Family: Simuloviridae
Ying Liu, Shishen Du, Xiangdong Chen, Mike Dyall-Smith, Matti Jalasvuori and Hanna M. Oksanen
The citation for this ICTV Report chapter is the summary published as:
Corresponding author: Hanna M. Oksanen ([email protected])
Edited by: Mart Krupovic and Stuart Siddell
Posted: March 2023
Summary
The family Simuloviridae includes tailless icosahedral viruses with an internal lipid membrane. The capsid is constructed from two major capsid proteins both with a single jelly-roll fold. The genome is a circular double-stranded DNA molecule of 16−19 kbp. All members infect halophilic archaea in the class Halobacteria (phylum Euryarchaeota) and are temperate viruses, their proviruses residing in host cells as extrachromosomal episomes. Once the lytic life cycle is triggered, production of virions causes cell lysis. Members of the family were previously classified as members of the family Sphaerolipoviridae, genus Betasphaerolipovirus.
Table 1. Simuloviridae. Characteristics of members of the family Simuloviridae
| Characteristic | Description |
| Example | Saline Natrinema sp. J7‐1 virus 1 (SNJ1; AY048850), species Yingchengvirus sinense |
| Virion | Tailless icosahedral virion with an internal lipid membrane, diameter about 70 nm, the capsids are constructed from two major structural proteins, both with a single jelly-roll fold |
| Genome | Circular dsDNA of 16−19 kbp |
| Replication | Rolling circle replication |
| Translation | Prokaryotic translation using viral mRNA and host ribosomes |
| Host range | Halophilic archaea of the genera Natrinema and Haloterrigena, order Natrialbales |
| Taxonomy | Realm Singelaviria, kingdom Helvetiavirae, phylum Dividoviricota, class Laserviricetes, order Halopanivirales: one genus, three species |
Virion
Morphology
Virions of Saline Natrinema sp. J7‐1 virus 1 (SNJ1) are tailless icosahedra with a diameter of approximately 70 nm (Figure 1.Simuloviridae). The virion consists of an icosahedral protein shell constructed from two single jelly-roll major capsid proteins PB2 and PB6, and an internal membrane vesicle enclosing the viral circular dsDNA genome (Zhang et al., 2012, Liu et al., 2015).
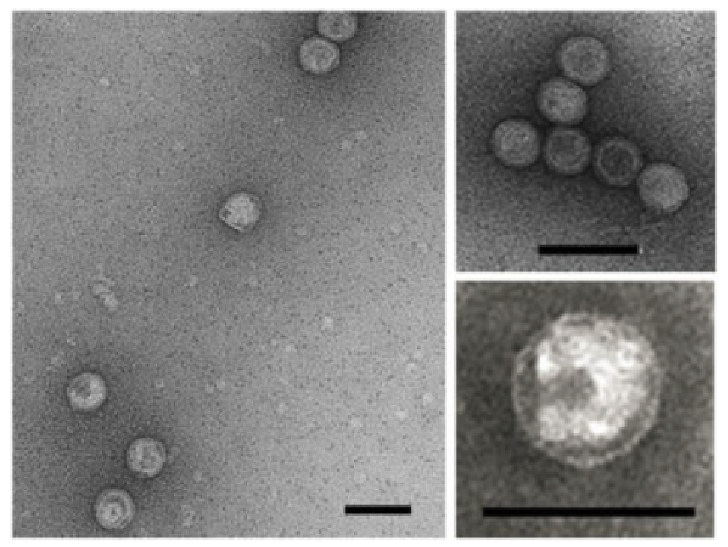 |
| Figure 1.Simuloviridae. Electron micrographs of Saline Natrinema sp. J7-1 virus 1 virions. Samples were negatively stained with 2% (wt/vol) uranyl acetate. Scale bars, 100 nm. Reproduced with permission from (Liu et al., 2015). |
Figure 1.Simuloviridae. Electron micrographs of Saline Natrinema sp. J7-1 virus 1 virions. Samples were negatively stained with 2% (wt/vol) uranyl acetate. Scale bars, 100 nm. Reproduced with permission from (Liu et al., 2015).
Physicochemical and physical properties
SNJ1 virions are typically stable at high ionic strength (above 4 M NaCl), however, infectivity is lost within 5 hours when NaCl concentration is lowered to 2.5 M (Mei et al., 2007, Mei et al., 2015). Mg2+ is also required for virus infectivity (Mei et al., 2007). Virions are stable at temperatures from 4–40 °C and under slightly alkaline conditions (pH 7.5–9). Infectivity is sensitive to organic solvent chloroform (Mei et al., 2007). The buoyant density of virions in CsCl is 1.30 g/ml (Liu et al., 2015).
Nucleic acid
The circular dsDNA genomes of SNJ1, Natrinema versiforme icosahedral virus 1 (NVIV1) and Haloterrigena jeotgali icosahedral virus 1 (HJIV1) are 16 492 bp, 18 925 bp and 17 189 bp, respectively (Zhang et al., 2012, Chen et al., 2020). The G+C content of these genomes is 61–64%, comparable to or slightly lower than those of their corresponding hosts, which have G+C contents of 64–65% (Feng et al., 2012, Cha et al., 2015, DasSarma et al., 2019).
Proteins
The SNJ1 virion contains two major capsid proteins (PB2 and PB6), both having single jelly-roll fold, as well as eight minor capsid proteins (PB1 and VP1–VP7). The virus encodes an A32-like genome packaging ATPase which is a signature for icosahedral membrane containing dsDNA viruses (Zhang et al., 2012).
Lipids
The lipids that form the internal membrane underneath the icosahedral capsid are selectively acquired from the pool of host lipids. The lipid species phosphatidylglycerol is enriched in SNJ1 virions (Liu et al., 2015).
Carbohydrates
No information available.
Genome organization and replication
The circular dsDNA genome of members of the family Simuloviridae is 16–19 kbp and contains about 30 open reading frames (ORFs) (Figure 2.Simuloviridae). Simuloviruses share 17–21 homologous ORFs including replication, regulation and virion structure related genes, and these gene modules are generally collinear among different viruses (Chen et al., 2020) (Figure 3.Simuloviridae).
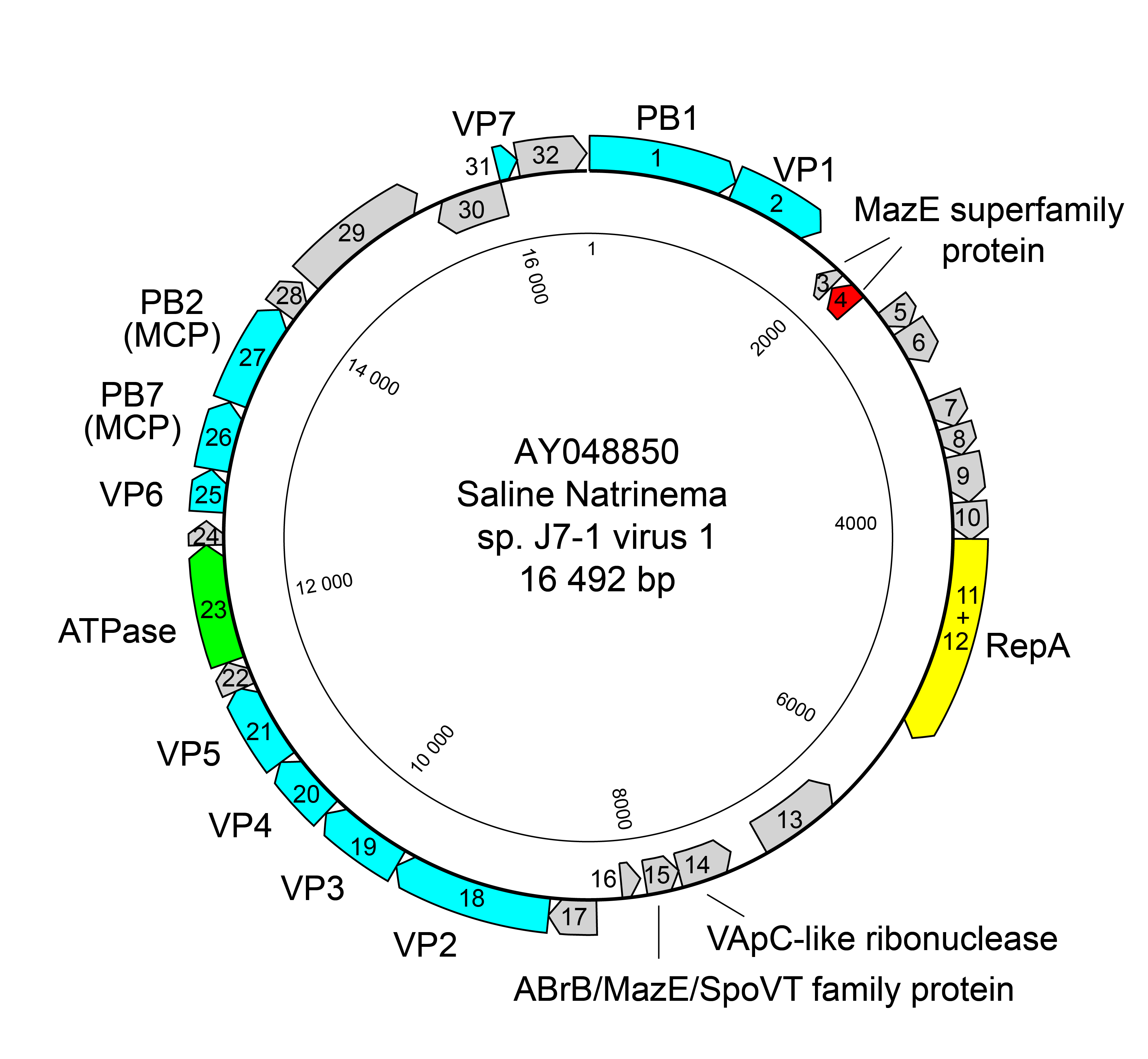 |
| Figure 2.Simuloviridae. Genome map of Saline Natrinema sp. J7‐1 virus 1. Blue - structural proteins; yellow - replication initiation protein RepA; green - putative genome packaging ATPase; pink - transcriptional regulator controlling virus lysis-lysogeny switch. |
Members of the family Simuloviridae have a temperate life cycle, their proviruses residing in host cells as extrachromosomal episomes (Ye et al., 2003, Zhang et al., 2012, Chen et al., 2020). The SNJ1-encoded ORF4 is a transcriptional regulator belonging to the MazE superfamily and controls the lysis-lysogeny switch. The virus is triggered to undergo the lytic life cycle upon treatment with DNA damaging agent mitomycin C (Chen et al., 2020). Homologs of SNJ1 ORF4 are also encoded by HJIV1 and NVIV1, implying a common lysis-lysogeny regulatory mechanism employed by simuloviruses (Chen et al., 2020). SNJ1 genome replication employs a rolling-circle mechanism, which depends on the virus-encoded replication initiation protein RepA, which belongs to the HUH endonuclease superfamily (Wang et al., 2016, Wang et al., 2018). A RepA homolog has been identified in HJIV1, but not in NVIV1, suggesting that a rolling-circle replication mechanism is also employed by HJIV1.
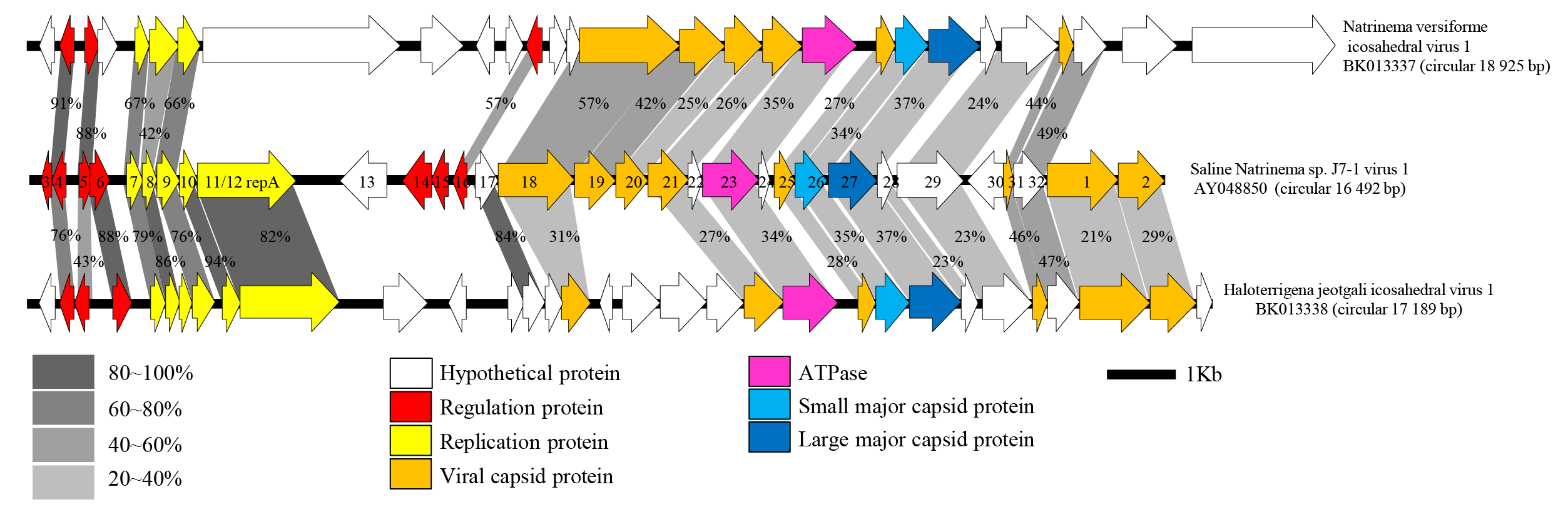 |
| Figure 3.Simuloviridae. Genome alignment of SNJ1, NVIV1 and HJIV1. The circular genome of SNJ1 is shown with ORFs numbers linearized between ORFs 2 and 3. Genes encoding transcriptional regulators and those involved in genome replication are indicated with red or yellow arrows, respectively. Capsid proteins identified by mass spectrometric analysis in the SNJ1 virion (Zhang et al., 2012) are shown in orange. The two major capsid proteins are shown in light and dark blue, whereas the putative genome packaging ATPase is shown in pink. Homologous ORFs are connected with grey shading, with the numbers representing the percentages of amino acid sequence identities. The figure is reproduced from (Chen et al., 2020). |
Biology
Members of the family Simuloviridae were initially identified as plasmids in their host cells and were later proved to be bona fide viruses; virion production has been observed for SNJ1. The plasmid-like episomes are their proviral form (Ye et al., 2003, Zhang et al., 2012, Chen et al., 2020). The lysogenized hosts are halophilic archaea belonging to the genera Natrinema and Haloterrigena (order Natrialbales) isolated from hypersaline environments in China (SNJ1), South Korea (HJIV1) and Bolivia (NVIV1). Simuloviruses encode an antitoxin MazE superfamily protein (SNJ1 ORF4) that governs the lysis-lysogeny switch and superinfection immunity by acting as a repressor of SNJ1 replication (Chen et al., 2020). Active SNJ1 virus replication can be induced by mitomycin C treatment, producing approximately 104 to 1011 PFU/ml (Zhang et al., 2012). The infection cycle of SNJ1 on the sensitive host Natrinema sp. J7-2 is about 10 hours with a release of an average of 100 virions per cell (Zhang et al., 2012).
Derivation of names
Simuloviridae: from the Latin simulo meaning pretend, referring to the fact that genomes of the viruses in this genus were initially thought to be plasmids, i.e., viruses pretending to be plasmids
Yingchengvirus: after Yingcheng city (Hubei province, China), location of the salt mine from which the virus Saline Natrinema sp. J7-1 virus 1, the first representative of the family, was isolated.
Genus demarcation criteria
There is only one genus in the family.
Species demarcation criteria
95% global genome identity is used as a species demarcation criterion, consistent with the thresholds used for other prokaryotic viruses.
Relationships within the family
The family includes three species. NVIV1 shows 82.9% nucleotide sequence identity to SNJ1 over 31% of the genome length, whereas HJIV1 displays 87.9% sequence identity to SNJ1 over 46% of the genome. NVIV1 and HJIV1 are 80% identical over 67% of the genome.
Relationships with other taxa
Simuloviruses have similar virion structures to those of viruses in the families Sphaerolipoviridae and Matsushitaviridae. Members of these three families all have external icosahedral capsids constructed from two single-jelly-roll major capsid proteins and have an internal lipid membrane enclosing dsDNA genomes of either circular or linear form (Rissanen et al., 2013, De Colibus et al., 2019, Santos-Pérez et al., 2019). Members of the families Simuloviridae and Sphaerolipoviridae both infect halophilic archaea, while those of the family Matsushitaviridae infect thermophilic bacteria. Although similar at structural level, viruses from these three families are distinct at genomic sequence level. Genome Relationship Applied to Virus Taxonomy (GRAViTy) analysis, which has been designed to recognize family level groups (Aiewsakun and Simmonds 2018), reveals divergence among members of the three families equivalent to that between most families of eukaryotic viruses (Aiewsakun et al., 2018). These three families, Simuloviridae, Sphaerolipoviridae and Matsushitaviridae, are assigned to the order Halopanivirales, class Laserviricetes, phylum Dividoviricota, kingdom Helvetiavirae. The kingdoms Helvetiavirae and Bamfordvirae share a common principle of capsid architecture and form the realm Varidnaviria. Unlike members of the kingdom Helvetiavirae, the capsids of viruses in the kingdom Bamfordvirae are composed of single major capsid protein species of double jelly-roll fold.

