Family: Solinviviridae
Katherine Brown, Ingrida Olendraite, Steven M. Valles, Andrew E. Firth, Yanping Chen, Diego M. A. Guérin, Yoshifumi Hashimoto, Salvador Herrero, Joachim R. de Miranda and Eugene Ryabov
Corresponding author: Andrew E. Firth ([email protected])
Edited by: Nick J. Knowles and Sead Sabanadzovic
Posted: February 2019
PDF: ICTV_Solinviviridae.pdf
Summary
Solinviviridae is a family of picorna/calici-like viruses with non-segmented, linear, positive-sense RNA genomes of approximately 10–11 kb (Table 1. Solinviviridae). Unusually, their capsid proteins are encoded towards the 3′-end of the genome and comprise a single jelly-roll fold domain with a large extension domain, plus an additional capsid-associated protein. A subgenomic capsid-encoding RNA is produced, but the capsid proteins can normally also be expressed from the genomic RNA as an extension of the replication (picorna-like helicase-protease-polymerase) polyprotein. Members of two species within the family infect ants but related unclassified virus sequences derive from a large variety of insects and other arthropods.
Table 1. Solinviviridae. Characteristics of members of the family Solinviviridae.
| Characteristic | Description |
| Typical member | Solenopsis invicta virus 3, DM (FJ528584), species Invictavirus solenopsis |
| Virion | Non-enveloped, 26–30 nm diameter with apparent projections |
| Genome | 10–11 kb of positive-sense, non-segmented RNA |
| Replication | Not studied; presumed to be similar to other picorna/calici-like viruses |
| Translation | From genomic (replication and capsid proteins) and subgenomic (capsid proteins) RNAs |
| Host range | Arthropoda |
| Taxonomy | Realm Riboviria, kingdom Orthornavirae, phylum Pisuviricota, class Pisoniviricetes, order Picornavirales; includes the genera Invictavirus and Nyfulvavirus, each with one species |
Virion
Morphology
Particles are icosahedral with a diameter of 26–30 nm and apparent projections (Figure 1. Solinviviridae). Similar morphologies have been noted in the related unclassified viruses, kelp fly virus (Scotti et al., 1976, Hartley et al., 2005) and Riptortus pedestris virus 1 (Yang et al., 2016), with the projections in kelp fly virus - present at some but not all five-fold axes - protruding approximately 6 nm above the surface of the 33 nm diameter virion. Particles are presumed to be T=3 containing 180 copies of viral protein 1 (VP1) (Valles et al., 2014a).
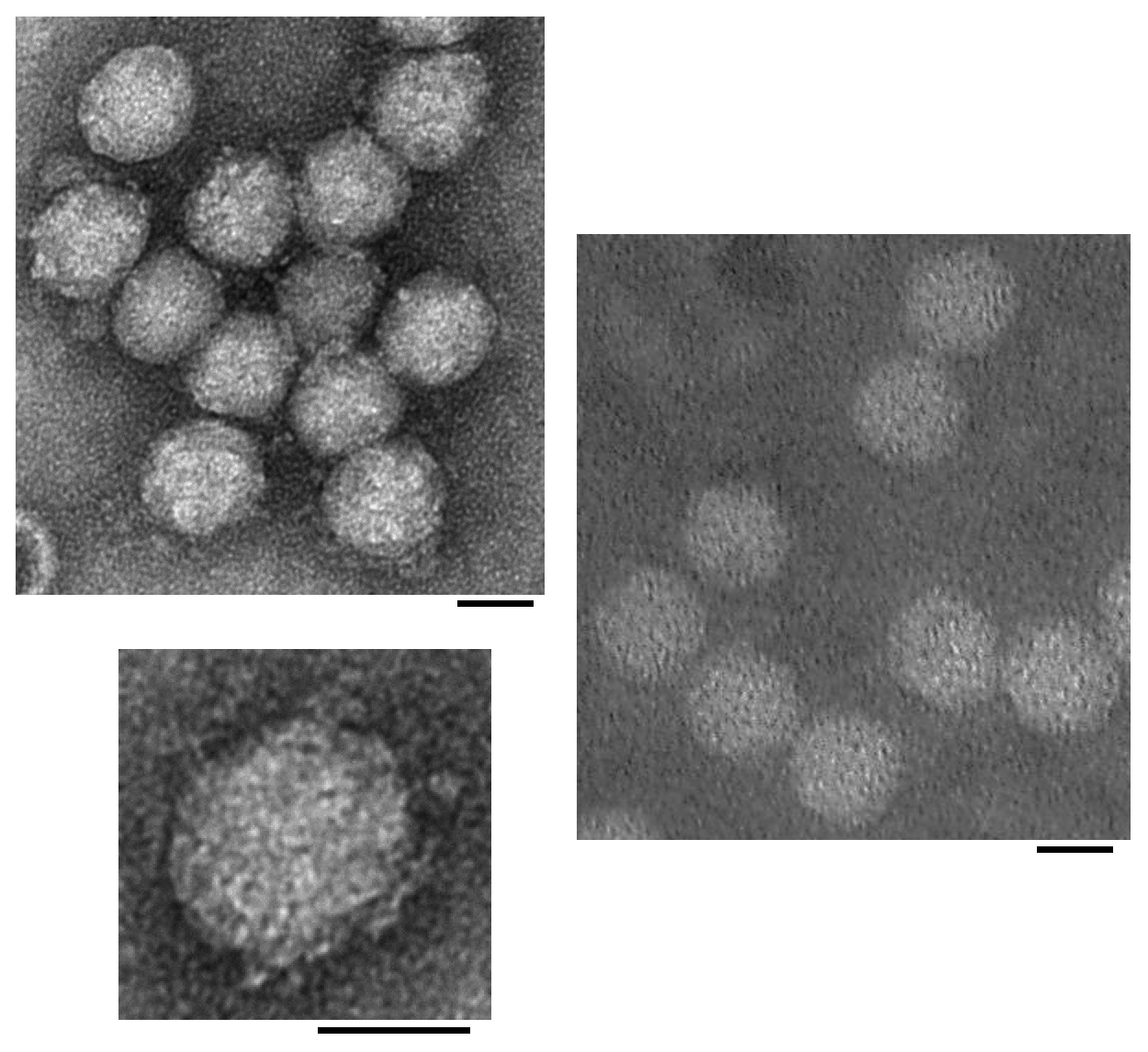 |
| Figure 1. .Solinviviridae. Transmission electron micrographs of negatively-stained Solenopsis invicta virus 3 (left and inset below) and Nylanderia fulva virus 1 (right) particles (scale bars = 20 nm) [reproduced from (Valles and Hashimoto 2009, Valles et al., 2016); U.S. government material as public domain content]. |
Physicochemical and physical properties
Buoyant density in CsCl has been measured at 1.39±0.02 g ml−1 for Solenopsis invicta virus 3 (SINV3) (Valles and Hashimoto 2009) and 1.287±0.012 g ml−1 for Nylanderia fulva virus 1 (NfV1) (Valles et al., 2016). A high buoyant density (1.425±0.002 g ml−1 in neutral CsCl) has also been noted for the related unclassified virus, kelp fly virus (Scotti et al., 1976). Kelp fly virus is reported to be unusually heat stable, surviving for 10 min at 90º C but not at 100º C (Scotti et al., 1976).
Nucleic acid
The genome comprises a single positive-sense RNA molecule of 10–11 kb. The genome has a 3′-poly(A) tail, and is presumed to have a viral protein genome-linked (VPg) covalently attached to the 5′-end. Related unclassified viruses have genomes in the range 9–12 kb.
Proteins
SINV3 virions contain a major component viral protein 1 (VP1) with a jelly-roll fold, and a minor component VP1-FSD where FSD (frame shift domain) is an extension domain that is appended to a proportion of the VP1 proteins as a result of programmed ribosomal frameshifting (Valles et al., 2014a). VP1 is thought to form the virion shell and FSD the projections (Valles et al., 2014a). The ratio of VP1 to VP1-FSD in virions has been estimated at around 1:5 in SINV3 (Valles et al., 2014a) and kelp fly virus (Scotti et al., 1976), and between 1:5 and 1:10 in Acyrthosiphon pisum virus (van den Heuvel et al., 1997). A further virion-associated protein, VP2, is encoded downstream of FSD from which it is presumed to be proteolytically cleaved. As VP2 has been detected only at low levels in SINV3 virions, it is not clear yet whether this protein is lost during virion purification or only a fraction of expressed VP2 associates with virions. In NfV1, there is no frameshift separating the VP1 and FSD open reading frames (ORFs), and so all of VP1 is expected to be produced in the form VP1-FSD, although it is possible that this is subsequently cleaved (by analogy to kelp fly virus where VP1 and FSD are also encoded in-frame yet most VP1 in the virion appears to lack the FSD domain). Related, but yet unclassified virus sequences exhibit a mixture of the two genome organizations i.e. with FSD-VP2 encoded in a frameshift ORF (SINV3-like) or with FSD-VP2 encoded in-frame with VP1 (NfV1-like) (Valles et al., 2016).
Lipids
None reported.
Carbohydrates
None reported.
Genome organization and replication
The genome of SINV3 (genus Invictavirus) contains a long 5′-proximal ORF covering approximately three quarters of the genome, encoding superfamily III helicase (Hel), 3C-like chymotrypsin-related cysteine protease (Pro), and superfamily I RNA-dependent RNA polymerase (RdRP) domains followed by the jelly-roll fold capsid protein (VP1) (Figure 2. Solinviviridae). Similar to other picorna/calici-like viruses, the ORF1 polyprotein is presumably cleaved by the virus 3C-like protease. Cleavage sites have been predicted (Valles et al., 2016) but not yet confirmed empirically. A dsRNA binding domain with homology to the 1A suppressor of silencing protein of Drosophila C virus (family Dicistroviridae) is present between the RdRP and jelly-roll fold (JR) domains (Valles et al., 2014a). Additional proteins are likely encoded upstream of Hel (one protein) and between Hel and Pro (two proteins, one presumed to be a VPg). ORF2 appears to be expressed via ribosomal frameshifting leading to an ORF1-ORF2 polyprotein. The 5′- and 3′-halves of ORF2 encode FSD (frameshift domain) and VP2, respectively (see above). A subgenomic RNA encoding the dsRNA-binding protein, VP1, FSD and VP2 is produced during virus infection (Valles et al., 2014a). NfV1 (genus Nyfulvavirus) has a single long ORF encoding similar domains to the SINV3 ORF1-ORF2 fusion and in the same order (Valles et al., 2016). In NfV1, the region upstream of helicase has homology to ovarian tumor (OTU) domain. Related, but yet unassigned virus sequences have similar Hel-Pro-RdRP-JR domain organization, normally with either a single long ORF or a second ORF downstream of JR accessible via ribosomal frameshifting (Valles et al., 2016, van der Wilk et al., 1997).
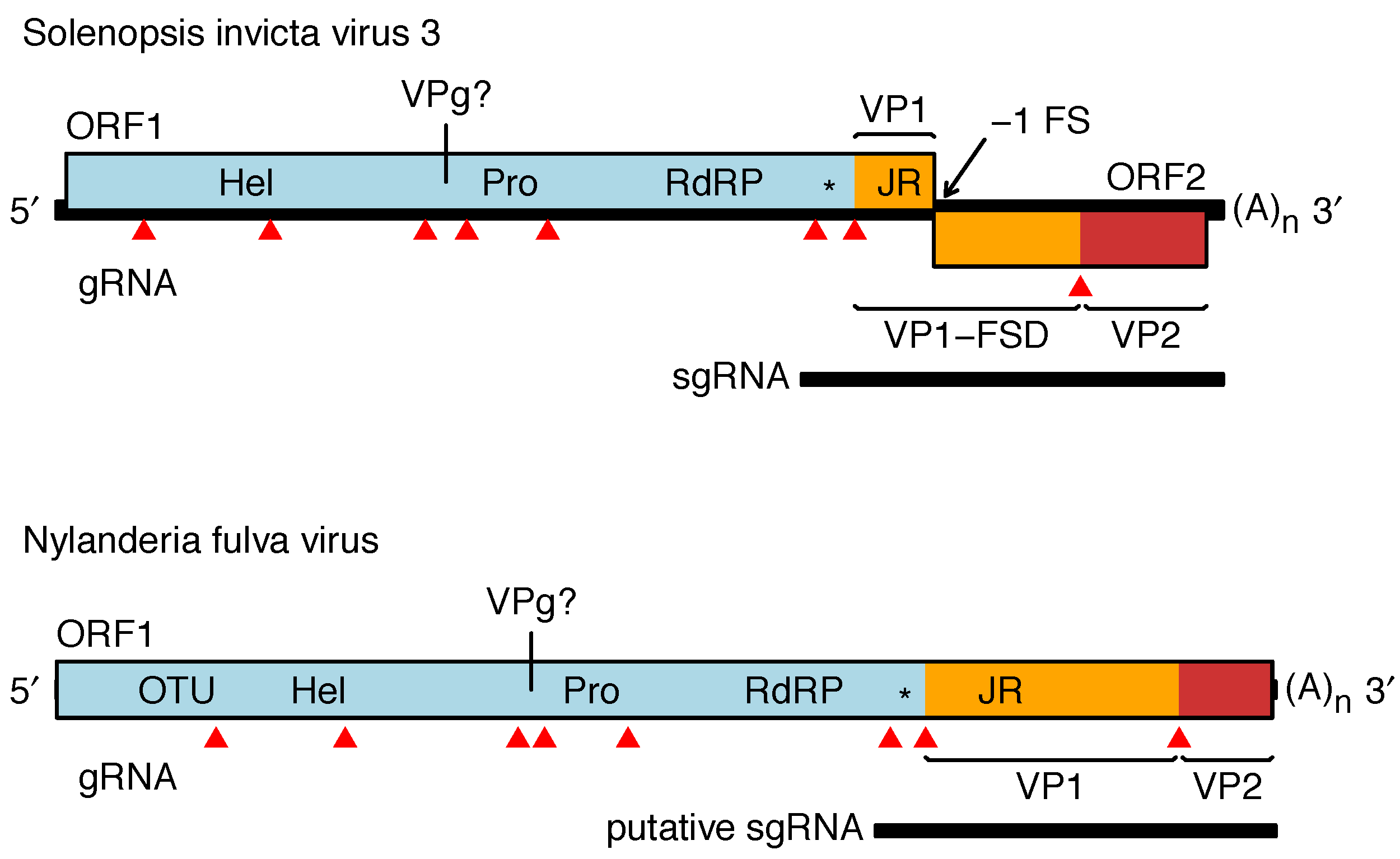 |
| Figure 2. .Solinviviridae. Genome organizations of Nylanderia fulva virus 1 and Solenopsis invicta virus 3. Predicted protein domains are indicated as follows: Hel - helicase, Pro - protease, RdRP - RNA-dependent RNA polymerase, JR - jelly-roll fold capsid, * - dsRNA binding protein (potentially a suppressor of host RNAi), OTU - ovarian tumor domain. Mapped virion proteins in Solenopsis invicta virus 3 are coloured as follows VP1 and VP1-FSD - orange, VP2 - red (VP = viral protein, FSD = frameshift domain). Predicted cleavage sites are indicated by red triangles. The genome is polyadenylated and believed to have a viral protein genome-linked (VPg) covalently attached to the 5′-end. |
Biology
Both SINV3 and NfV1 infect ants. Both viruses appear to have high host specificity and are easily transmitted in the lab by feeding (Valles et al., 2016, Porter et al., 2013). SINV3 infection is largely specific to workers and is highly pathogenic, leading to cessation of solid food feeding, significant mortality of adults and brood, and weight loss in queens (Valles et al., 2014b). NfV1 has been detected in larvae, pupae, worker and queen ants, with the highest viral load and active replication detected in larvae (Valles et al., 2016). NfV1 has not been detected in eggs, indicating absence of vertical transmission. Pathology has not been fully determined though it appears less pathogenic than SINV3 (Valles et al., 2016). Related unclassified viruses and virus-derived sequences have been obtained from a variety of insects and other arthropods (Shi et al., 2016) and have been reported to infect epithelial cells of the digestive tract and retard growth in pea aphids (van den Heuvel et al., 1997), to possibly reduce fecundity in rosy apple aphids (Ryabov et al., 2009), and to cause systemic infection in bean bugs (Yang et al., 2016).
Derivation of names
Solinvi: from the type species Solenopsis invicta virus 3.
Genus demarcation criteria
Genera are separated based on phylogenetic clustering based on protein sequence identities. Members of the genera Invictavirus and Nyfulvavirus have different genome organisations. However formal genus demarcation criteria have not yet been established.
Relationships within the family
Phylogenetic analysis of RdRP amino acid sequences indicates that members of the two classified solinvivirus species form part of a large and very diverse group of arthropod-infecting viruses within the picorna/calici-like group of viruses (Figure 3. Solinviviridae). The taxonomy of this larger group is currently uncertain. Analysis of genome structure using HHpred (Zimmermann et al., 2018) reveals a common Hel-Pro-RdRP-JR domain organization (with a few exceptions where the JR capsid cannot be detected and may have been replaced with unrelated capsid proteins). However, monophyly of the group within the larger picorna/calici-like group is not completely certain (Valles et al., 2016).
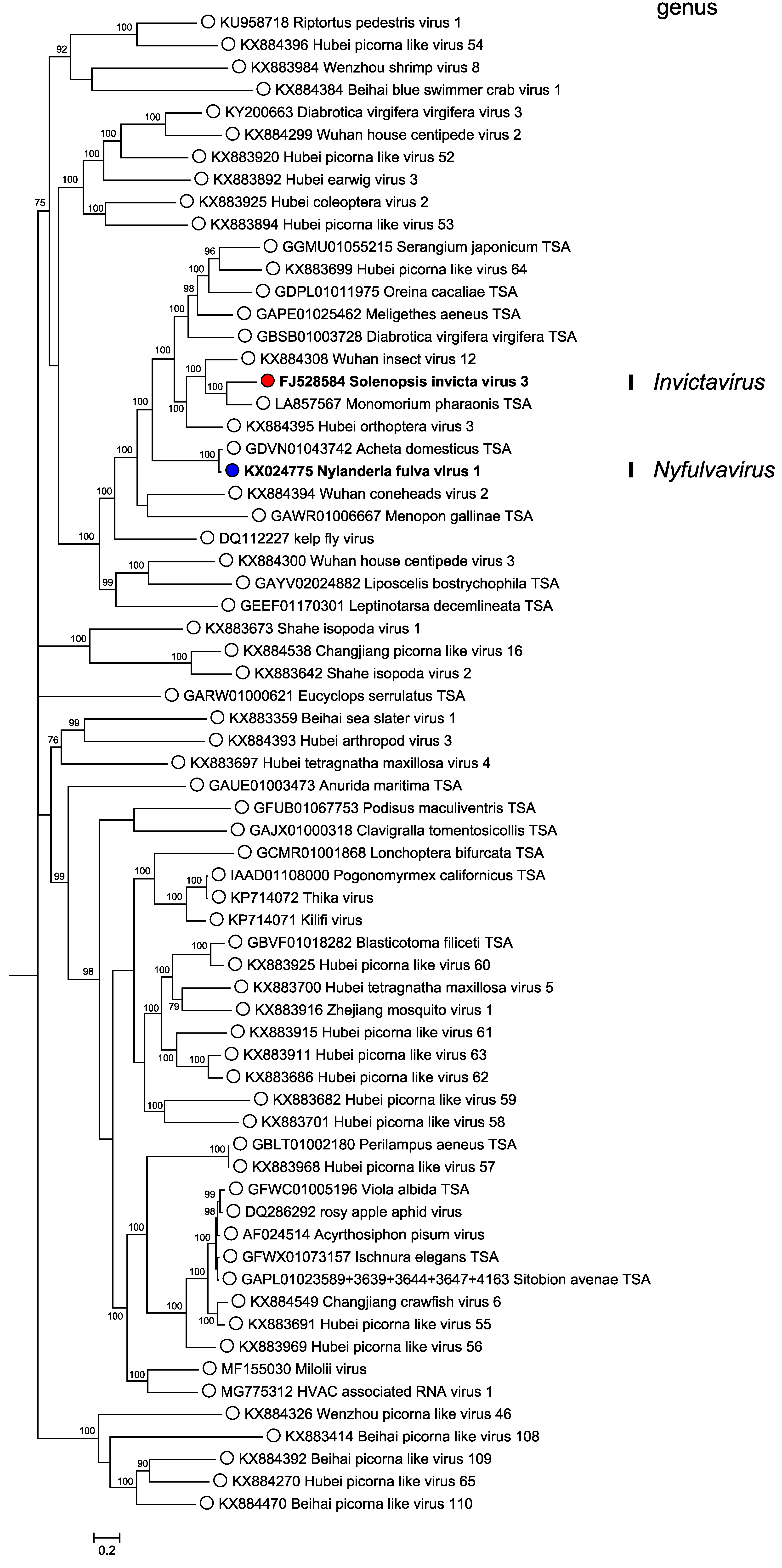 |
| Figure 3. . Solinviviridae. Mid-point rooted phylogenetic tree of soliniviruses (bold) and related unclassified virus sequences. RdRP amino acid sequences were obtained as reported previously (Valles et al., 2016), with additional sequences cropped to represent the same region. Sequences were aligned using MAFFT v7.310 (Katoh et al., 2002) with the L-INS-i algorithm and 1000 iterations. A Bayesian Markov chain Monte Carlo-based phylogenetic tree was generated using MrBayes v3.2.6 (Ronquist and Huelsenbeck 2003) with one million generations, discarding the first 25% as burn-in, with a ‘mixed’ amino acid rate matrix (averaged across ten models) and inverse gamma rate distribution. Posterior probabilities are indicated at nodes. Genome structure for all sequences was confirmed using HHpred (Zimmermann et al., 2018). This phylogenetic tree and corresponding sequence alignment are available to download from the Resources page. |
Relationships with other taxa
Genomes encode picorna/calici-like superfamily III helicase, 3C-like chymotrypsin-related cysteine protease, and superfamily I RdRP domains (Valles et al., 2014a, Valles et al., 2016). In contrast to members of the order Picornavirales, but similar to members of the family Caliciviridae, there is only a single jelly-roll fold capsid domain. Encoding of capsid proteins at the 3′-end of the genome, production of a capsid-encoding subgenomic RNA, and presence of a capsid projection domain are also calicivirus-like features. It is possible that Solinviviridae forms a sister group to Caliciviridae, though phylogenetic clustering is inconclusive due to difficulty in resolving tree topologies at this depth (Valles et al., 2014a).
Related, unclassified viruses
| Virus name | Accession number | Virus abbreviation |
| Acyrthosiphon pisum virus | AF024514 | APV |
| Perilampus aeneus TSA* | GBLT01002180 | |
| kelp fly virus | DQ112227 | KFV |
| rosy apple aphid virus | DQ286292 | RAAV |
| Clavigralla tomentosicollis TSA | GAJX01000318 | |
| Meligethes aeneus TSA | GAPE01025462 | |
| Sitobion avenae TSA | GAPL01023589; GAPL01023639; GAPL01023644; GAPL01023647; GAPL01024163 | |
| Eucyclops serrulatus TSA | GARW01000621 | |
| Anurida maritima TSA | GAUE01003473 | |
| Menopon gallinae TSA | GAWR01006667 | |
| Liposcelis bostrychophila TSA | GAYV02024882 | |
| Diabrotica virgifera virgifera TSA | GBSB01003728 | |
| Blasticotoma filiceti TSA | GBVF01018282 | |
| Lonchoptera bifurcata TSA | GCMR01001868 | |
| Oreina cacaliae TSA | GDPL01011975 | |
| Acheta domesticus TSA | GDVN01043742 | |
| Leptinotarsa decemlineata TSA | GEEF01170301 | |
| Podisus maculiventris TSA | GFUB01067753 | |
| Viola albida TSA | GFWC01005196 | |
| Ischnura elegans TSA | GFWX01073157 | |
| Serangium japonicum TSA | GGMU01055215 | |
| Pogonomyrmex californicus TSA | IAAD01108000 | |
| Kilifi virus | KP714071 | |
| Thika virus | KP714072 | |
| Riptortus pedestris virus 1 | KU958718 | RiPV1 |
| Beihai sea slater virus 1 | KX883359 | |
| Beihai picorna like virus 108 | KX883414 | |
| Shahe isopoda virus 2 | KX883642 | |
| Shahe isopoda virus 1 | KX883673 | |
| Hubei picorna like virus 59 | KX883682 | |
| Hubei picorna like virus 62 | KX883686 | |
| Hubei picorna like virus 55 | KX883691 | |
| Hubei tetragnatha maxillosa virus 4 | KX883697 | |
| Hubei picorna like virus 64 | KX883699 | |
| Hubei tetragnatha maxillosa virus 5 | KX883700 | |
| Hubei picorna like virus 58 | KX883701 | |
| Hubei earwig virus 3 | KX883892 | |
| Hubei picorna like virus 53 | KX883894 | |
| Hubei picorna like virus 63 | KX883911 | |
| Hubei picorna like virus 61 | KX883915 | |
| Zhejiang mosquito virus 1 | KX883916 | |
| Hubei picorna like virus 52 | KX883920 | |
| Hubei coleoptera virus 2 | KX883922 | |
| Hubei picorna like virus 60 | KX883925 | |
| Hubei picorna like virus 57 | KX883968 | |
| Hubei picorna like virus 56 | KX883969 | |
| Wenzhou shrimp virus 8 | KX883984 | |
| Hubei picorna like virus 65 | KX884270 | |
| Wuhan house centipede virus 2 | KX884299 | |
| Wuhan house centipede virus 3 | KX884300 | |
| Wuhan insect virus 12 | KX884308 | |
| Wenzhou picorna like virus 46 | KX884326 | |
| Beihai blue swimmer crab virus 1 | KX884384 | |
| Beihai picorna like virus 109 | KX884392 | |
| Hubei arthropod virus 3 | KX884393 | |
| Wuhan coneheads virus 2 | KX884394 | |
| Hubei orthoptera virus 3 | KX884395 | |
| Hubei picorna like virus 54 | KX884396 | |
| Beihai picorna like virus 110 | KX884470 | |
| Changjiang picorna like virus 16 | KX884538 | |
| Changjiang crawfish virus 6 | KX884549 | |
| Diabrotica virgifera virgifera virus 3 | KY200663 | DvvV3 |
| Monomorium pharaonis TSA | LA857567 | |
| Milolii virus | MF155030 | |
| HVAC associated RNA virus 1 | MG775312 |
Virus names and virus abbreviations are not official ICTV designations.
*TSA: Transcriptome shotgun assembly

