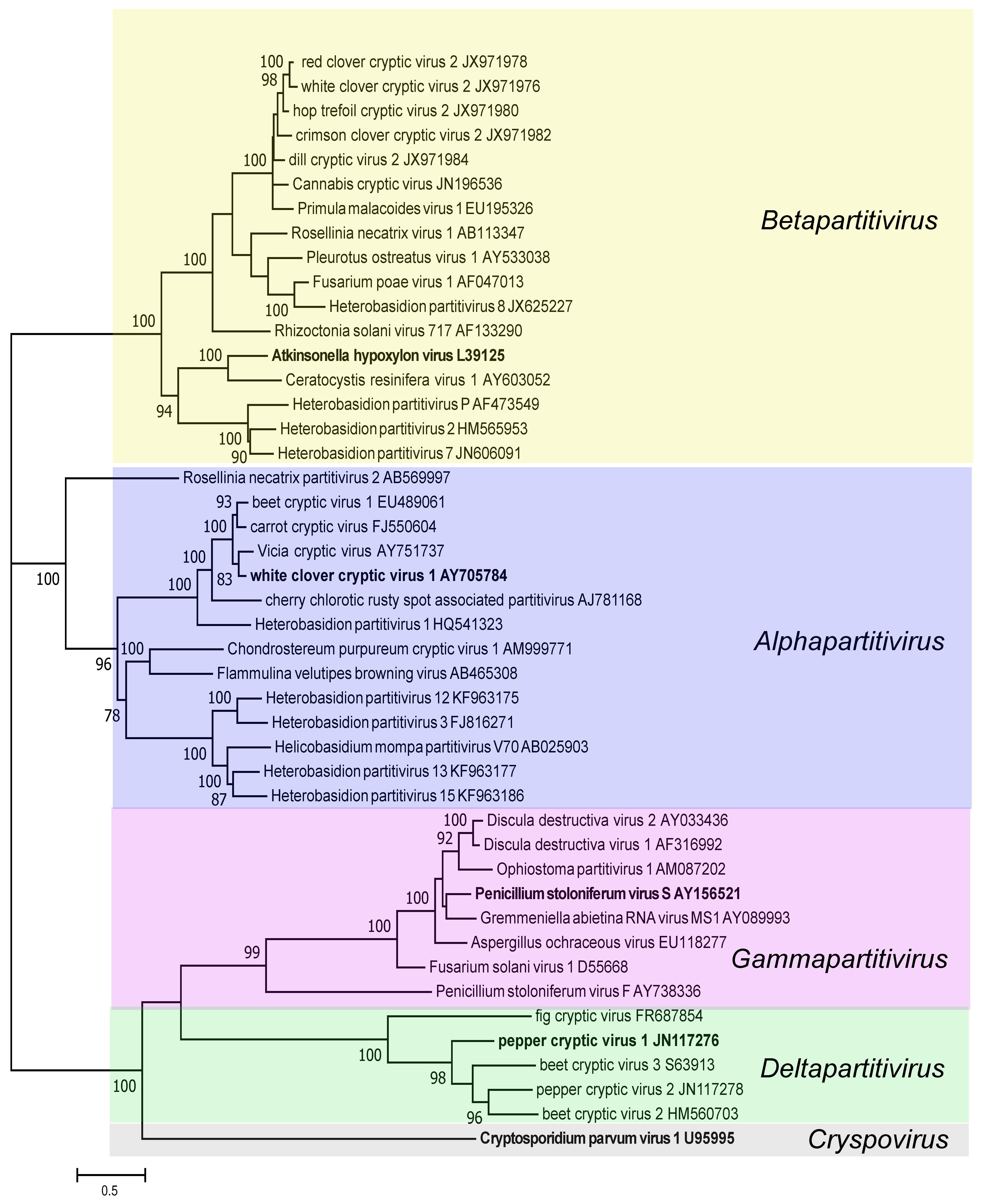 |
| Figure 4. Partitiviridae. Unrooted Maximum Likelihood dendrogram based on the complete amino acid sequences of RdRPs of classified members of the family Partitiviridae. The sequences were aligned using MAFFT v6.814b in Geneious Pro 5.5.8 (Biomatters Ltd.) and the evolutionary history was inferred using MEGA 7 (Kumar et al., 2016). The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (150 replicates) are shown next to the branches (values of ≥70% shown). The evolutionary distances were computed using the Le Gascuel 2008 model (LG+G+I) and are in the units of the number of amino acid substitutions per site. Virus names and GenBank accession numbers corresponding to RdRP nucleotide sequences are given; an isolate of the type species of each genus is shown in bold. This phylogenetic tree and corresponding sequence alignment are available to download from the Resources page. |
Comparison of phylogenetic trees separately constructed from either RdRP or CP sequences revealed essentially the same grouping, corresponding to the current five genera, regardless of the proteins used for analyses (Nibert et al., 2014). The cryo-EM structures for the gammapartitiviruses Penicillium stoloniferum virus F and Penicillium stoloniferum virus S, and the betapartitivirus Fusarium poae virus 1, suggest that the CP subunits of all three viruses have similar folds.
Relationships with other taxa
Based on genome organization and virion morphology, members of the family Partitiviridae have properties similar to members of the family Picobirnaviridae, e.g., the genomes are bisegmented, and the capsids are small (<45 nm in diameter) with 120-subunit T=1 symmetry and share similar structural details. The picobirnaviruses, however, are phylogenetically distinct and infect vertebrates rather than plants, fungi and protozoa; they probably have other basic differences including co-packaging of both genome segments and capacity for extracellular transmission.
Members of the family Megabirnaviridae and genus Botybirnavirus also have bisegmented genomes. Megabirnavirus genomes are substantially larger than those of partitiviruses (8931 bp and 7189 bp for Rosellinia necatrix megabirnavirus 1, a representative of the type species of the genus, Megabirnavirus), and the larger segment encodes the CP and RdRP, whereas the smaller segment encodes two proteins of unknown functions (Chiba et al., 2009). Megabirnaviruses have particles of approximately 50 nm in diameter. Botrytis porri RNA virus 1 is the only member of the genus Botybirnavirus (Wu et al., 2012). The two genome segments are 6215 bp and 5879 bp and virus particles have a diameter of approximately 35 nm. The larger genome segment appears to encode a CP and RdRP, and the smaller segment seems to encode another CP and another protein of unknown function.
A separate and currently unclassified group of fungal dsRNA viruses with two small genome segments (each 1700–2400 bp) includes the mutualistic Curvularia thermal tolerance virus (Marquez et al., 2007) as well as seven other viruses described thus far. The larger genome segment of these viruses has a single ORF encoding the RdRP, whereas the smaller segment has 1–2 ORFs of unknown function. Isometric virus particles of approximately 27 nm in diameter appear to be associated with Curvularia thermal tolerance virus, but no virus particles have been detected with isolates from the other viruses. The RdRP sequences of these viruses form a distinct phylogenetic clade, and a formal proposal to assign them to a new taxon seems likely to be forthcoming.
Based on sequence similarity, partitivirus polymerases resemble those of family Amalgaviridae (Sabanadzovic et al., 2009), which have unsegmented genomes organized similarly to those of many members of family Totiviridae. Members of a recently described group of unsegmented viruses provisionally designated as ‘Unirnaviruses’ (Kotta-Loizou et al., 2015) also share moderate levels of RdRP sequence homology with partitiviruses.
Related, unclassified viruses
Botryosphaeria dothidea partitivirus 1 and Alternaria alternata partitivirus 1 are related to other partiviruses but seem to belong to a lineage distinct from established partitivirus genera (Wang et al., 2014, Xavier et al., 2017).
| Virus name | Accession number | ||
| dsRNA1/RdRP | dsRNA2/CP | dsRNA3 | |
| Botryosphaeria dothidea partitivirus 1 | KF688740 | KF688741 | KF688742 |
| Alternaria alternata partitivirus 1 | KY352402 | KY352403 | |
Virus names and virus abbreviations are not official ICTV designations.

