Family: Pneumoviridae
Bert Rima, Peter Collins, Andrew Easton, Ron Fouchier, Gael Kurath, Robert A. Lamb, Benhur Lee, Andrea Maisner, Paul Rota and Linfa Wang
The citation for this ICTV Report chapter is the summary published as Rima et al., (2017):
ICTV Virus Taxonomy Profile: Pneumoviridae, Journal of General Virology, 98, 2912–2913.
Corresponding author: Bert Rima ([email protected])
Edited by: Stuart G. Siddell and Peter Simmonds
Posted: December 2017, updated June 2023
PDF: ICTV_Pneumoviridae.pdf
Summary
Pneumoviridae is a family of large enveloped negative-strand RNA viruses (Table 1.Pneumoviridae). This taxon formerly was a subfamily within the family Paramyxoviridae, but was reclassified as a separate family in 2016. Members of the genus Orthopneumovirus infect mammals while members of the genus Metapneumovirus are specific for mammals or birds. Some viruses are specific and pathogenic for humans such as human respiratory syncytial virus and human metapneumovirus. There are no known vectors for these viruses and transmission is thought to be primarily by aerosol droplets and contact.
Table 1.Pneumoviridae. Characteristics of members of the family Pneumoviridae.
| Characteristic | Description |
| Example | human respiratory syncytial virus-A2 (M74568), species Orthopneumovirus hominis, genus Orthopneumovirus |
| Virion | Enveloped, both spherical and filamentous virions with a helical ribonucleoprotein (RNP) core. |
| Genome | Negative-sense unsegmented RNA genomes; from 13.2 to 15.3 kb |
| Replication | Cytoplasmic. The RNA-directed RNA polymerase comprises the phospho- (P) and large (L) proteins; M2-1 protein is a processivity factor. |
| Translation | The cellular translation machinery translates the capped and poly-adenylated messenger RNAs in the cytoplasm. |
| Host range | Mammals (both genera); birds (avian metapneumovirus) |
| Taxonomy | Realm Riboviria, kingdom Orthornavirae, phylum Negarnaviricota, subphylum Haploviricotina, class Monjiviricetes, order Mononegavirales; two genera, Orthopneumovirus and Metapneumovirus, which include 3 and 2 species, respectively. |
Virion
Morphology
Taking human respiratory syncytial virus, strain A2 (HRSV-A2) as a representative isolate, virions consist of spherical and filamentous forms and can have considerable variation in size and shape. Filamentous forms usually predominate and most often are 70–190 nm in diameter and up to 2 µm in length. Spherical forms are reported to be 80–140 nm in diameter (which may be cross-sections of filaments), but there also can be larger forms of 250–600 nm. Virions consist of a lipid envelope surrounding a nucleocapsid (Figure 1. Pneumoviridae). The envelope is derived directly from the host cell plasma membrane by budding and contains three transmembrane viral glycoproteins. These are present as homo-oligomers forming spike-like projections, 10–14 nm in length, spaced 8–11 nm apart for HRSV and 13–17 nm in length for metapneumoviruses. Underneath the lipid envelope is a layer of a non-glycosylated membrane or matrix protein (M), and underneath that is a second layer composed of a second non-glycosylated membrane or matrix protein (M2-1); these two layers have center-to-center distances from the lipid membrane of 5 nm and 13 nm, respectively (Kiss et al., 2014). The M2-1 protein is unique to the family. The viral nucleocapsid consists of the single-stranded viral RNA genome tightly bound by associated viral proteins including the viral polymerase. In cryo-EM the nucleocapsid has helical symmetry and is 15 nm in diameter with a 6.8 nm pitch for HRSV (Bakker et al., 2013). No structural data are available for metapneumoviruses but variable estimates from negatively stained EM images average about 14 nm in diameter with a 7 nm pitch. Multiploid virions are found, although the vast majority of virions contain a single functional genome, based on the observation that UV-inactivation of RSV has single-hit kinetics.
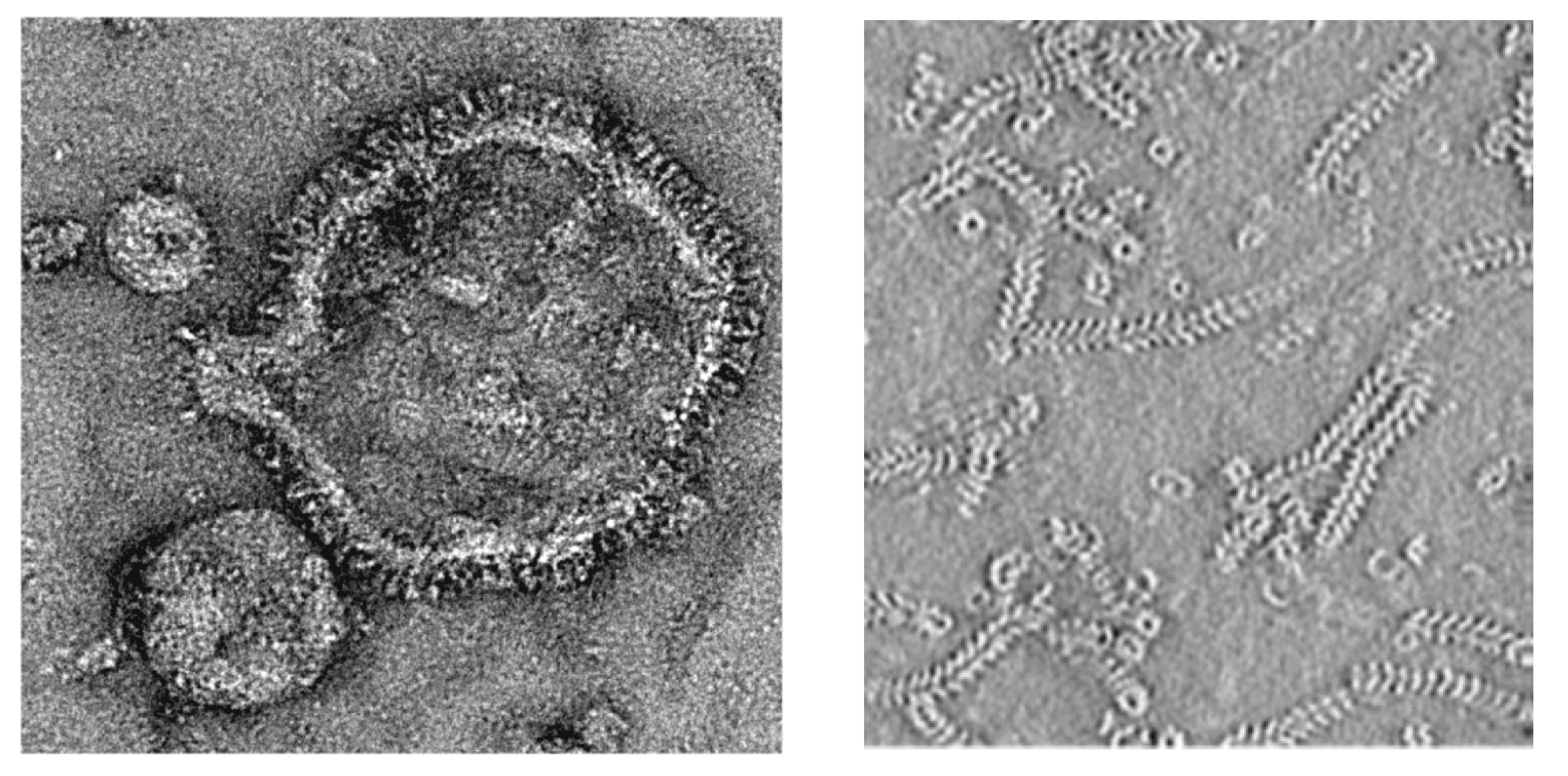 |
| Figure 1. Pneumoviridae. Left panel: negative contrast electron micrographs of intact human respiratory syncytial virus (HRSV) (courtesy of Kyle Dent, Neil Ranson and John Barr, University of Leeds). Right panel: the HRSV nucleocapsid in a denoised tomogram (reproduced from (Bakker et al., 2013) with permission from the Microbiology Society). |
Physicochemical and physical properties
Classical virological experiments describe the HRSV virion as having a Mr of about 500 ×106, and much greater for multiploid virions. Virion buoyant density in sucrose is 1.18–1.20 g cm−3. Virion S20,w is at least 1000S. Such values have not been established for human metapneumovirus (HMPV). Virions are very sensitive to freeze-thawing, heat, lipid solvents, ionic and non-ionic detergents, formaldehyde and oxidizing agents.
Nucleic acid
Virions contain the single-stranded non-segmented negative-sense viral RNA genome. This is not infectious as naked RNA. It is only infectious in the form of the nucleocapsid protein RNA complex (RNP) but as a negative-stranded RNA virus it also requires the presence of the P and L proteins in the complex to become replication and transcription competent. The RNA genome length varies substantially between members of the two genera and to a lesser extent within genera: e.g., 15 191–15 277 nt for HRSV and 13 280–13 406 nt for HMPV. Genome-size RNA is found exclusively in nucleocapsids both intracellularly and in virions
Proteins
Pneumovirus genomes encode 9–11 proteins of 4.8–250 kDa (64–2165 amino acids for HRSV), including two proteins (M2-1 and M2-2) encoded by overlapping ORFs in the M2 locus. Virion proteins common to both genera include: three nucleocapsid-associated proteins, namely a nucleoprotein (N) that tightly binds genome-length RNA along its entire length, a phosphoprotein (P) that is a polymerase co-factor and chaperonin for nucleocapsid protein monomers, and a large polymerase protein (L) that contains the enzymatic domains for polymerization and capping; two non-glycosylated membrane or matrix proteins (M and M2-1) that form separate layers underlying the virion envelope; and three glycosylated transmembrane surface envelope proteins, namely a fusion protein (F) that mediates viral penetration, an attachment protein (G), and a small hydrophobic protein (SH) that is a putative viroporin and forms ion channels of unknown significance. The M2-2 protein encoded by an internal ORF in the M2 mRNA is a non-abundant species that helps shift RNA synthesis from transcription to genome replication; its status as a virion component is unknown. The M2-1 protein also serves as an essential transcription processivity factor for members of the Orthopneumovirus genus; its role for viruses in the Metapneumovirus genus is unclear. The F protein is synthesized as a precursor (F0) that is activated following cleavage by cellular protease(s) to produce the virion disulfide-linked F1 and F2 subunits (order: amino-F2-F1-carboxyl). For HRSV and bovine respiratory syncytial virus (BRSV) but not mouse pneumonia virus (MPV)), cleavage occurs at two furin-compatible sites separated by 27 amino acids. Cleavage releases a p27 peptide that, for BRSV, has tachykinin-like activities. The F proteins of members of the Metapneumovirus genus contain a single cleavage site; members vary as to whether cleavage has specificity for trypsin or furin. The pneumovirus G protein is a heavily glycosylated protein, with N- and O-linked sugars, and a high content of serine, threonine, and proline residues contributing to a predicted non-globular structure. G has a high degree of sequence variation within and between viral species. For HRSV, G is expressed as an abundant secreted form in addition to the membrane-anchored form, and the G ectodomain contains a conserved cysteine noose with mimicry to the CX3C cytokine fractalkine. The cysteine noose and the potential for a secreted form are conserved in BRSV but otherwise these features are not found in other members of the family. Two apparent non-structural proteins (NS1, NS2), encoded by separate genes, are present only in members of the genus Orthopneumovirus. Most members lack a hemagglutinin (a hemagglutinin has been reported only for MPV, involving the G protein), and all members lack a neuraminidase.
Lipids
Lipids in the viral envelope are derived from host cell plasma membrane.
Carbohydrates
F and G proteins are both glycosylated by several N-linked carbohydrate side chains. The G protein is also heavily glycosylated by O-linked carbohydrate side chains spread along most of the ectodomain. The SH protein is present in infected cells and virions in multiple forms, including non-glycosylated forms, glycosylated forms with N-linked carbohydrate, and glycosylated forms with N-linked carbohydrate modified by the addition of polylactosaminoglycan. The dense (and variable) glycosylation and unusual amino acid composition of the G protein resemble cellular mucin.
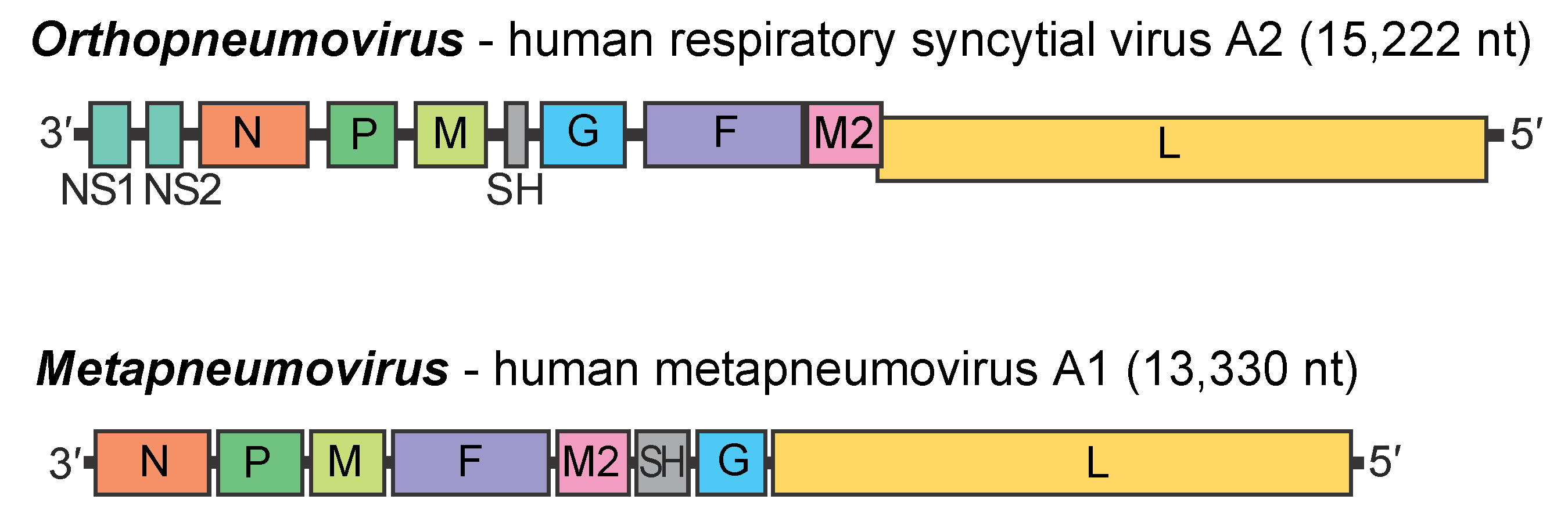
Genome organization and replication
The genome organization is illustrated in Figure 2.Pneumoviridae for viruses representing the two genera of the family. The viral genome contains 8 (genus Metapneumovirus) or 10 (genus Orthopneumovirus) genes that each encode a separate mRNA. Each mRNA has a single ORF with two exceptions: (i) the M2 mRNA encodes the M2-1 and M2-2 proteins from separate ORFs that have a small overlap, and (ii) the MPV P mRNA has an additional ORF encoding a small protein of unknown significance. For HRSV, translation of the downstream M2-2 ORF is mediated by ribosomes that exit the upstream M2-1 ORF and re-initiate at the M2-2 ORF. The genome is transcribed processively from the 3′ end by the virion-associated polymerase complex. There is a polar, decreasing gradient of gene expression across the genome. Transcription is guided by short (9–13 nt) conserved transcription start and termination/polyadenylation signals flanking each gene. The mRNAs are capped and methylated by the viral polymerase and possess 3′-poly(A) tracts synthesized by reiterative copying of the polyadenylation site. The genes are separated by short intergenic regions that are poorly conserved in sequence and length. An exception is that there is overlap between the M2 and L genes in HRSV and BRSV, but not in MPV (Figure 2. Pneumoviridae) these viruses give rise to separate mRNAs because polymerase complexes that exit the M2 gene can backtrack and re-initiate at the L gene. RNA replication by the viral polymerase involves synthesis of an intermediate, the antigenome that is a complete positive sense copy of the genome. The genomic and antigenomic RNAs do not contain a 5′ cap, nor a covalently linked protein, and the 3′ ends are not polyadenylated.
Nucleocapsid assembly is tightly linked to RNA replication
Virus replication
Receptors and the process of virus attachment are incompletely understood for members of the Pneumoviridae. The fusion F protein may be involved in attachment in addition to the so-called attachment G protein, and a number of members can replicate in vivo, albeit with reduced efficiency, when the G gene has been deleted. Attachment to cell lines in vitro involves cell surface glycosaminoglycans, but these glycans may not be similarly accessible in vivo. A number of additional cellular proteins have been suggested to serve as receptors: most recently, the cellular proteins nucleolin (Tayyari et al., 2011) and integrin avb1 (Yun et al., 2016) have been reported as receptors for HRSV and the avian and human metapneumoviruses, respectively, in each case involving the F protein. CX3CR1 may serve as a more relevant attachment receptor mediated by the G protein (Johnson et al., 2015). Following attachment, virus entry is achieved by fusion of the virus envelope with the cell membrane. This typically occurs at neutral pH, and may occur at the cell surface or in internal vesicles following endocytosis. However, fusion by some strains of HMPV requires low pH. Membrane fusion by pneumoviruses can be mediated by the F protein independent of the G protein. Transcription and RNA replication are cytoplasmic and are contemporaneous. RNA synthesis and nucleocapsid formation appear to occur in dense cytoplasmic viral inclusion bodies. Nucleocapsids associated with the M protein are enveloped by budding at the cell surface plasma membrane at sites containing virus envelope proteins. HRSV in particular buds inefficiently in vitro, with >90% of progeny virions remaining cell-associated. Pneumovirus replication in vitro does not lead to high virus titres.
Biology
Pneumoviruses have been identified conclusively only in in mammals and birds. Most viruses have a narrow host range in nature, but they are capable of infecting a wide range of cell types in vitro. Infection of cultured cells generally results in cell destruction from syncytia formation and apoptosis but temperate or persistent infections in vitro can occur. Transmission is horizontal, mainly through contact and airborne routes; no vectors are known. Transmission between host species may occur in some cases (e.g., humans and chimpanzees for HRSV and HMPV). Pneumoviruses of humans do not have animal reservoirs. Human pneumovirus infection takes place in the superficial epithelial cells of the respiratory tract and causes respiratory tract disease, and generally does not spread significantly beyond that site; avian metapneumovirus infection can be more complex and associated with disease beyond the respiratory tract. In general, pneumovirus infections are limited and eliminated by host immunity. However, virus sometimes can be shed for periods of weeks in otherwise healthy individuals, and for months in immunocompromised individuals; apart from this, latent infection and long term persistent infections are unknown.
Antigenicity
The fusion F protein is the major viral neutralization and protective antigen. The attachment G protein is a secondary neutralization and protective antigen for orthopneumoviruses but not for metapneumoviruses. Antibodies to N and, variably, to other viral proteins also are induced by infection but generally do not neutralize infectivity in vitro. Cell-mediated immunity is important for viral clearance, and various proteins of members of the Pneumoviridae have been shown to contain epitopes for cytotoxic or helper T cells.
There are two antigenic subgroups of HRSV, called A and B, which exhibit genome-wide divergence in nucleotide and amino acid sequence identity. Between the subgroups, the F protein has 89% amino acid sequence identity and has a high level of antigenic cross-reactivity, whereas the G protein has only 53% sequence identity and has a lower level of antigenic cross-reactivity. Comparable antigenic subgroups appear to exist involving BRSV: specifically, BRSV and isolates from sheep exhibit sequence dimorphism approaching that of the HRSV subgroups (e.g., 60-64% amino acid sequence identity for G protein) and appear to be two subgroups of ungulate RSV.
HMPV strains also can be divided in two genetic lineages or antigenic subgroups, A and B, each separating into sub-lineages (A1, A2a and A2b, B1 and B2). Between the two antigenic subgroups, the amino acid sequence identity for most of the proteins ranges between 92 and 98%, except for the G and SH proteins for which amino acid sequence identity can be as low as 28% and 54%, respectively. Although antibodies against the F protein are highly cross-reactive, antisera raised against virus from one antigenic subgroup cross-neutralizes virus from the other subgroup with a 16- to 128-fold reduction in efficiency, compared to only a ~4-fold reduction for the HRSV subgroups.
Four subgroups, A-D, have been described for avian metapneumovirus (AMPV), based primarily on genetic and antigenic differences. G is the most divergent protein, with 33–38% amino acid sequence identity between subgroups. Members of subgroups A and B cross react with monoclonal antibodies directed against the F glycoprotein. Subgroup C and D viruses are distinct and do not show cross reactivity against the other subgroups. In western blot analysis, cDNA-expressed F protein from AMPV subgroup C also reacted with antibodies specific for HMPV. Sequence analysis confirmed that AMPV-C is more closely related to HMPV than to AMPV-A, B, and D: for example, for the F protein, AMPV-C has 98-99% amino acid sequence identity within the subgroup, 81-82% identity with HMPV, and 71-73% identity with AMPV subgroups A, B, and D.
Derivation of names
Metapneumovirus: from Greek meta, “after”.
Orthopneumovirus: from Greek orthos, “straight”.
Pneumoviridae: from Greek pneuma, “breath”.
Respiro: from Latin respirare, “respire, breathe”.
Genus demarcation criteria
Orthopneumoviruses are distinguished from metapneumoviruses by (i) possession of 10 genes in members of the Orthopneumovirus genus as compared to 8 in members of the Metapneumovirus genus, (ii) the presence of the NS1 and NS2 genes, (iii) the SH, G, F and M2 genes being in the order SH-G-F-M2 as opposed to F-M2-SH-G for metapneumoviruses, (iv) a greater genome length (15,191–15,277 nt compared to 13,280–13,406 nt for metapneumoviruses, (v) differences in intergenic region length (up to ~57 nt versus up to ~190 nt) and (iv) a higher degree of nucleotide and amino acid sequence relatedness within each genus than between genera.
Relationships within the family
The literature on the relationships of members of the family Pneumoviridae generally is consistent with the phylogeny established on the basis of the amino acid sequence comparison of the RdRPs of these viruses. (Figure 3.Pneumoviridae). One incongruity is that the RdRP of the AMPV-C subgroup of viruses that cause turkey rhinotracheitis is phylogenetically closer to the RdRPs of the members of the species Metapneumovirus hominis. It has been classified as a member of the species Metapneumovirus avis on the basis of its host range rather than on the basis of sequence alignments.
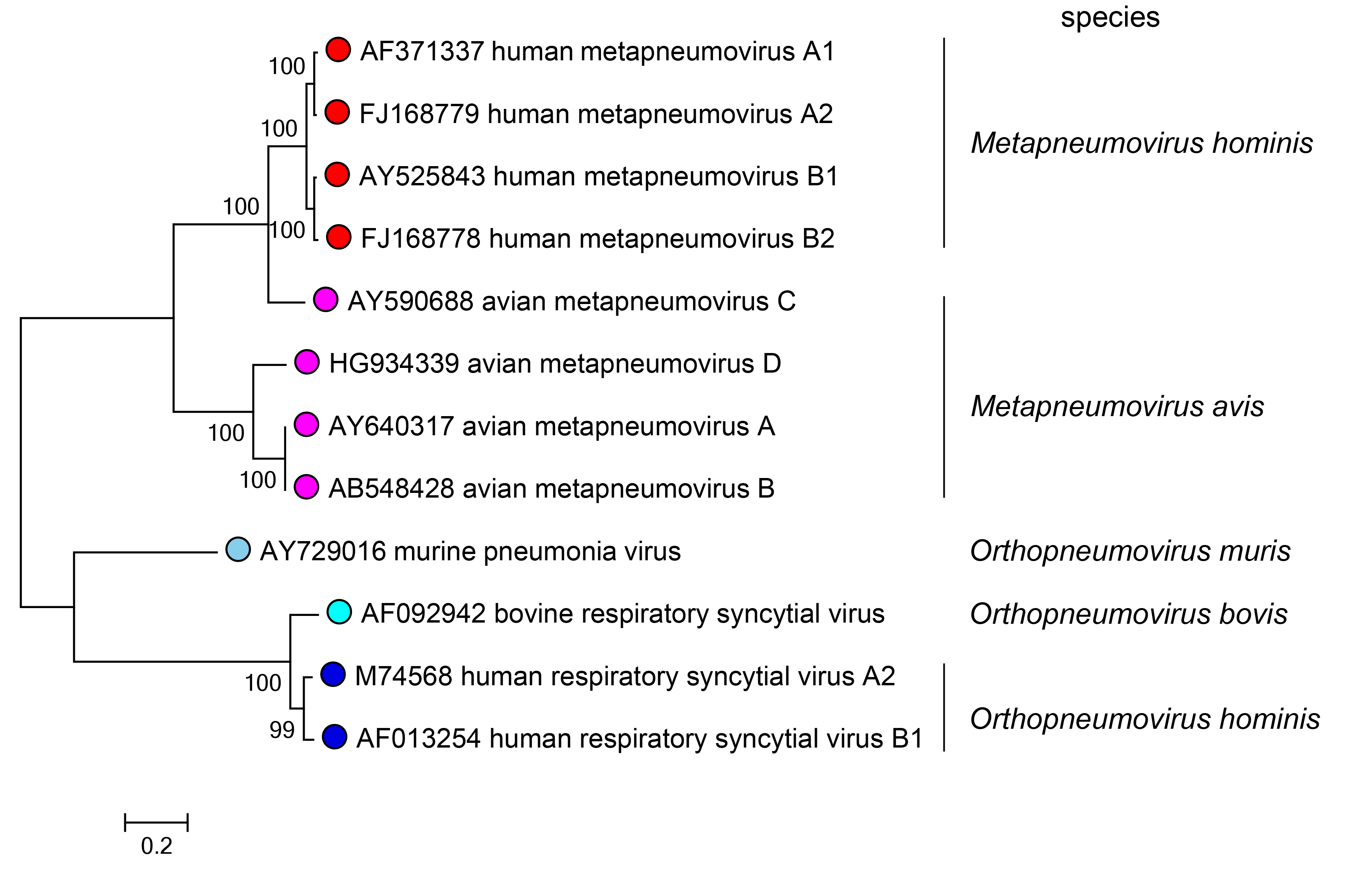 |
| Figure 3.Pneumoviridae. Phylogeny of viruses in the family Pneumoviridae. Aligned L (RdRP) sequences from representative members of the Pneumoviridae were used to produce a phylogenetic tree by Maximum Likelihood based on the Le_Gascuel_2008 model with a Gamma distribution of evolutionary rate variation between sites including invariant sites using MEGA7 (Kumar et al., 2016). The percentage of trees in which the associated taxa clustered together is shown for values over 70%. Bars indicate the species assignments of these virus isolates. This phylogenetic tree and corresponding sequence alignment are available to download from the Resources page. |
Relationships with other taxa
The member viruses of the family Pneumoviridae have a similar strategy of gene expression and replication and gene order to those of viruses in other families in the order Mononegavirales, specifically in the families Paramyxoviridae, Rhabdoviridae and Filoviridae.