Family: Lipothrixviridae
Chapter Version: ICTV Ninth Report; 2009 Taxonomy Release
General properties
Virions are flexible filaments that vary from 410 to 2200 nm in length and 24 to 38 nm in diameter. They are enveloped, and the envelope consists of viral proteins and host derived lipids. The helical nucleoprotein core contains a single molecule of linear dsDNA, 15.9–56 kbp long. Virion ends have specific structures that vary between genera (Figure 1).
Genus Alphalipothrixvirus
Type species Thermoproteus tenax virus 1
Distinguishing features
The width of the virion significantly exceeds that of other members of the family, most likely reflecting exceptional properties of the helical core. The terminal structures are also exceptional: both ends are rounded and do not taper, as in case of members of the other genera. None of the putative genes of the sole member of the genus shows any sequence similarity to sequences in extant databases.
Virion properties
Morphology
The virion is a rigid rod, about 410 nm long and 38 nm in diameter, with apparently asymmetric extensions at each end. The envelope encloses a helical core consisting of dsDNA and two DNA-binding proteins (Figures 1, 2).
Physicochemical and physical properties
Virion Mr is 3.3 ×108 and buoyant density in CsCl is 1.25 g cm−3. The virions are stable at 100 °C and a fraction remains viable after autoclaving at 120 °C. Particles maintain their structure in 6M urea and 7M guanidine hydrochloride. Detergents (e.g. Triton X-100 and octylglycoside), dissociate virions into viral cores, containing the DNA and DNA-binding proteins, and viral envelopes, containing isoprenyl ether lipids and capsid proteins (Figure 2). The virion consists of about 3% (w/w) DNA, 75% protein and 22% isoprenyl ether lipids.
Nucleic acid
The virion contains one molecule of linear dsDNA of 15.9 kbp. The ends of the molecule are masked in an unknown manner.
Proteins
The virion contains at least four proteins with molecular masses of 12.9, 16.3, 18.1 and 24.5 kDa. The two smallest form the virion core and highly hydrophobic 18.1 kDa protein is present in the envelope. Additional minor proteins may be present.
Lipids
The virion envelope carries the same lipids as the host membrane, essentially diphytanyl tetraether lipids. The envelope has a bilayer structure. The phosphate residues of the phospholipids are oriented towards the inside and the glycosyl residues towards the outside of the particles.
Carbohydrates
No information available.
Genome organization and replication
About 85% of the total genome sequence has been determined. The genome contains several transcription units. So far, the function of only a few genes is known, among them those encoding the four structural proteins. These genes are not linked. The genome carries two low complexity sequence regions consisting of the repetitive sequence CCNACN (where N is any nucleotide), one located within the tpx gene and the other in a noncoding region. Frequent recombination appears to occur between these two regions which generates multiple variants of the TPX protein, which is present in infected cells in high amounts, however, is absent among virion proteins. No information is available on genome replication.
Antigenic properties
No information available.
Biological properties
Host range is limited to the hyperthermophilic archaeaon Thermoproteus tenax. Adsorption and infection appear to proceed via interaction of the terminal protrusions of the virion with the host pili. The virus is temperate and virions are released by cell lysis. The viral genome is present in the host cells in a linear non-integrated form, and only its fragments were found to be integrated in the host chromosome. Mature virions assemble in host cell lumen prior to their release.
List of species demarcation criteria in the genus
Not applicable.
List of species in the genus Alphalipothrixvirus
| Thermoproteus tenax virus 1 |
|
|
| Thermoproteus tenax virus 1 | [X14855] | (TTV-1) |
Species names are in italic script; names of isolates are in roman script. Sequence accession numbers [ ] and assigned abbreviations ( ) are also listed.
Other related viruses which may be members of the genus Alphalipothrixvirus but have not been approved as species
None reported.
Genus Betalipothrixvirus
Type species Sulfolobus islandicus filamentous virus
Distinguishing features
The virions are the longest in the family. Their terminal structures, as well as DNA arrangement in the virion core, differ from those of members of the other genera. The genomes show a limited sequence similarity to the genomes of members of the other genera.
Virion properties
Morphology
The virion is a flexuous filament, about 2000 nm long and 23 nm wide (Figures 1, 3). It consists of a cylindrical 3.1 nm thick envelope which encases an inner core consisting of DNA and DNA-binding proteins. Each end of the virion is tapered and carries short filaments; their number is either three or six, dependent on the species.
Physicochemical and physical properties
The buoyant density of the virions is about 1.3 g cm−3. Short treatment with detergents removes the terminal structures and the envelope and uncovers a filament about 17 nm in width. The cylindrical envelope contains small globular subunits arranged in a helical formation. The envelope encases an inner core with two parallel rows, producing a zigzag pattern, which is likely to be the projection of a double row of nucleosome-like particles (Figure 3). The linear dsDNA appears to wrap around the nucleosome-like particles.
Nucleic acid
The genome is linear dsDNA, varying is size from 36.895 to 41.050 kbp. The termini of the two strands are not covalently linked, and are strongly associated with protein(s).
Proteins
The virion carries two major proteins, of about 17.0–18.5 and 22.5–25.0 kDa, in equal amounts, which are involved in formation of the virion core. Among six minor proteins with molecular masses of in the range of 8.0–19.0 and 30.5 kDa; four are conserved in all known species and two are exclusive to only two viruses.
Lipids
In the virion, presumably in the envelope, modified host lipids were detected.
Carbohydrates
No information available.
Genome organization and replication
The genomes of different species reveal a high level of conservation in both gene content and gene order over large regions (Figure 4). Other shared features of the genomes are: (i) large inverted terminal repeats exhibiting conserved, regularly spaced direct repeats; (ii) a highly conserved operon encoding the two major structural proteins; and (iii) multiple overlapping ORFs which may be indicative of gene recoding. A few predicted gene products of each species, in addition to the structural proteins, could be assigned functions, including a putative helicase, a nuclease, a protein phosphatase, glycosyltransferase and transcription regulators. No information is available on DNA replication.
Antigenic properties
No information available.
Biological properties
The host range is restricted to hyperthermophilic archaea from the genera Sulfolobus and Acidianus. The virus genome does not integrate into the host chromosome. Host cell lysis was not observed. A population of infected host cells continuously produces the virus. Details of virus–host interactions are not known.
List of species demarcation criteria in the genus
Species in the genus differ in the length and the terminal structure of the virion, the size and nucleotide sequence of the genomes and their host range.
List of species in the genus Betalipothrixvirus
| Acidianus filamentous virus 3 |
|
|
| Acidianus filamentous virus 3 | [AM087120] | (AFV-3) |
| Acidianus filamentous virus 6 |
|
|
| Acidianus filamentous virus 6 | [AM087121] | (AFV-6) |
| Acidianus filamentous virus 7 |
|
|
| Acidianus filamentous virus 7 | [AM087122] | (AFV-7) |
| Acidianus filamentous virus 8 |
|
|
| Acidianus filamentous virus 8 | [AM087123] | (AFV-8) |
| Acidianus filamentous virus 9 |
|
|
| Acidianus filamentous virus 9 | [EU545650] | (AFV-9) |
| Sulfolobus islandicus filamentous virus |
|
|
| Sulfolobus islandicus filamentous virus | [AF440571] | (SIFV) |
Species names are in italic script; names of isolates are in roman script. Sequence accession numbers [ ] and assigned abbreviations ( ) are also listed.
Other related viruses which may be members of the genus Betalipothrixvirus but have not been approved as species
| Desulfurolobus ambivalens filamentous virus | (DAFV) |
| Thermoproteus tenax virus 2 | (TTV-2) |
| Thermoproteus tenax virus 3 | (TTV-3) |
Genus Gammalipothrixvirus
Type species Acidianus filamentous virus 1
Distinguishing features
The virion has distinctive claw-like structures at its termini (Figures 1, 5). The virion is the shortest in the family, and the genome is the smallest and significantly differs in sequence from the genomes of other members of the family. Moreover the genome does not carry long ITRs, as other members of the family.
Virion properties
Morphology
The filamentous virion is 900 nm long and 24 nm wide (Figures 1, 5). Terminal structures at both ends of the linear virion are identical and resemble claws. They are 20 nm in diameter. Each claw is connected to the virion body by a 60 nm long tapering appendage and a collar (Figure 5). The body of the virion is covered with an envelope about 3.4 nm thick (Figure 6).
Physicochemical and physical properties
The buoyant density is about 1.3 g cm−3. The virion envelope is partially degraded by treatment with 0.3% Triton X-100 and completely removed by treatment with 0.1% SDS. Concomitant with the removal of the envelope, the terminal structures are removed. Conformational changes occur in the virion termini at an initial phase of adsorption: the claw-like terminal structures fold and keep the virion tightly attached to the host cell pili.
Nucleic acid
The genome is linear dsDNA of 20.869 kbp. The ends are modified in an unknown manner. The short inverted terminal repeat CG10 is present at the termini, preceded by about 350-bp-long sequence which contains clusters of short direct repeats of the sequence TTGTT or close variants thereof. The structural organisation resembles the telomeric ends of linear eukaryotic chromosomes.
Proteins
The virion carries two major proteins P132 and P140, and several minor proteins. They bind DNA and form filaments when incubated with linear dsDNA. A C-terminal domain is identified in their crystal structure with a four-helix-bundle fold. In the topological model, the genomic dsDNA superhelix wraps around P132 basic core, and the P140 basic N-terminus binds genomic DNA, while its lipophilic C-terminal domain is embedded in the lipidic outer shell (Figure 6). The four-helix bundle fold of P132 and P140 is identical to that of the coat protein of the members of the family Rudiviridae.
Lipids
In the virion, presumably in the envelope, modified host lipids were detected.
Carbohydrates
No information available.
Genome organization and replication
From 40 putative genes, seven have homologs in betalipothrixviruses and rudiviruses (Figure 7). Three of these, ORFs 135, 426 and 223, are ordered similarly to their respective homologs on the genomes of the rudiviruses SIRV1 and SIRV2.
Antigenic properties
No information available.
Biological properties
The host range is restricted to hyperthermophilic archaea of the genus Acidianus. Virions adsorb to the pili of the host cell, their adsorption to cell surface was never observed. The unusual termini of the virion appear to have a special function in the process of adsorption: the claw folds and keeps the virion firmly attached to a pilus.
During a cycle of productive infection neither formation of any significant amounts of cell debris, nor decrease in cell density were observed. Latent period is about 4 h. Infected host cell culture was not cured of the virus after several successive transfers into fresh medium and prolonged, continuous growth. Integration of viral genome into the host chromosome was not observed.
List of species demarcation criteria in the genus
Not applicable.
List of species in the genus Gammalipothrixvirus
| Acidianus filamentous virus 1 |
|
|
| Acidianus filamentous virus 1 | [AJ567472] | (AFV-1) |
Species names are in italic script; names of isolates are in roman script. Sequence accession numbers [ ] and assigned abbreviations ( ) are also listed.
Other related viruses which may be members of the genus Gammalipothrixvirus but have not been approved as species
None reported.
Genus Deltalipothrixvirus
Type species Acidianus filamentous virus 2
Distinguishing features
The virion carries exceptional structures at the termini, not observed in the virions of other members of the family. Moreover, the virion core does not exhibit any regular structure on its surface and, thereby, differs from both the nucleoprotein complex of the alphalipothrixvirus TTV1 and the nucleosome-like arrangement of the betalipothrixvirus virion cores.
Virion properties
Morphology
The virion is a flexuous filament, about 1100 nm in length and 24 nm in width (Figures 1, 8). It is enveloped with a 3–4 nm thick envelope. At the two ends of the virion identical structures are present, constituting a complex collar with two sets of filaments, resembling a bottle brush with a solid cap 17 nm in diameter on its top (Figure 8).
Physicochemical and physical properties
The buoyant density of the virions is about 1.3 g cm−3. Short treatment with detergents removes the envelope and uncovers a thinner filament about 17 nm in width.
Nucleic acid
The genome is linear dsDNA of 31.787 kbp. No information is available on modification of the ends of the molecule.
Proteins
The virion carries seven proteins with apparent molecular masses of 65, 50, 45, 40, 35, 26 and 6 kDa.
Lipids
Lipids could not be detected.
Carbohydrates
None reported.
Genome organization and replication
From 34 putative genes, eight have homologs in betalipotrixviruses and rudiviruses. The genome contains an unusual 1.008 kbp region close to the centre, which carries two large 46-bp direct repeats and multiple imperfect short repeats throughout the region (Figure 9). Its base composition is strongly biased, with one DNA strand containing only 7% guanosines. The region is bordered by short inverted repeats and may constitute a replication initiation site. In the central part of the genome is present also a tRNALys gene, and it contains a 12-bp archaeal intron. No information is available on genome replication.
Antigenic properties
No information available.
Biological properties
The host range is restricted to hyperthermophilic archaea of the genus Acidianus. The virus host is devoid of pilus-like structures and virions attach directly to the cell surface. The terminal structures of the virion appear to be involved in adsorption. No conformational changes could be detected in these structures upon adsorption.
During a cycle of productive infection neither formation of any significant amounts of cell debris, nor decrease in cell density was observed. Infected host cell culture was not cured of the virus after several successive transfers into fresh medium and prolonged, continuous cultivation. Integration of the viral genome into the host chromosome was not observed.
List of species demarcation criteria in the genus
Not applicable.
List of species in the genus Deltalipothrixvirus
| Acidianus filamentous virus 2 |
|
|
| Acidianus filamentous virus 2 | [AJ854042] | (AFV-2) |
Species names are in italic script; names of isolates are in roman script. Sequence accession numbers [ ] and assigned abbreviations ( ) are also listed.
Other related viruses which may be members of the genus Deltalipothrixvirus but have not been approved as species
Not applicable.
List of unassigned species in the family Lipothrixviridae
None reported.
Phylogenetic relationships within the family
The alphalipothrixvirus does not share any homologous gene with the members of the three other genera. There is limited similarity in gene content between beta-, gamma- and deltalipothrixviruses, and only a few ORFs are highly conserved in all of them. A dendrogram constructed by alignment of the sequence of ORF235 of the betalipothrixviruses and its homologs in the gamma- and deltalipothrixviruses reflects relationships within the family (Figure 10).
Similarity with other taxa
Originally, the two families of dsDNA viruses with linear genomes, the Rudiviridae and the Lipothrixviridae, were distinguished by differences in virion structure and this was later supported by comparative genomics. Nevertheless, a substantial fraction of orthologous genes, including some encoding glycosyl transferases and transcriptional regulators, are shared by the rudiviruses and lipothrixviruses. For example, of the 45 predicted genes of the rudivirus Sulfolobus islandicus rod-shaped virus 1, nine share orthologs with the lipothrixvirus Sulfolobus islandicus filamentous virus. Moreover, the structure of the major virion protein of the rudiviruses turned out to be identical to the structures of the two major virion proteins of the lipothrixvirus Acidianus filamentous virus 1. These observations indicate that the known linear dsDNA viruses, all of which infect hyperthermopilic members of the domain Archaea, share a common ancestor. One can suggest a sequence of evolutionary events in which the major virion protein of a “simpler” non-enveloped virion of the Rudiviridae has been duplicated and evolved so as to facilitate interactions with a hydrophobic envelope, producing the more complex virion of the Lipothrixviridae. Therefore, it has been suggested that the two the families Rudiviridae and Lipothrixviridae should be placed into a new order.
Derivation of names
Lipo: from Greek lipos, “fat”.
Thrix: from Greek thrix, “hair”.
Further reading
Arnold, H.P., Zillig, W., Ziese, U., Holz, I., Crosby, M., Utterback, T., Weidmann, J.F., Kristjanson, J.K., Klenk, H.P., Nelson, K.E. and Fraser, C.M. (2000). A novel lipothrixvirus, SIFV, of the extremely thermophilic crenarchaeon Sulfolobus. Virology, 267, 252-266.
Bettstetter, M., Peng, X., Rachel, R., Garrett, R.A. and Prangishvili, D. (2003). AFV1, a novel virus of the hyperthermophilic crenarchaeon Acidianus. Virology, 315, 68-79.
Bize, A., Peng, X., Prokofeva, M., MacLellan, K., Lucas, S., Forterre, P., Garrett, R.A., Bonch-Osmolovskaya, E.A. and Prangishvili, D. (2008). Viruses in acidic geothermal environments of the Kamchatka Peninsula. Res. Microbiol., 159, 358-366.
Goulet, A., Blangy, S., Redder, P., Prangishvili, D., Felisberto-Rodriguesa, C., Forterre, P. Campanacci, V. and Cambillau, C. (2009). Acidianus filamentous virus 1 coat proteins display a helical fold spanning the filamentous archaeal viruses lineage. Proc. Natl Acad. Sci. USA, 106, 21155-21160.
Häring M., Rachel, R., Vestergaard, G., Brügger, K., Rachel, R., Garrett, R.A. and Prangishvili (2005). Structure and genome organisation of AFV2, a novel archaeal virus with unusual terminal and core structures. J. Bacteriol., 187, 3855-3858.
Janekovic, D., Wunderl, S., Holz, I., Zillig, W., Gierl, A. and Neumann, H. (1983). TTV1, TTV3 and TTV3, a family of viruses of the extremely thermophilic, acidophilic sulfur-reducing archaeobacterium Thermoproteus tenax. Mol. Gen. Genet., 192, 39-45.
Prangishvili, D., Forterre, P. and Garrett, R.A. (2006). Viruses of the Archaea: a unifying view. Nat. Rev. Microbiol., 4, 837-848.
Prangishvili, D., Garrett, R.A. and Koonin, E.V. (2006). Evolutionary genomics of archaeal viruses: Unique viral genomes in the third domain of life. Virus Res., 117, 52-67.
Vestergaard, G., Aramayo, R., Basta, T., Häring, M., Peng, X., Brügger, K., Chen, L., Rachel, R., Biosset, N., Garrett, R.A. and Prangishvili, D. (2008). Structure of the Acidianus Filamentous Virus 3 and comparative genomics of related archaeal lipothrixviruses. J. Virol., 82, 371-381.
Zillig, W., Prangishvili, D., Schleper, C., Elferink, M., Holz, I., Albers, S., Janekovic, D. and Götz, D. (1996). Viruses, plasmids and other genetic elements of thermophilic and hyperthermophilic Archaea. FEMS Microbiol. Rev., 18, 225-236.
Contributed by
Prangishvili, D.
Figures
Figure 1 Negative contrast electron micrographs of virions of representatives of four genera of the family Lipothrixviridae. (Top, left) Thermoproteus tenax virus 1 from the genus Alphalipothrixvirus; (center; left) Sulfolobus islandicus filamentous virus from the genus Betalipothrixvirus; (top, right) Acidianus filamentous virus 1 from the genus Gammalipothrixvirus; (bottom) Acidianus filamentous virus 2 from the genus Deltalipothrixvirus. The bars represent 100 nm.
(Modified from Arnold et al., 2000; Bettstetter et al., 2003, Hring et al., 2005; Vestergaard et al. 2008.)
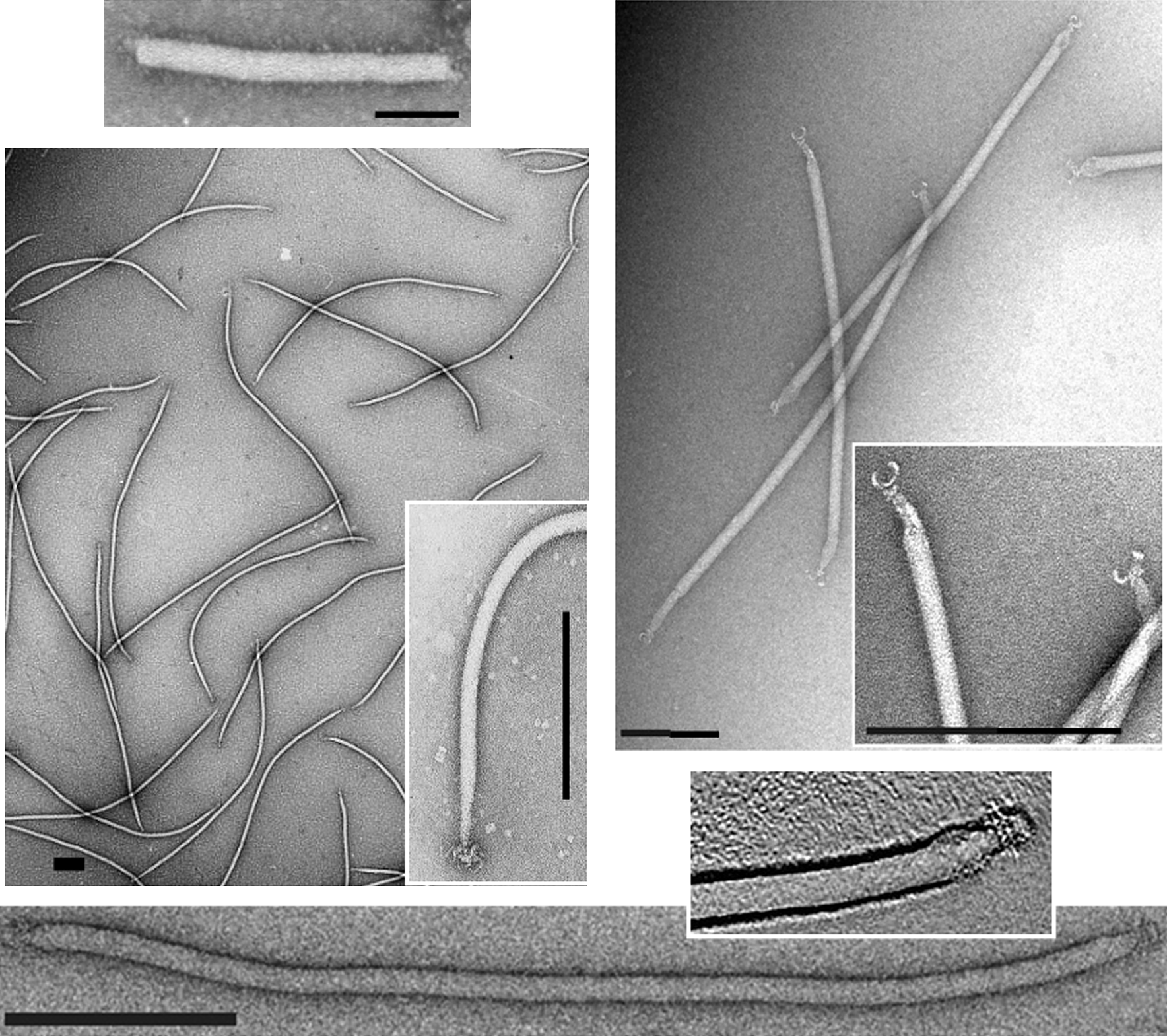

Figure 2 Negative contrast electron micrograph of intact and partially deteriorated virions of Thermoproteus tenax virus 1, exhibiting the envelope and the core. The bar represents 100 nm.
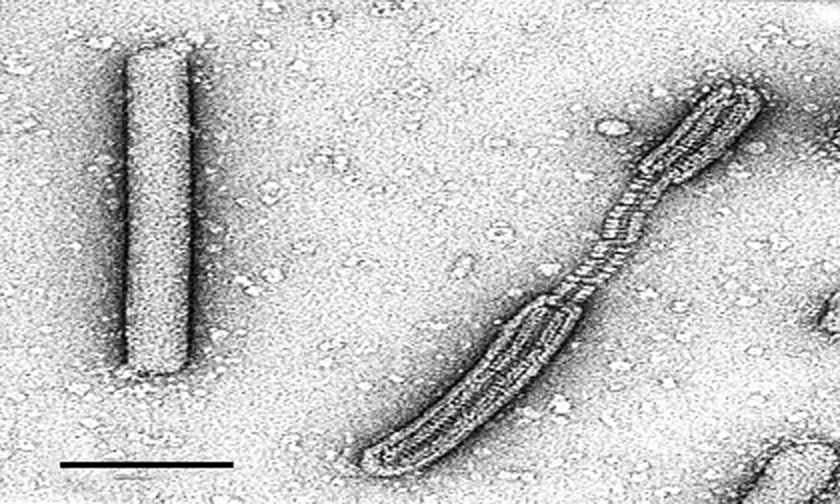
Figure 3 (Top) Negative contrast electron micrograph of AFV3 virion. The bar represents 100 nm. (Middle row) Image processing of AFV3 virion: (left) average 2D map computed from a set of the negatively stained sample, and (right) average 2D map computed from a set of images taken from cryo-EM micrographs; the contrast is inverted. The bars represent 5 nm. (Bottom) Schematic model of the virion of the Acidianus filamentous virus 3.
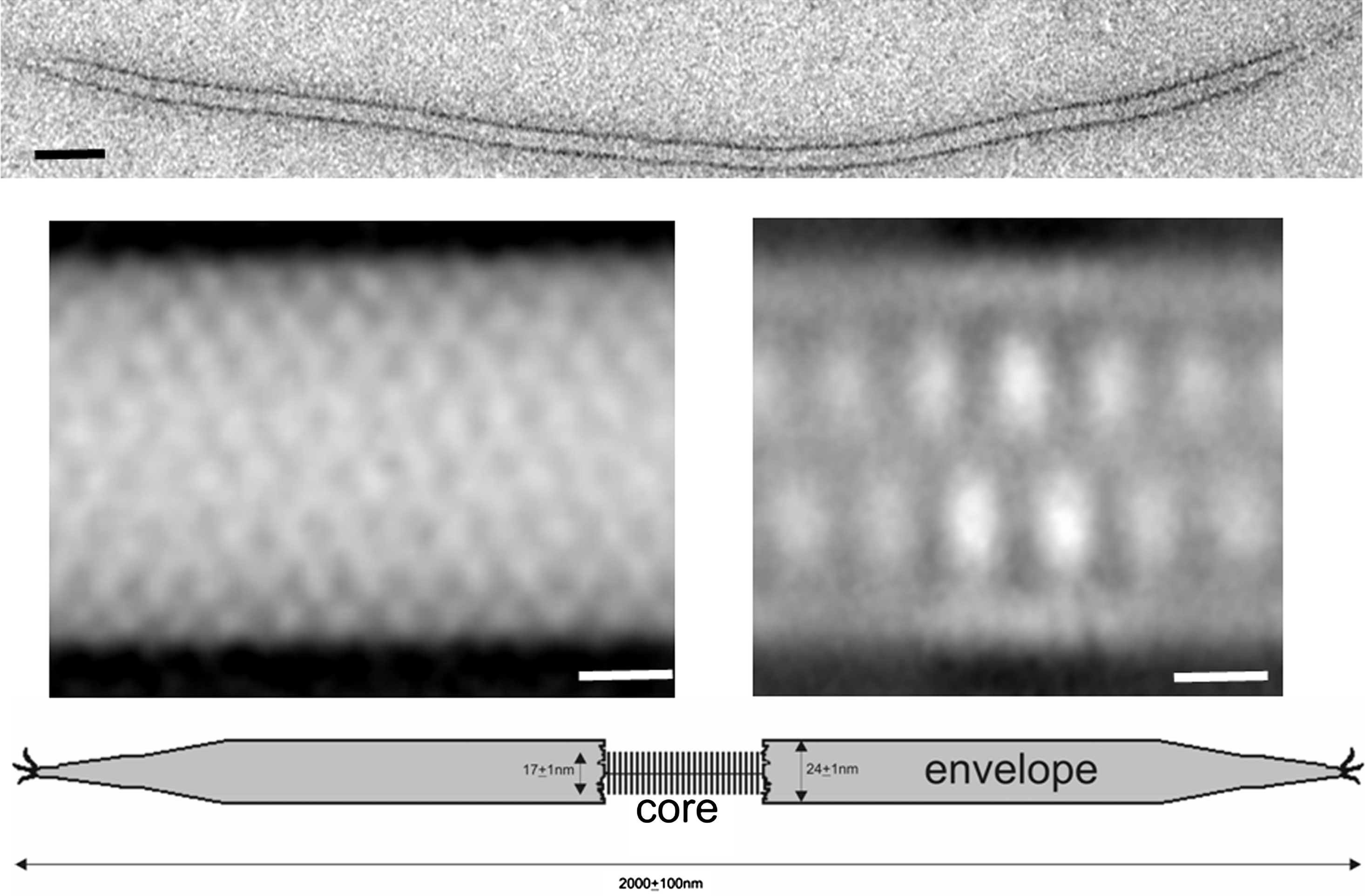

Figure 4 Genome maps aligned for the betalipothrixviruses, showing the relative size, location and orientation of the predicted genes. Homologous genes are coded with identical colors. Homologous operons carrying three or more genes are displayed as a consecutive set of genes with the same color. White boxes indicate that no homologous genes were detected in the other genomes. The ORFs of AFV-3 are labelled according to their amino acid lengths. Predicted protein functions are shown as follows: hc, helicase; gt, glycosyltransferase; mt, SAM-dependent methyltransferase; nc, nuclease; pp, protein phosphatase; sp, structural protein; and tr, transcriptional regulator.
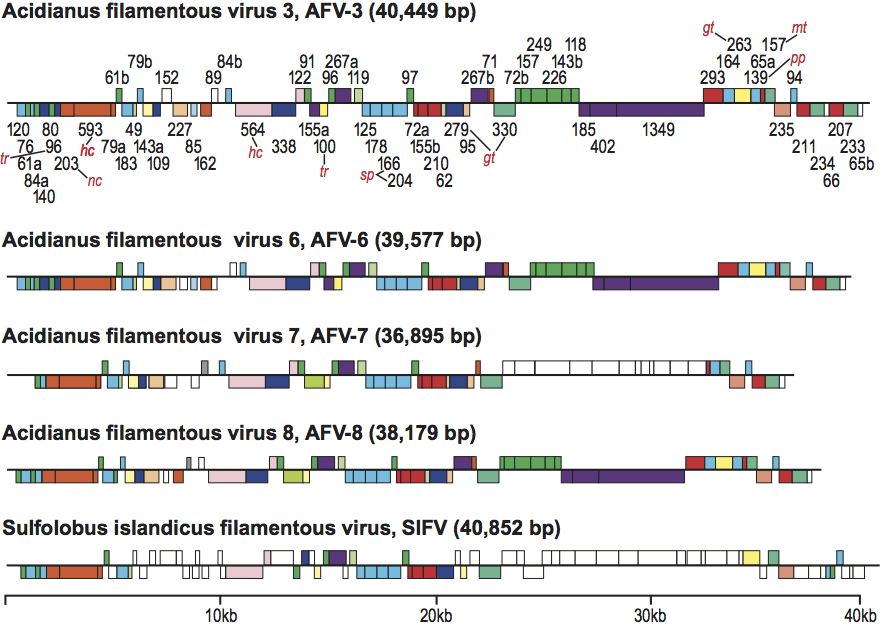
Figure 5 Schematic model of the virion of the Acidianus filamentous virus 1.
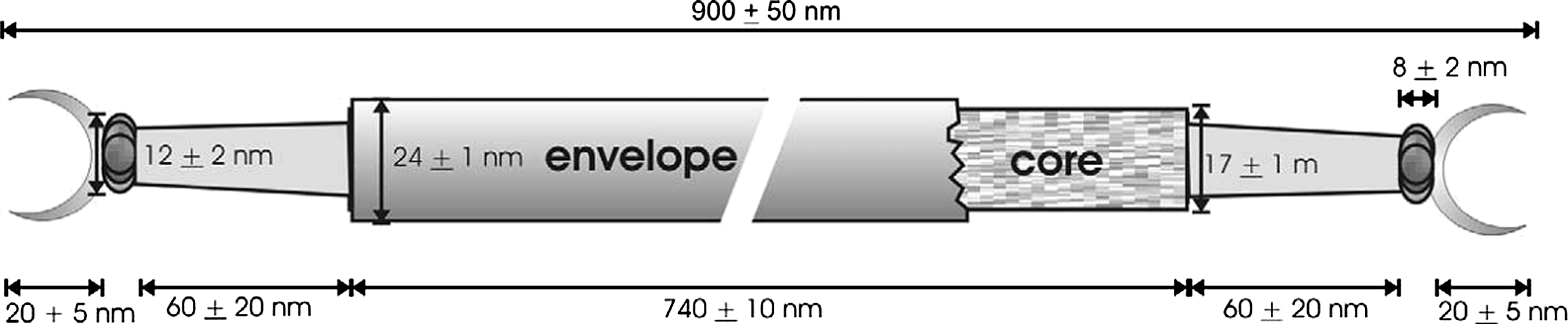
Figure 6 Topological model of the virion of Acidianus filamentous virus 1, indicating location of the two major structural proteins, P132 and P140. The P132 patch is represented by the blue surface in the center. The dsDNA backbone is orange and the P140 is represented by the grey surface. P140 contacts both the lipid-containing outer shell (in green) and the genomic DNA.
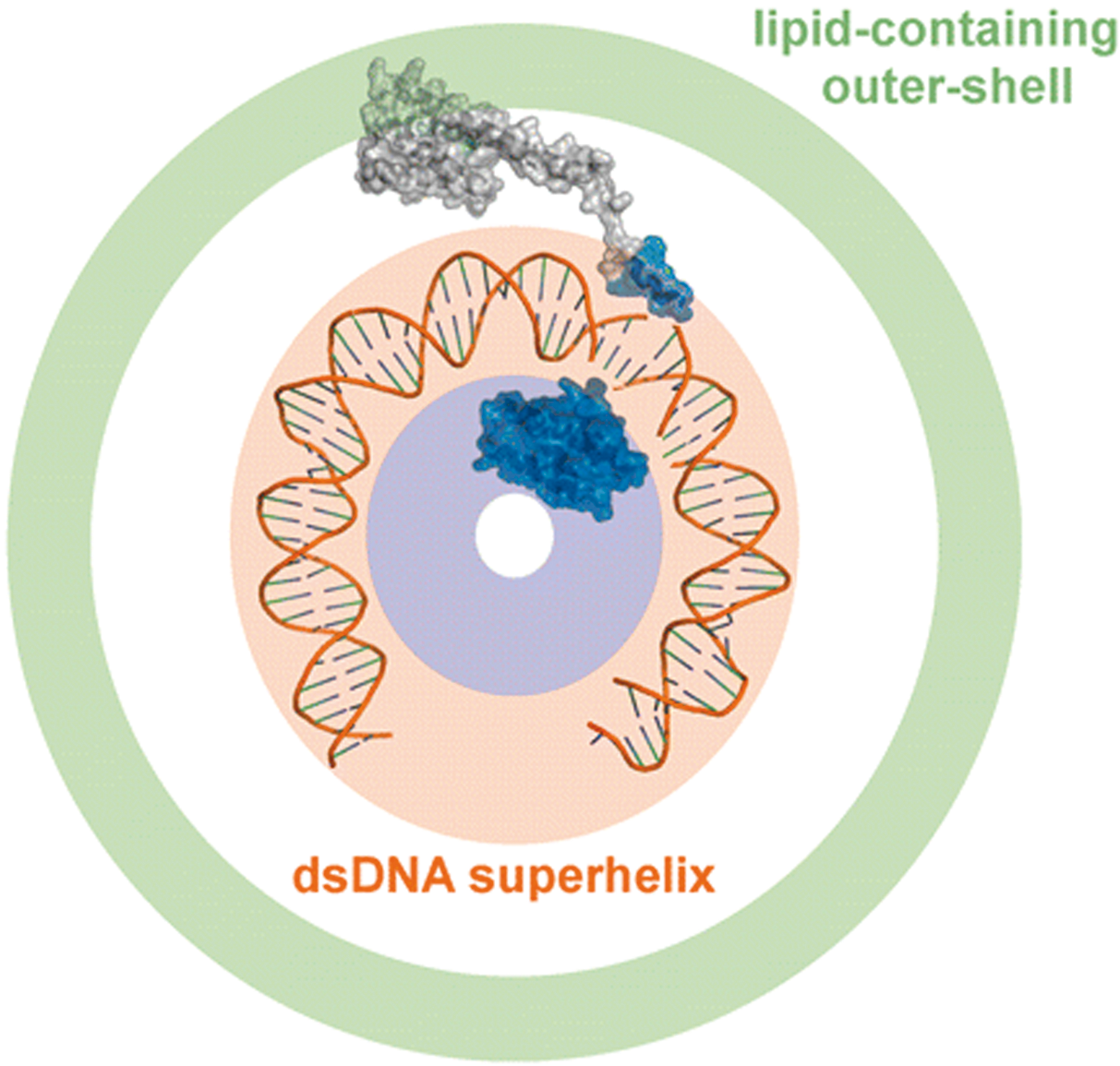
(From Goulet et al., 2009.)
Figure 7 Genome map of Acidianus filamentous virus 1, showing relative size, location, and orientations of the predicted genes. ORFs with significant sequence similarities to ORFs in other viruses are shaded and labeled.
(From Bettstetter et al., 2003.)
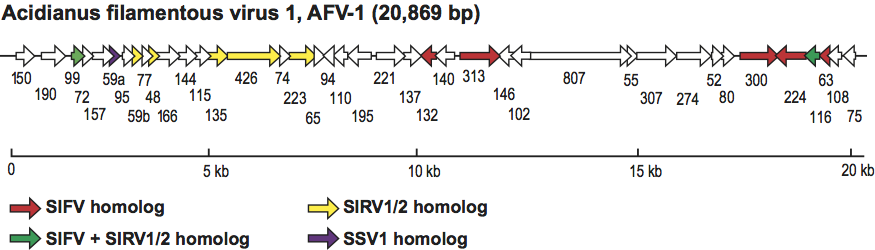
Figure 8 Schematic model of the virion of the Acidianus filamentous virus 2.
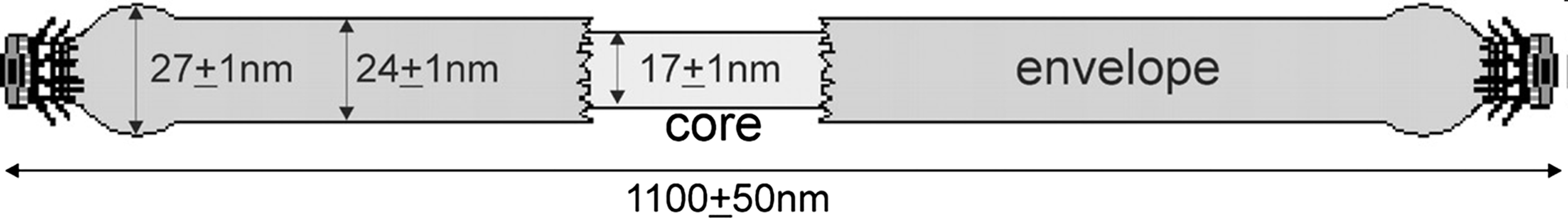
Figure 9 Genome map of Acidianus filamentous virus 2, showing relative size and location of the predicted genes. Genes are expressed from left to right in the upper row and from right to left in the lower row. ORFs shared with the beta- and gammalipothrixviruses, and with the rudiviruses, are represented by filled rectangles.
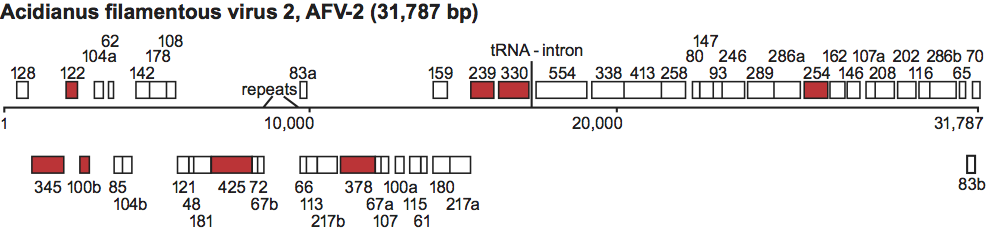
Figure 10 Dendrogram for genera Betalipothrixvirus, Gammalipothrixvirus, and Deltalipothrixvirus, derived from comparison of sequences of the only well-conserved putative gene shared by all members of the genera.
(Modified from Vestergaard et al., 2008.)