Family: Spiraviridae
David Prangishvili, Tomohiro Mochizuki and Mart Krupovic
The citation for this ICTV Report chapter is the summary published as Prangishvili et al., (2019):
ICTV Virus Taxonomy Profile: Spiraviridae, Journal of General Virology, 101, 240–241
Corresponding authors: David Prangishvili ([email protected]) and Mart Krupovic ([email protected])
Edited by: Andrew M. Kropinski and Stuart G. Siddell
Posted: December 2019
PDF: ICTV_Spiraviridae.pdf
Summary
The family Spiraviridae includes viruses that replicate in hyperthermophilic archaea from the genus Aeropyrum (Table 1. Spiraviridae). The non-enveloped, hollow, cylindrical virions are formed from a coiling fibre which consists of two intertwining halves of a single circular nucleoprotein filament. A short appendage protrudes from each end of the cylindrical virion. The genome is a circular, positive-sense single-stranded DNA of 24,893 nucleotides. The coil-shaped morphology represents one of the archaea-specific virion morphotypes.
Table 1. Spiraviridae. Characteristics of members of the family Spiraviridae
| Characteristic | Description |
| Typical member | Aeropyrum coil-shaped virus (HE681887), species Alphaspiravirus yamagawaense |
| Virion | Coil-shaped, 230±10 × 19±1 nm wide, with appendages of 20±2 nm at each end; non-enveloped |
| Genome | Circular, positive-sense single-stranded DNA of 24,893 nucleotides |
| Replication | Non-lytic, chronic infection |
| Translation | Not characterised |
| Host range | Hyperthermophilic archaea from the genus Aeropyrum |
| Taxonomy | Family Spiraviridae, genus Alphaspiravirus, single species |
Virion
Morphology
Virions of Aeropyrum coil-shaped virus (ACV) are non-enveloped, hollow cylindrical particles, 230±10 x 19±1 nm, formed by coiling of a nucleoprotein fiber as a helical spring (Figure 1. .Spiraviridae) (Mochizuki et al., 2012). The spring-forming nucleoprotein fiber itself has a helical structure being formed by two intertwining halves of circular single-stranded DNA molecule covered by capsid proteins (Figure 1. .Spiraviridae). An appendage of 20±2 nm protrudes from each end at 45° angles to the axis of the cylindrical virion (Figure 1. .Spiraviridae, inset). The architectural solution used by ACV to package its circular genome is unprecedented among viruses of bacteria and eukaryotes (King et al., 2012). The coil-shaped morphology represents a group of archaea-specific virion morphotypes (Prangishvili et al., 2017).
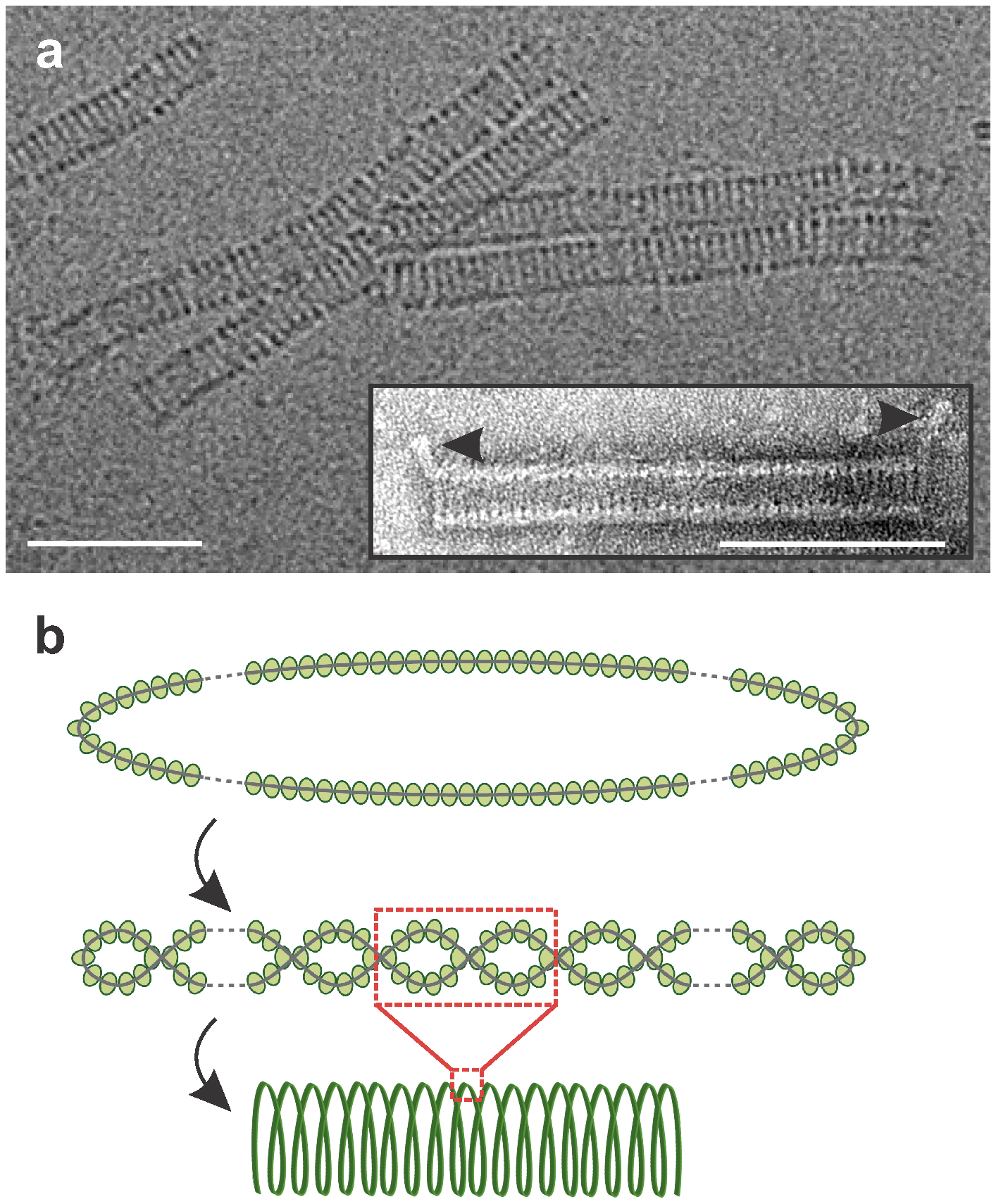 |
| Figure 1. .Spiraviridae. Virions of Aeropyrum coil-shaped virus. A. Electron micrographs of virions negatively-stained with uranyl acetate (in inset) and embedded in vitreous ice. Scale bars, 100 nm. B. Schematic representation of different levels of virion organization: the two halves of the circular nucleoprotein (top) intertwine with each other to form a filament (middle), which is condensed into the cylindrical helix of the virion (bottom). Modified with permission from (Mochizuki et al., 2012). |
Physicochemical and physical properties
The ACV virion buoyant density in caesium chloride is about 1.28 g cm−3 (Mochizuki et al., 2012). The flexible coil of the native virion is prone to contraction and stiffening upon dehydration due to uranyl acetate staining, as is revealed by differences in the size and appearance of virions embedded in vitreous ice and those stained with uranyl acetate (Figure 1. .Spiraviridae). The virions are fragile and often partially disassembled after high-speed ultracentrifugation (Mochizuki et al., 2012).
Nucleic acid
Virions contain a single molecule of circular single-stranded (ss) DNA of 24,893 nucleotides. The G+C content of the genome is 46.7% (Mochizuki et al., 2012).
Lipids
Not reported.
Proteins
The virions have two major proteins with molecular masses of about 23 and 18.5 kDa, and a few minor proteins with molecular masses of 5–13 kDa (Mochizuki et al., 2012). However, the identity of these proteins could not been determined by MALDI-TOF mass spectrometry analysis, potentially due to extensive posttranslational modification.
Genome organization and replication
The genome contains 57 predicted open reading frames (ORFs) larger than 40 codons that occupy 93.5 % of the genome (Figure 2. .Spiraviridae) (Mochizuki et al., 2012). All but one of these ORFs are present on the DNA strand that is packaged into the ACV virions, indicating that the genome is of positive-sense. The functional complexity of the ACV genome is far greater than that observed for any other known ssDNA virus. ACV encodes putative a trypsin-like serine protease, tyrosine recombinase, two thioredoxin-like proteins, proteins involved in carbohydrate metabolism, and several DNA-binding proteins. The virus does not encode any protein with significant sequence similarity with the known Rep proteins involved in rolling circle replication mechanism, operating in most of the known ssDNA viruses (Krupovic et al., 2018, Kazlauskas et al., 2019). Thus, ACV might employ a novel mechanism of genome replication, which is likely to depend on the host machinery.
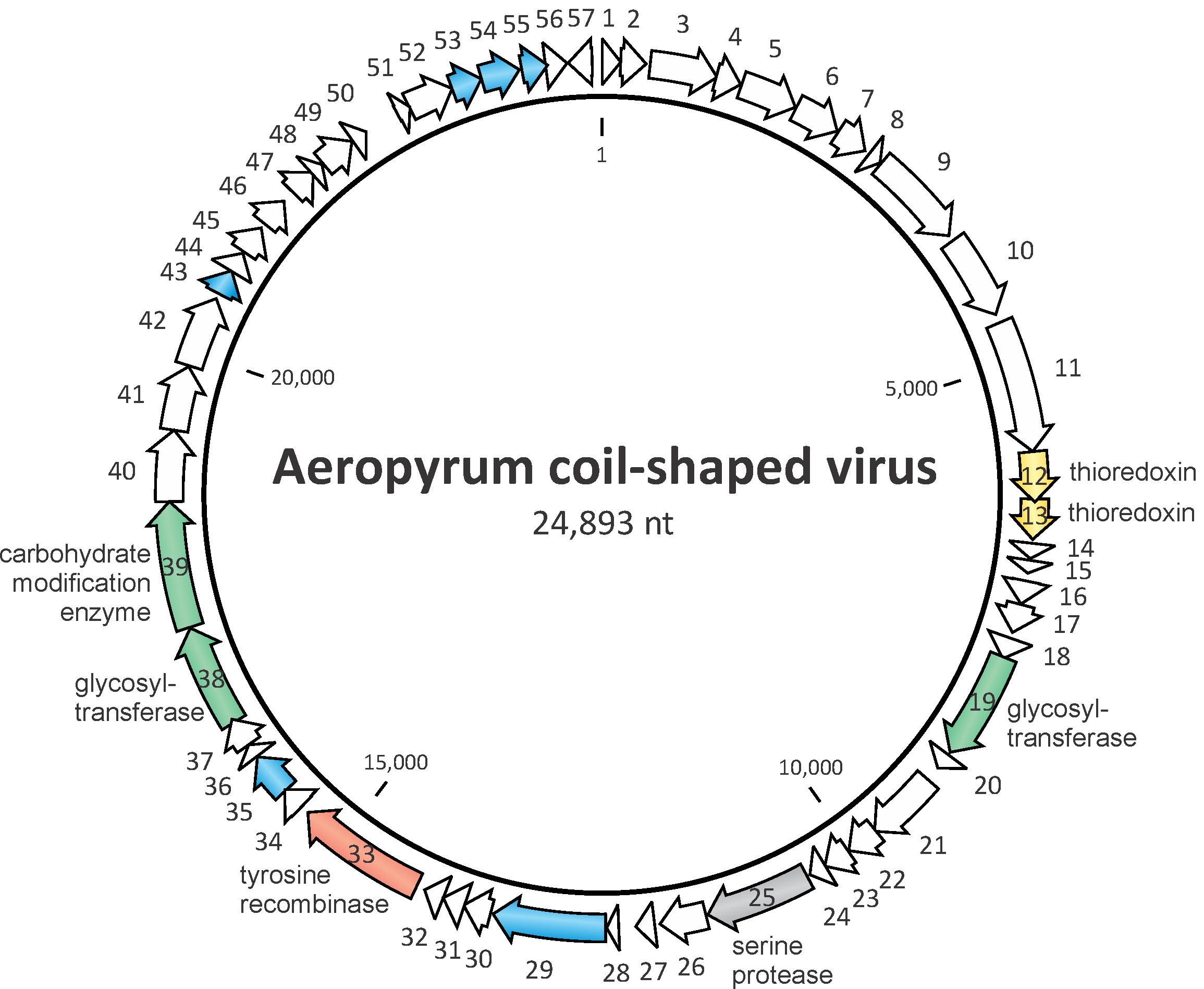 |
| Figure 2. Spiraviridae. Circular genome map of Aeropyrum coil-shaped virus. The open reading frames (ORFs) are marked with arrows indicating the direction of transcription. The ORFs encoding putative proteins for which functions could be predicted are color-coded and labeled on the figure; blue arrows represent ORFs encoding DNA-binding proteins. |
Biology
Aeropyrum coil-shaped virus was isolated from a coastal hot spring with temperatures reaching 100°C at Yamagawa, Kyushu island, Japan (Mochizuki et al., 2012, Mochizuki et al., 2010). The virus host Aeropyrum permix, is the most thermophilic species among known aerobic organisms, growing optimally at 90–95°C. ACV virions are released without apparent host cell lysis. The virus does not encode identifiable DNA and RNA polymerases and is likely to depend on the host machineries for genome replication and transcription.
Derivation of names
Spira: From Latin spira, for “coil”.
Aeropyrum coil-shaped virus: genus of archaeal host Aeropyrum and shape of the virion
Relationships within the family
ACV represents the sole known species of the family.
Relationships with other taxa
The family Spiraviridae comprises a single genus Alphaspiravirus, with one species. No relationships with other viruses have been revealed (Krupovic et al., 2018, Iranzo et al., 2016). Spiraviruses, along with other archaeal viruses, may represent ancestral virus forms no longer observed amongst extant prokaryotic or eukaryotic viruses (Prangishvili 2015).

