Family: Pleolipoviridae
Ying Liu, Mike Dyall-Smith and Hanna M. Oksanen
The citation for this ICTV Report chapter is the summary published as Liu et al., (2022):
ICTV Virus Taxonomy Profile: Pleolipoviridae 2022, Journal of General Virology (2022) 103:001793.
Corresponding author: Hanna M. Oksanen ([email protected])
Edited by: Mart Krupovic
Posted: December 2017, updated August 2022
Summary
Members of the family Pleolipoviridae are pseudo-spherical and pleomorphic archaeal viruses composed of a membrane vesicle enclosing a DNA genome (Table 1.Pleolipoviridae). The virions do not have a protein capsid. The genome is either circular single-stranded or double-stranded, or linear double-stranded DNA molecules of approximately 7 to 17 kilonucleotides or kbp. Typically, virions contain a single type of spike protein at the envelope and a single type of internal membrane protein embedded in the envelope. All viruses infect extremely halophilic archaea in the class Halobacteria (phylum Euryarchaeota). Pleolipoviruses have a narrow host range and a persistent, non-lytic life cycle. Some viruses are temperate and can integrate into the host chromosome.
Table 1.Pleolipoviridae. Characteristics of members of the family Pleolipoviridae.
| Characteristic | Description |
| Example | Halorubrum pleomorphic virus 1 (HRPV-1; FJ685651), species Alphapleolipovirus finnoniense |
| Virion | Enveloped, pseudo-spherical and pleomorphic, virions (diameters 40–70 nm), typically with a single type of spike protein at the envelope and a single type of internal membrane protein species embedded in the envelope |
| Genome | Circular ssDNA, circular dsDNA or linear dsDNA of 7–17 kilonucleotides or kbp |
| Replication | Possibly rolling circle replication for circular genomes; protein primed replication for linear genomes |
| Translation | Prokaryotic translation using viral mRNA and host ribosomes |
| Host Range | Archaea, euryarchaeal Halorubrum, Haloarcula, Halogeometricum or Natrinema strains |
| Taxonomy | Realm Monodnaviria, kingdom Trapavirae, phylum Saleviricota, class Huolimaviricetes, order Haloruvirales; three genera Alphapleolipovirus, Betapleolipovirus and Gammapleolipovirus including 16 species |
Virion
Morphology
Virions consist of a genome surrounded by a pleomorphic membrane vesicle of 40–70 nm in diameter with irregularly distributed spike structures (Pietilä et al., 2009, Pietilä et al., 2012, El Omari et al., 2019, Demina and Oksanen 2020) (Figure 1.Pleolipoviridae). Virions lack a capsid or a nucleocapsid structure. There are two to four types of major structural protein that either form the spikes or are integral membrane proteins. The spikes are formed of one or two types of membrane-anchored protein species. The spike protein is anchored to the lipid membrane with a C-terminal transmembrane domain (Pietilä et al., 2010, Senčilo et al., 2012). The integral membrane proteins (either one or two types) are located in the internal side of the membrane vesicle.
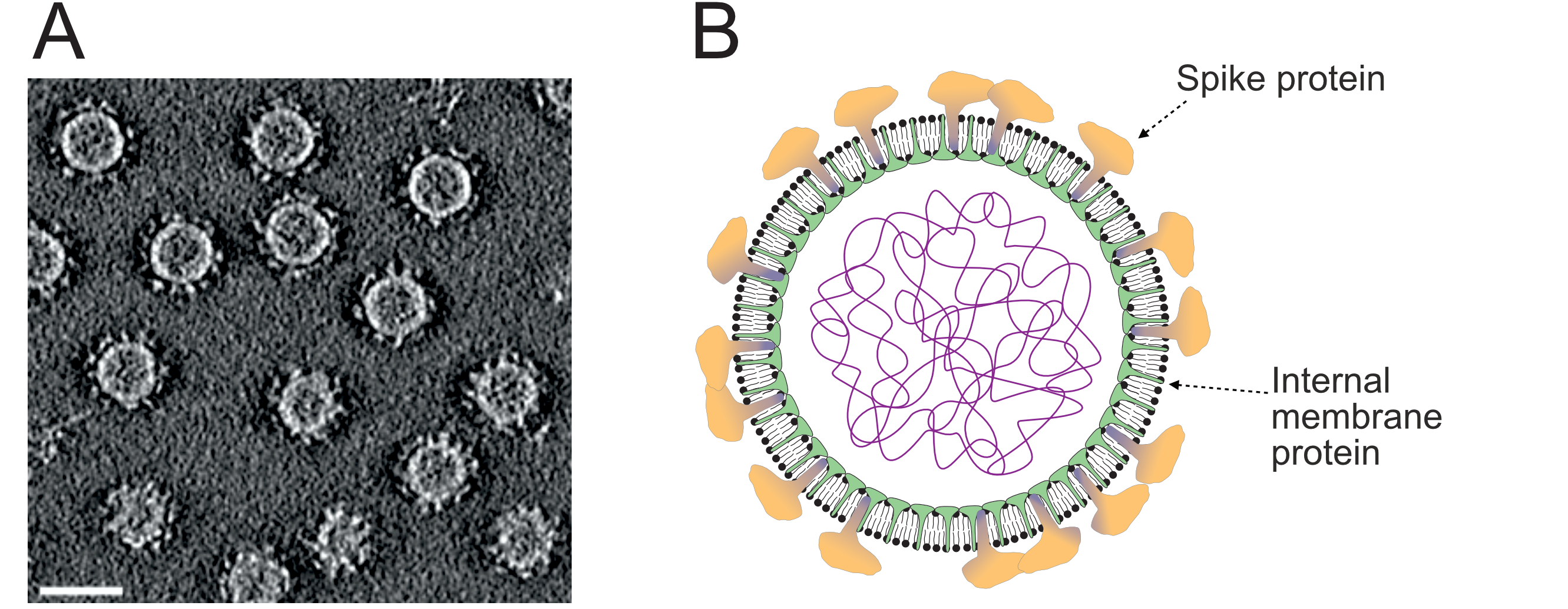 |
| Figure 1.Pleolipoviridae. Morphology of pleolipovirus virions. (a) A slice through a three-dimensional cryo-electron microscopy tomogram of Halorubrum pleomorphic virus 6 reconstructed from tilt series data. Reproduced under Creative commons license 4.0 from (El Omari et al., 2019). Scale bar is 40 nm. (b) Schematic presentation of the pleolipovirus virion. |
Physicochemical and physical properties
Virions are typically stable at high ionic strengths (e.g. above 2.5 M NaCl) or even in saturated NaCl but some virions are sensitive to lowered levels of NaCl or CaCl2. Virion buoyant densities in CsCl are from 1.26 to 1.34 g cm-3. Infectivity is sensitive to detergents and the organic solvent chloroform. Virions are stable at temperatures below 60 °C.
Nucleic acid
Members of family Pleolipoviridae have different types of DNA genomes (Roine et al., 2010, Senčilo et al., 2012). Genomes are circular single-stranded DNA of 7-11 kilonucleotides, circular double-stranded DNA of approximately 8-17 kbp, or linear double-stranded DNA of approximately 16 kbp (Figure 2.Pleolipoviridae). Members of the genus Alphapleolipovirus have either single-stranded or double-stranded circular DNA genomes. Genomes of members of the genus Betapleolipovirus are circular double-stranded DNA molecules that may contain single-stranded discontinuities, and the linear genomes of members of the genus Gammapleolipovirus contains inverted terminal repeat sequences and terminal proteins attached to the genome ends.
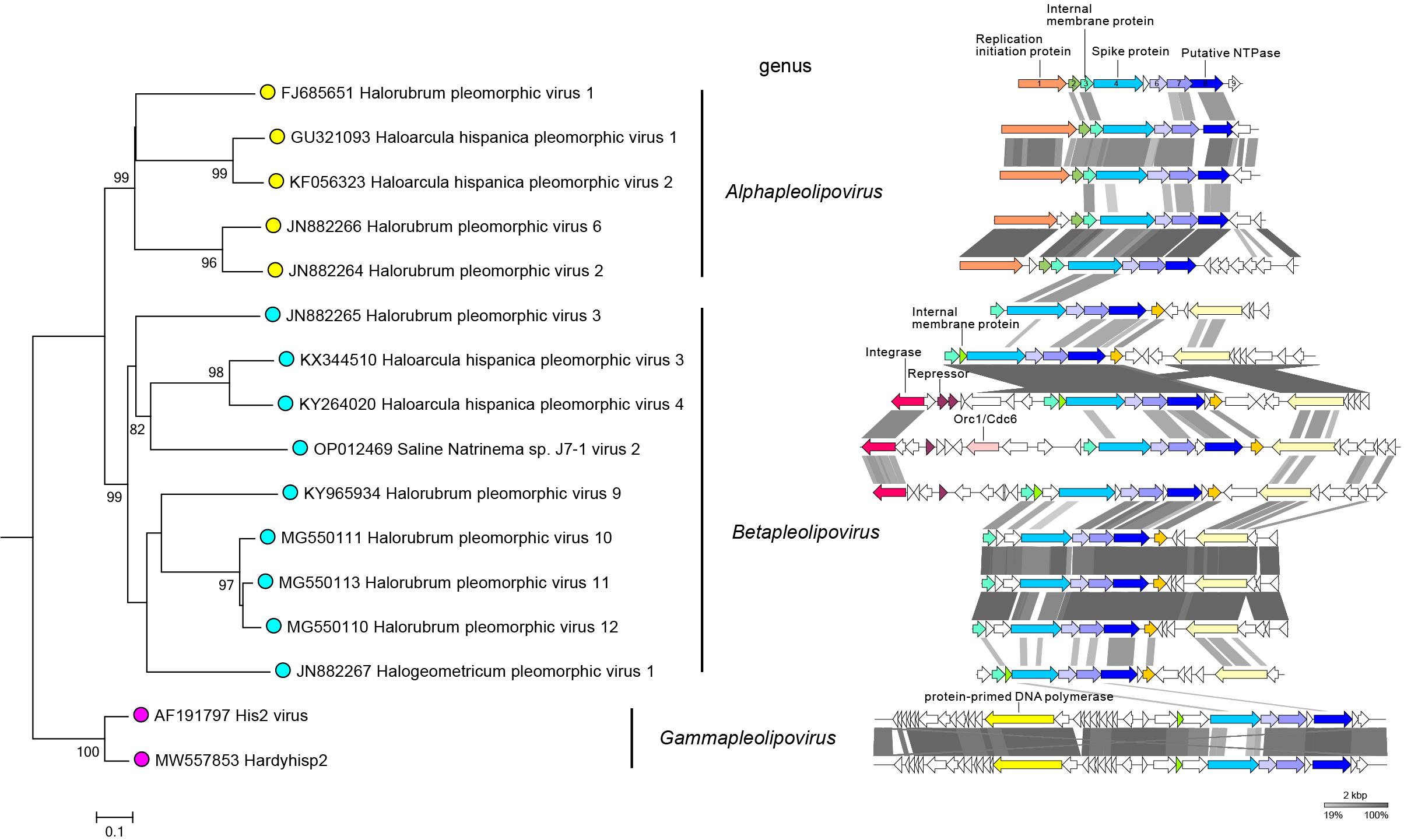 |
| Figure 2.Pleolipoviridae. Genomes of pleolipoviruses. The phylogenomic tree of pleolipoviruses (left panel) based on whole genome analysis at the amino acid level was performed using VICTOR (Meier-Kolthoff and Göker 2017). The tree is rooted with gammapleolipoviruses, and the branch length is scaled in terms of the Genome BLAST Distance Phylogeny (GBDP) distance formula D6. The bootstrap support values >80% are shown. The treefile is available on the Resources page. The right panel shows the linear representation of the pleolipovirus genomes depicted in the tree. Gammapleolipoviruses have linear genomes. The genomes of other viruses are circular. Genes and ORFs are shown as arrows. Homologous genes are indicted with the same colors, and are connected between viruses by shadings of different degrees of grey based on the amino acid sequence identity. Genes and ORFs are named in HRPV-1. HRPV-1 genes 3, 4 and 8, and ORFs 6 and 7 form a group of conserved genes characteristic of pleolipovirus genomes. |
Proteins
Pleolipovirus genomes have from 8 to 35 predicted open reading frames (ORFs), of which 2-4 have been shown to code for structural proteins and the rest have unknown functions or predicted function based on sequence similarity. Virions contain one or two types of internal membrane protein, with several predicted transmembrane regions, and one or two types of spike protein, which are processed during maturation. Halorubrum pleomorphic virus 1 (HRPV-1) has also been identified to contain a minor structural protein VP8, which is predicted to be an NTPase (Pietilä et al., 2009).
The non-structural proteins of the members of the genus Alphapleolipovirus include putative rolling circle replication initiation proteins (RCR Reps) and members of the genus Gammapleolipovirus encode a putative type B DNA polymerase (Bath et al., 2006, Pietilä et al., 2009, Roine et al., 2010). The fusion-inducing spike protein has a unique V-shaped fold and belongs to a new structural class of fusion proteins (El Omari et al., 2019). The betapleolipoviruses Haloarcula hispanica pleomorphic virus 4 (HHPV4), Saline Natrinema sp. J7-1 virus 2 (SNJ2), and Halorubrum pleomorphic virus 9 (HRPV9) have genes encoding integrases (Liu et al., 2015, Atanasova et al., 2018a, Atanasova et al., 2018b).
Lipids
Virions contain lipids that are derived non-selectively from host cell membranes. Viral lipid species compositions are similar to that of the host archaeon (Pietilä et al., 2012). Lipids form the sole outer layer of the virion with virus-specific spike and internal membrane proteins. Polar lipids identified in HRPV-1 virions are cardiolipin (CL), phosphatidylglycerol (PG), phosphatidylglycerophosphate methyl ester (PGP-Me), and phosphatidylglycerosulfate (PGS) (Pietilä et al., 2010). In addition, virions of HRPV-1 contain neutral lipids and glycolipids (Pietilä et al., 2010). Based on thin-layer chromatography, CL, PG, PGP-Me, and PGS are typical lipid species in the virions of the pleolipoviruses (Pietilä et al., 2012, Atanasova et al., 2018b, Mizuno et al., 2019). The spike protein VP4 of Halogeometricum pleomorphic virus 1 (HGPV-1) contains a lipid modification (Pietilä et al., 2012).
Carbohydrates
Virions can contain carbohydrates in the form of glycolipids (Pietilä et al., 2010).
The spike protein VP4 of HRPV-1 is glycosylated (Pietilä et al., 2010) with a pentasaccharide as the major N-glycan (Kandiba et al., 2012).
Genome organization and replication
All pleolipovirus genomes carry a conserved block of collinear genes, encoding major structural proteins, a putative NTPase, and other putative proteins (genes 3, 4 and 8; ORFs 6 and 7 of Halorubrum pleomorphic virus 1) (Figure 2.Pleolipoviridae). Viruses enter the host cell by membrane fusion of the viral envelope with the host cell cytoplasmic membrane (Pietilä et al., 2009, El Omari et al., 2019). Virus genomes are predicted to employ different genome replication strategies, including rolling circle replication (RCR; circular genomes) and protein-primed replication carried out by family B-type polymerase (linear genomes), although direct experimental evidence is missing (Bath et al., 2006, Pietilä et al., 2009, Roine et al., 2010). His2 virus genes are expressed in early, middle, and late phases and by 4.5 hr after infection the majority of virus genes are actively transcribed (Lee et al., 2020). Relatively few host genes show significant differential regulation during His2 infection with most of them predicted to be involved in transport, translation, and metabolism. During virus assembly, the lipids are unselectively acquired from the host cell membrane (Pietilä et al., 2012). Viruses have non-lytic life cycles and they exit the cells continuously starting 3–4 hours post infection (Pietilä et al., 2012, Svirskaitė et al., 2016). During the infection cycle, host growth can be slightly retarded, although the infected cells are consuming oxygen as much as the non-infected ones (Pietilä et al., 2009, Roine et al., 2010, Pietilä et al., 2012, Svirskaitė et al., 2016). The non-lytic life cycle is also evidenced by the hazy appearance of the plaques the viruses form on the host lawn. The characteristics of the non-lytic life cycle and the presence of lipid envelope in the virions suggest that pleolipoviruses exit host cells using budding. Some of the pleolipoviruses have a temperate life cycle and have genes encoding a putative integrase (Liu et al., 2015, Atanasova et al., 2018a, Atanasova et al., 2018b). The mechanism of integration of betapleolipovirus SNJ2 has been experimentally characterized (Wang et al., 2018).
Biology
The current members of family Pleolipoviridae have a narrow host range. They typically infect only their original isolation hosts, all of which belong to the class Halobacteria. Pleolipoviruses display a wide geographical distribution and originate from hypersaline environments such as solar salterns and hypersaline lakes in Europe, Asia, Africa, and Australia (Bath et al., 2006, Atanasova et al., 2012, Pietilä et al., 2012, Li et al., 2014, Mizuno et al., 2019). In addition, pleolipovirus-related proviral regions are commonly found in haloarchaeal genomes.
Derivation of names
Pleolipoviridae: from the Greek pleo, meaning "many" and the Greek lipos, meaning "lipid".
Genus demarcation criteria
Genera are identified by the gene content and well-supported monophyletic groups based on phylogenomic analysis of the whole genome sequences. The following criteria are used to differentiate genera in the family:
- Genome type i.e. linear or circular
- Alphapleolipoviruses have either single-stranded or double-stranded circular DNA.
- Betapleolipoviruses have circular double-stranded DNA genomes that can contain single-stranded discontinuities
- Gammapleolipoviruses have linear double-stranded DNA genomes
- Besides the conserved cluster of five genes shared by all pleolipoviruses (genes 3, 4 and 8; ORFs 6 and 7 of Halorubrum pleomorphic virus 1):
- Alphapleolipovirus genomes share an ORF coding for a rolling circle replication initiation proteins (RCR Rep).
- Betapleolipovirus genomes share two ORFs coding for proteins of unknown function (e.g. Halorubrum pleomorphic virus 3 ORFs 6 and 9). One of those is predicted to contain a winged helix-turn-helix (wHTH) domain.
- Gammapleolipovirus genomes have a gene encoding a putative type B DNA polymerase
Relationships within the family
Members of family Pleolipoviridae have genomes that share a similar arrangement of structural protein and some other genes (genome synteny) as well as some sequence similarity between homologous genes. The sequence similarity between pleolipovirus genomes is typically low, but there are exceptions (Figure 3.Pleolipoviridae). The overall nucleotide sequence similarities between Haloarcula hispanica pleomorphic virus 1 (HHPV-1) and Haloarcula hispanica pleomorphic virus 2 (HHPV-2) and between HRPV-2 and HRPV-6 are 71% and 79%, respectively (Senčilo et al., 2012, Li et al., 2014). Halorubrum pleomorphic virus 10, 11, and 12 (HRPV10, HRPV11, HRPV12) have highly similar genomes with 84-89% nucleotide sequence similarity (Mizuno et al., 2019). Two gammapleolipoviruses His2 and Hardyhisp2 share 81% nucleotide similarity (Dyall-Smith et al., 2021). Furthermore, Haloarcula hispanica pleomorphic virus 3 and 4 (HHPV3 and HHPV4), although representing different species, share 100% identical genomic regions, but the overall pairwise genome identity is 76% (Figure 2.Pleolipoviridae and Figure 3.Pleolipoviridae) (Demina et al., 2016, Atanasova et al., 2018b).
Relationships with other taxa
Members of the family Pleolipoviridae form a distinct group of viruses. Their structural proteins share no significant amino acid similarity with the structural proteins of any other virus. Acholeplasma virus L2 (APVL2, family Plasmaviridae) infecting the bacterium Acholeplasma laidlawii displays similar virion morphology (quasi-spherical, pleomorphic) as that of pleolipoviruses. In both cases, the virion envelope is composed of host-derived lipids that are acquired unselectively (Al-Shammari and Smith 1981, Dybvig et al., 1985, Maniloff et al., 1994). The APVL2 genome is a circular double-stranded DNA molecule (~12 kbp) as the genomes of some members of the genus Alphapleolipovirus (Pietilä et al., 2009, Senčilo et al., 2012). Nevertheless, the genome sequence of APVL2 shares no identity with pleolipoviral sequences at the nucleotide sequence level. On the basis of virion morphology and structural protein patterns, it has been hypothesized that HRPV-1 could be related to mycoplasmavirus L172 which infects Acholeplasma laidlawii and contains a circular single-stranded DNA genome (Dybvig et al., 1985, Pietilä et al., 2009). Currently, there is no genome sequence data available for mycoplasmavirus L172 and it has not been classified to any taxon.
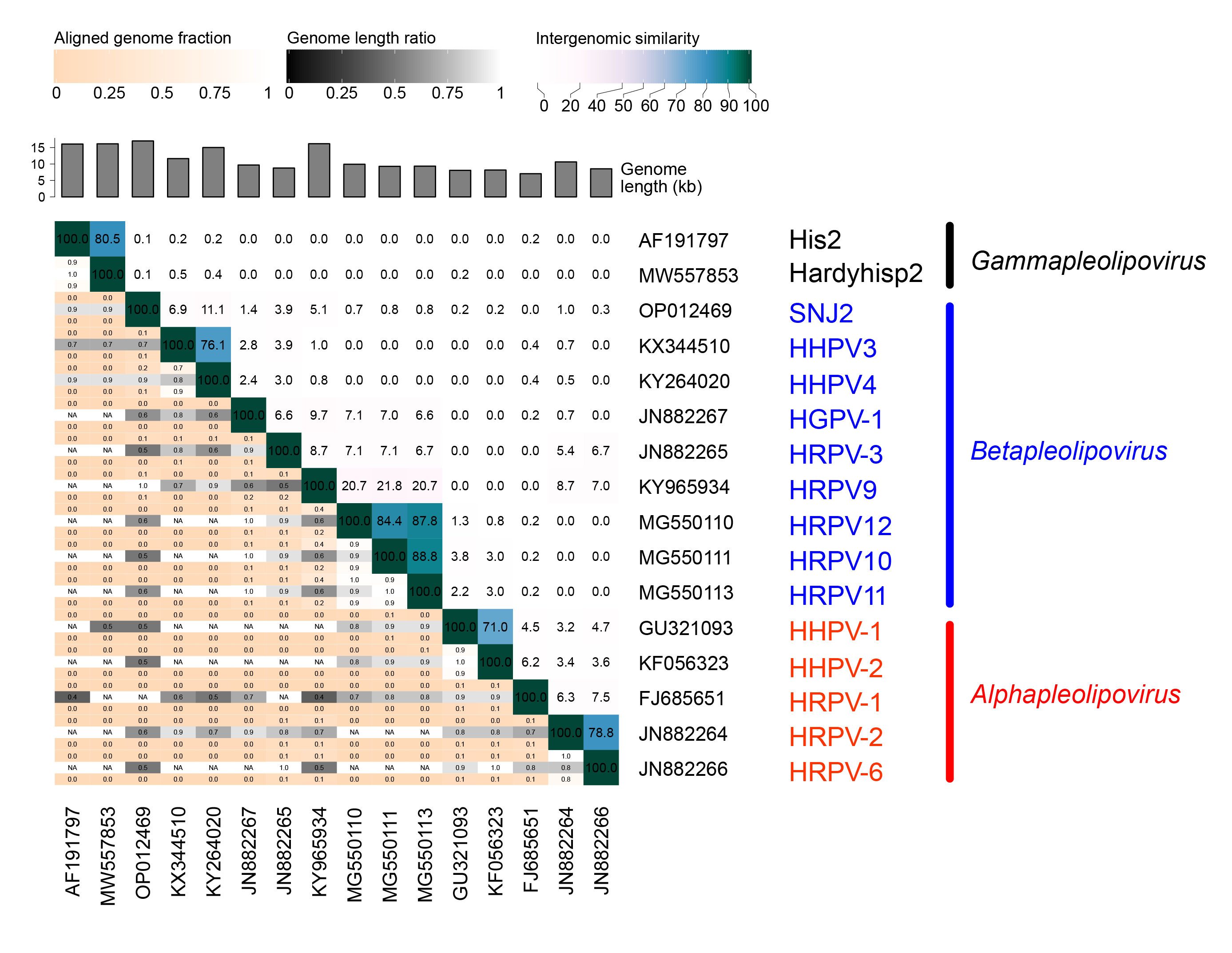 |
| Figure 3.Pleolipoviridae. Intergenomic similarities of the members of Pleolipoviridae, as output from the VIRIDIC webserver (Moraru et al., 2020). Virus names are shown at the right, colour coded and labeled to indicate their membership of the Alphapleolipovirus, Betapleolipovirus or Gammapleolipovirus genera. Accession codes are given at the lower edge. Genome length, and keys for aligned genome fraction, genome length ratio and intergenomic similarity are shown at top. |

