Family: Hepadnaviridae
Genus: Orthohepadnavirus
Distinguishing features
Viruses of this genus infect mammals, with a narrow host range for members of each virus species. The only known natural hosts of members of the species Hepatitis B virus are humans and apes (chimpanzees, gorillas, orangutans, and gibbons). Virions of hepatitis B virus (HBV) are 40–45 nm in diameter with a 32 or 36 nm internal nucleocapsid. Subviral HBV particles are typically spherical (16–25 nm diameter) or filamentous (20 nm diameter of variable length). The genome of HBV is 3.2 kb with a cohesive overlap of 240 bp. The viruses have an SHBs protein of approximately 226 amino acid residues (aa) as a major envelope protein, an MHBs protein of about 271 aa (which appears unnecessary for infection in experimental situations) and an LHBs protein of about 400 aa.
The surface, proteins are partially glycosylated, thus generating doublets upon gel electrophoresis, e.g. for HBV, P24/GP27 for SHBs, P39/GP42 for LHBs, and, in the case of MHBs, GP33/GP36, due to an additional glycosylation in the preS2 sequence. The HBV core protein is approximately 180 aa, and the virus encodes an X protein of 154 aa whose function in the virus life cycle is uncertain, but it has been shown to serve as a pleiotropic transactivator in vitro.
Virion
See discussion under family description.
Genome organization and replication
See discussion under family description.
Biology
HBV may cause, as a consequence of the host immune response to infection, acute and chronic hepatitis, and the immune complex diseases periarteritits nodosa, glomerulonephritis, and infantile papular acrodermatitis. Late sequelae are liver cirrhosis and hepatocellular carcinoma. An asymptomatic HBeAg positive carrier state with high viremia may develop, particularly after perinatal infection or under immune suppression.
Horizontal transmission of HBV usually occurs by: (1) percutaneous contact with infected blood or body fluids, e.g. intravenous drug abuse or use of infected blood or blood products; (2) sexual contact; (3) perinatal transmission from an infected mother; and (4) “inapparent horizontal” transmission, particularly between children in low socio-economic settings, thought to be due, at least in part, to unrecognized exposure to open skin breaks or mucous membranes. In communities with a high prevalence of infection, routes (3) and (4) predominate, while in low prevalence communities, infections are acquired later in life and involve particularly routes (1) and (2).
Acute hepatitis occurs in woodchucks and squirrels when infected with their respective viruses, and chronic infection may cause hepatocellular carcinoma even greater in frequency than that in chronic carriers of HBV. In infections with WHV, hepatocellular carcinoma frequently occurs within 2 years after infection. However, the proposed mechanisms for the oncogenicity of HBV, WHV and ground squirrel hepatitis B virus (GSHV) are different. Also, both the incidence and typical time scales differ, being greatest with WHV and least with HBV. Woolly monkey hepatitis B virus (WMHBV) causes hepatitis in its host but is not yet known to play a role in hepatocancerogenesis.
Differences in host range: HBV infection is limited to primates, but HBV may infect primary hepatocyte cultures from the tree shrew Tupaia belangeri, which belongs to a mammalian order, Scandentia, that is classified along with primates and Dermoptera in the Euarchonta. GSHV infection has been experimentally transferred to chipmunks and woodchucks but not to several related ground squirrel species. WHV also has a narrow host range, being reported not to infect Beechey ground squirrels (the source of GSHV) or other rodent species.
WMHBV preferentially infects the woolly monkey but may be transmitted to the spider monkey. WMHBV is only with difficulty transmitted to the chimpanzee, which is highly susceptible to human HBV.
Antigenicity
At least five antigenic specificities have been identified for HBsAg. A group determinant (a) is shared by virtually all HBsAg strains located in residues 120–136. Mutations in this region have been found in viruses present in immunized individuals who subsequently became infected, in HBV carriers, particularly in occult HBV infection without detectable HBsAg, and in infected individuals given immunotherapy. Two pairs of subtype determinants (d, y and w, r) have been demonstrated that are generally mutually exclusive and thus behaving as alleles. At the molecular level, the d/y and w/r expression has been shown to reside at residues 122 and 160, respectively, and at both these sites changes in serological subtype were mediated by a shift from lysine to arginine (Okamoto et al., 1987). Additional antigenic heterogeneity has incorrectly been attributed to variation of w. Thus, eight major serological subtypes were designated as ayw1, ayw2, ayw3, ayw4, ayr, adw2, adw4, and adr. The molecular basis for this serological variation was later shown to be located to residues 126 and 127 but also in interaction with other substitutions (Norder et al., 1992).
DNA sequence analysis has now replaced antigenic typing in defining nine viral genotypes differing from each other by 7.5–15% at the nucleotide level of complete genomes and designated with A to J. Different genotypes have different geographical distributions, and there is some, but not complete, correspondence between genotype and serological subtype
HBeAg and HBcAg proteins share common sequences and epitopes, but also contain epitopes that distinguish these two proteins from each other. HBeAg is a 16 kDa truncated derivative of HBcAg. It is found as a soluble antigen in the serum of viraemic patients. HBcAg is more conserved and has been found to cross-react more strongly with the woodchuck hepatitis B virus (WHV) core antigen than is seen between the corresponding surface antigens. In much of the earlier literature, the term surface antigen or HBsAg is used arbitrarily to refer to either the antigenic specificity, various protein products of the preS1/preS2/S gene, or the empty 17–22 nm HBsAg-bearing particles. The term “antigen” should not be used if “protein” or “particle” is intended. Similar considerations apply to the use of “core antigen”.
Species demarcation criteria
The species demarcation criterium in the genus is, as for avihepadnaviruses, more than about 20% nucleotide sequence divergence of complete genomes. The nucleotide sequence divergences between complete genomes are 38% between WHV and HBV and 22% between WMHBV and HBV, and 36% between WMHBV and WHV.
Although, GSHV and WHV differ by only 15% in the nucleotide sequence of their genomes, WHV has been reported not to infect Beechey ground squirrel as the mentioned source of GSHV. The currently unclassified Arctic ground squirrel hepatitis virus (AGSHV) differs from GSHV with about 15% at the nucleotide level.
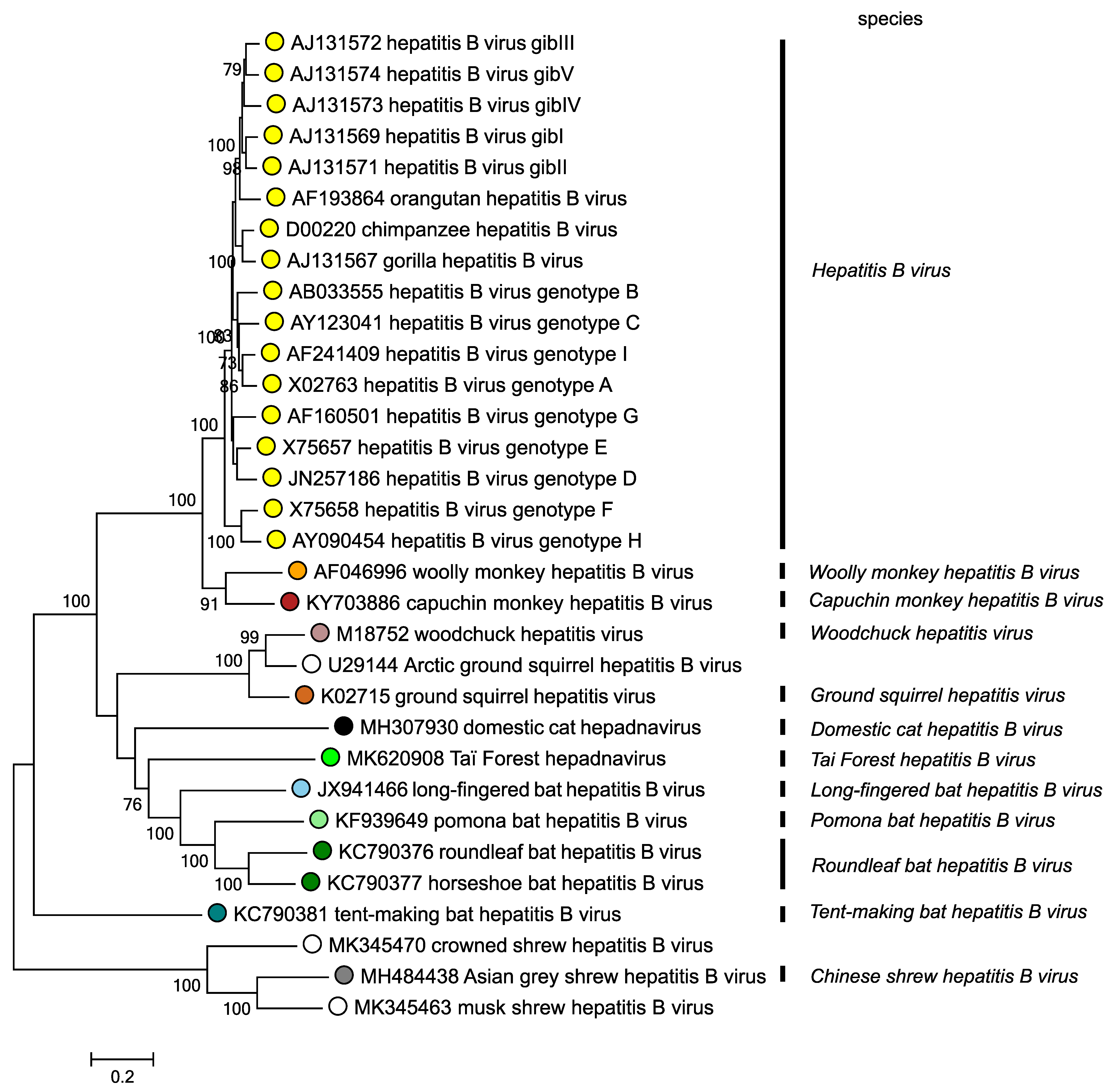
The type species of the genus is Hepatitis B virus. HBV infects humans and primates. There are an additional 11 species belonging to the genus Orthohepadnavirus, members of which all infect mammals (Figure 1. Orthohepadnavirus). Members of Orthohepadnavirus species are so far identified from six mammalian orders: insectivores (1 virus species), carnivores (1 virus species), artiodactyls (1 virus species), bats (4 virus species), rodents (2 virus species), and primates (3 virus species).
 |
|
Figure 1. Orthohepadnavirus. Phylogenetic tree based on polymerase gene sequences of 32 orthohepadnaviruses. . Maximum likelihood trees were produced using the General Time Reversible model with a gamma distribution of variation including invariant sites, and conducted using MEGA7 (Kumar et al., 2016). Numbers indicate where bootstrap support for branches was > 70%. Circles at tips are coloured according to species; unclassified viruses have unfilled circles. This phylogenetic tree and corresponding sequence alignment are available to download from the Resources page. |
Since the Amerindian HBV genotypes F and H are more divergent that those of apes and gibbons these are now regarded conspecific with HBV.
Related, unclassified virus
|
Virus name |
Accession number |
Virus abbreviation |
|
Arctic ground squirrel hepatitis virus |
AGSHV |
|
|
crowned shrew* |
CSHBV |
|
|
musk shrew* |
MSHBV |
Virus names and virus abbreviations are not official ICTV designations.

