Family: Nyamiviridae
Ralf G. Dietzgen, Andrew E. Firth, Dàohóng Jiāng, Sandra Junglen, Hideki Kondo, Jens H. Kuhn, Sofia Paraskevopoulou and Nikos Vasilakis
Corresponding author: Ralf G. Dietzgen ([email protected])
Edited by: Stuart G. Siddell and Peter Simmonds
Posted: December 2017, updated June 2019, August 2021, June 2022, May 2023, June 2024
PDF: ICTV_Nyamiviridae.pdf (2019 version)
Summary
Nyamiviridae is a family of viruses in the order Mononegavirales with non-segmented or segmented, negative-sense RNA genomes of approximately 10–13 kb (Table 1 Nyamiviridae). Several viruses in the family Nyamiviridae have been identified by next generation sequencing (Shi et al., 2016) and the family was recently expanded to include the genus Formivirus. Nyamiviridae now includes the genera Berhavirus, Crustavirus, Formivirus, Nyavirus, Orinovirus, Socyvirus, and Tapwovirus that form monophyletic clades on phylogenetic analysis of the RNA-directed RNA polymerase. Nyamivirids have a similar genome organization, including the number and locations of genes identified by homology with those of other mononegavirals. Only viruses of the genus Tapwovirus have segmented genomes. Viruses of the genus Nyavirus are tick-borne and some of them also infect birds (Takahashi et al., 1982). Members of the Socyvirus genus infect plant parasitic nematodes (Bekal et al., 2011). Other nyamivirids were identified in marine animals and have been classified in the genus Berhavirus (echinoderms and sipunculids) and Crustavirus (crustaceans). Nyamivirids found in moths and beetles are members of the genus Orinovirus, and those associated with tapeworms belong to the genus Tapwovirus. Members of the genus Formivirus are associated with ants and wasps. Members of the genera Nyavirus and Socyvirus produce enveloped, spherical virions (Kuhn et al., 2013). For the other nyamivirids, no virion information is available.
Table 1 Nyamiviridae. Characteristics of members of the family Nyamiviridae.
| Characteristic | Description |
| Example | Nyamanini virus (FJ554526), species Nyavirus nyamaniniense, genus Nyavirus |
| Virion | Enveloped, spherical particles, approximately 100–130 nm in diameter |
| Genome | Negative-sense, non-segmented or bi-segmented RNA of up to 13.3 kb |
| Replication | Nuclear: the large protein, which contains an RNA-directed RNA polymerase domain, engages with ribonucleoprotein at the genome 3′-end |
| Translation | Individual putatively polyadenylated mRNAs are translated in the cytoplasm |
| Host Range | Invertebrates: ticks, insects, echinonoderms, sipunculids, crustaceans, nematodes, tapeworms; vertebrates: land- and seabirds |
| Taxonomy | Realm Riboviria, phylum Negarnaviricota, subphylum Haploviricotina, class Monjiviricetes, order Mononegavirales. The family includes 7 genera and 29 species. The genus Crustavirus includes 3 species, the genera Berhavirus, Nyavirus and Formivirus include 7 species each, the genera Orinovirus and Tapwovirus include 2 species each, and the genus Socyvirus includes 1 species |
Virion
Morphology
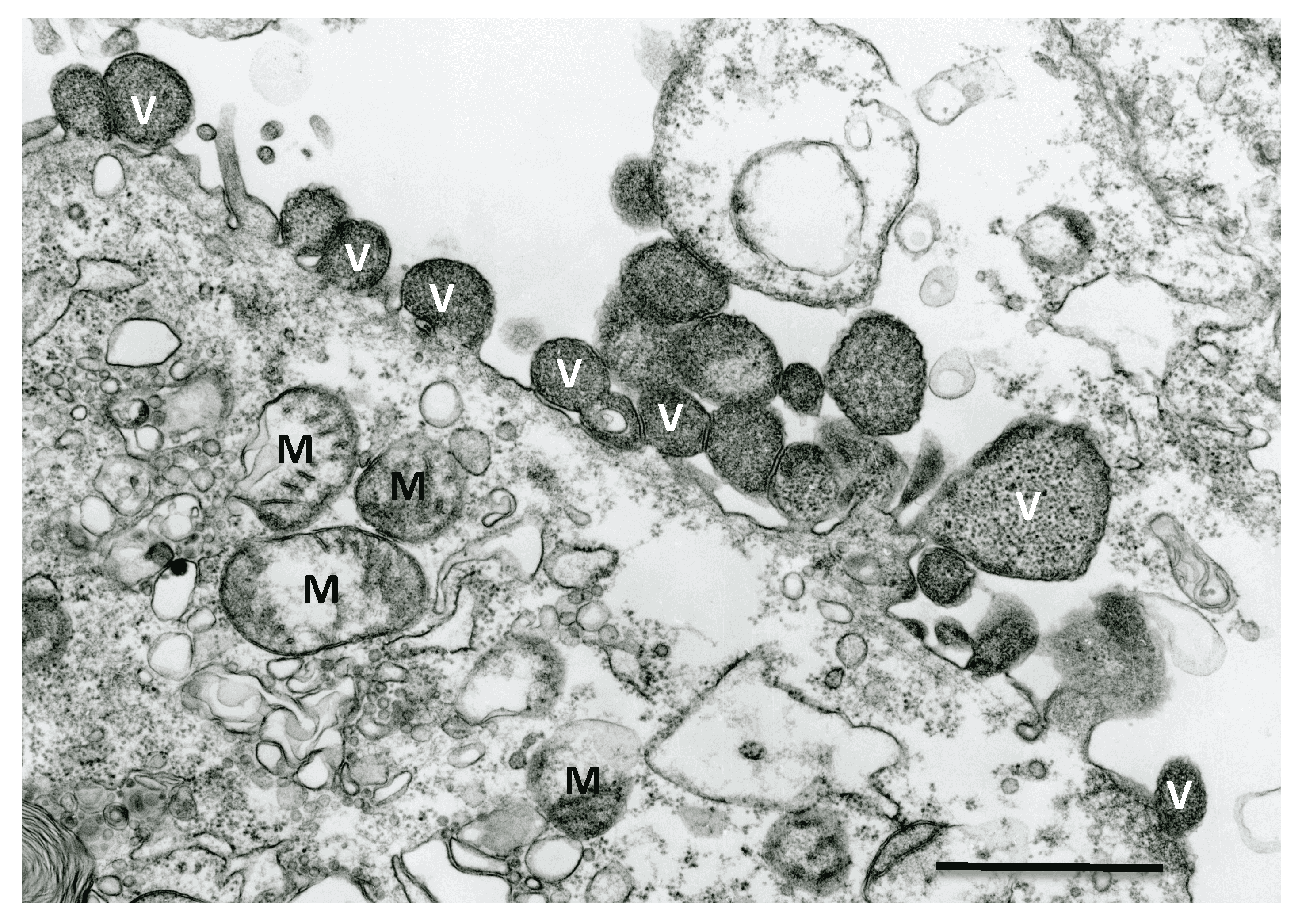
Virions of Nyamanini virus are enveloped and spherical with a diameter of 100–130 nm; virions of Sierra Nevada virus are of similar appearance (Figure 1.Nyamiviridae). San Jacinto virus-infected Vero E6 cells release large spherical (320–800 nm) or pleomorphic and elongated particles (350 × 540 nm to 280 × 1120 nm and 3280 × 1275 nm).
Physicochemical and physical properties
None reported.
Nucleic acid
Non-segmented, negative-sense RNA of 10–13 kb.
Proteins
Nyamivirids encode 4 to 8 structural proteins. Among them are the nucleocapsid protein (N), glycoprotein (G) and large protein (L), which contains an RNA-directed RNA polymerase (RdRP) domain, that are identified based on sequence similarity and structural properties shared with mononegavirus homologues. Functions of the other encoded proteins are largely unknown but may be those of matrix and polymerase cofactor proteins.
Lipids
None reported.
Carbohydrates
None reported.
Genome organization and replication
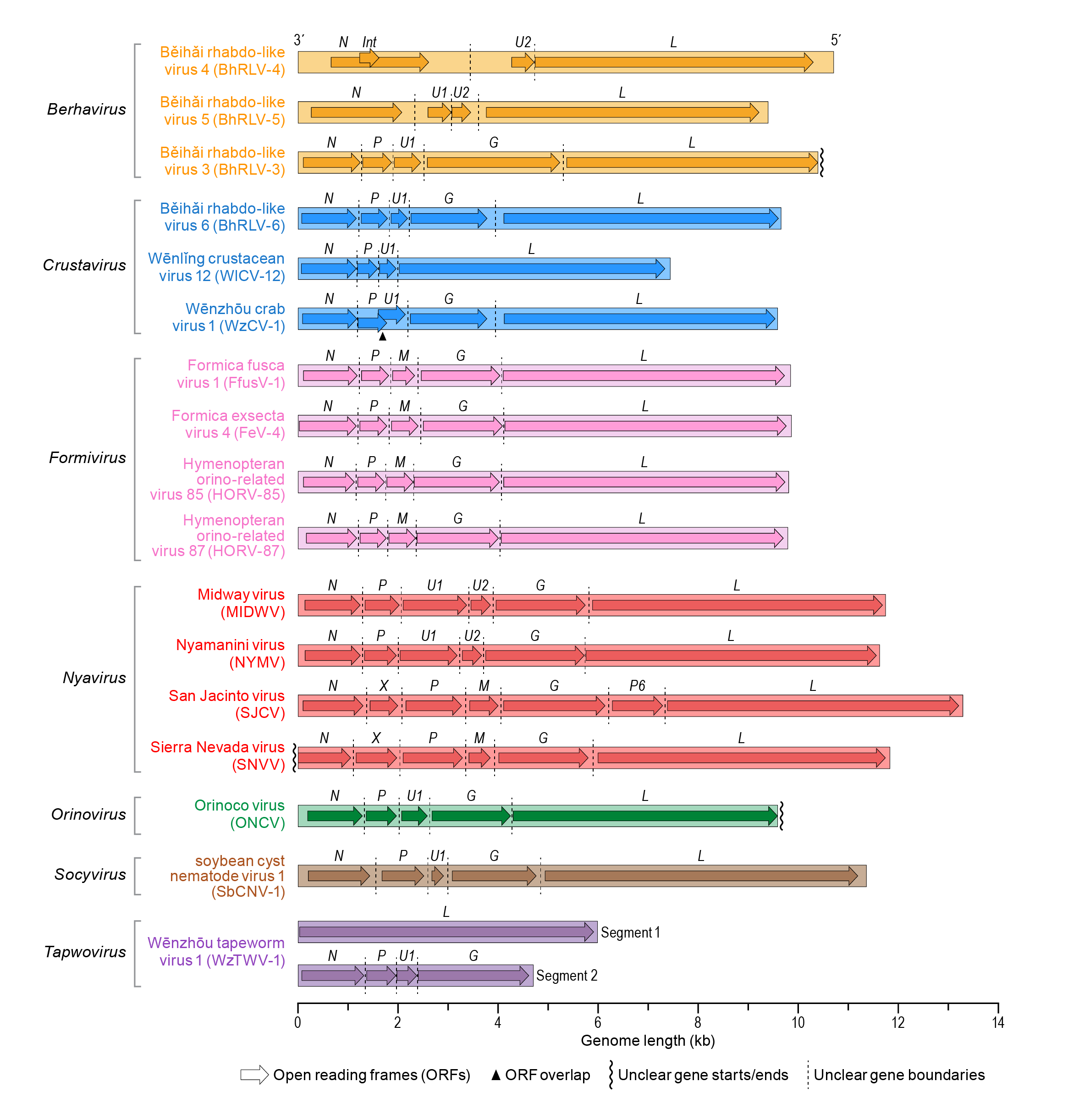
Nyamivirid negative-sense RNA genomes range from 10–13 kb (Figure 2.Nyamiviridae). All nyamivirids in the genera Nyavirus and Socyvirus have non-segmented genomes with 4 to 8 open reading frames (ORFs) that encode mostly structural proteins. Nyamivirids in the genera Berhavirus, Crustavirus, and Orinovirus also have non-segmented genomes with up to 5 ORFs. The only known member of Tapwovirus has a bi-segmented genome with RNA 1 encoding the large protein (L) and the other ORFs located on RNA 2. Knowledge about nyamivirid replication is limited. Nyamanini virus (genus Nyavirus) replicates in the nucleus of infected cells.
|
| Figure 2. Nyamiviridae. Genome organization of viruses in the family Nyamiviridae, genera Berhavirus, Crustavirus, Formivirus, Nyavirus, Orinovirus, Socyvirus and Tapwovirus. Box arrows indicate the position and length of each ORF. Putative nucleocapsid protein (N), phosphoprotein (P), glycoprotein (G) and large protein (L) ORFs are indicated; ORFs encoding hypothetical proteins with unknown function are numbered U1, U2; int - internal ORF, X - ORF with unknown function. |
Biology
Viruses of the genus Nyavirus are tick-borne and some of them also infect birds, whereas members of the Socyvirus genus infect plant parasitic nematodes. Viruses of the genus Berhavirus are associated with marine echinoderms and sipunculids, viruses of the genus Crustavirus are associated with crustaceans, viruses of the genus Orinovirus with moths, viruses of the genus Formivirus are associated with ants and wasps, and viruses of the genus Tapwovirus with tapeworms.
Antigenicity
An antigenic relationship between Nyamanini virus and Midway virus (genus Nyavirus) was demonstrated by complement fixation tests.
Genus demarcation criteria
Due to the small number of genus members, no firm genus demarcation criteria have been established beyond phylogeny and the hosts the viruses infect.
Derivation of names
Berhavirus: from Běihǎi rhabdo-like virus 3.
Crustavirus: from Crustacea.
Formivirus: from Formica, the ant genus name associated with the ant hosts in which Formica fusca virus 1 and Formica exsecta virus 4 were discovered.
Orinovirus: from Orinoco virus.
Nyamiviridae: from Nyamanini Pan (place of isolation of Nyamanini virus in South Africa) and Midway Atoll (place of isolation of Midway virus in the USA).
Nyavirus: from Nyamanini Pan (place of isolation of Nyamanini virus in South Africa. Species in the genus are named after the location where species members were first identified.
Socyvirus: from soybean cyst nematode virus. Species in the genus are named after the host.
Tapwovirus: from Wēnzhōu tapeworm virus 1.
Relationships within the family
Viruses assigned to each genus appear to form monophyletic clades but bootstrap support in the phylogenetic analyses of L polymerase amino acid sequences is weak (Figure 3.Nyamiviridae). Nyamivirids have a similar genome organization, including the number and locations of genes identified by homology with those of other mononegaviruses (Figure 2.Nyamiviridae).
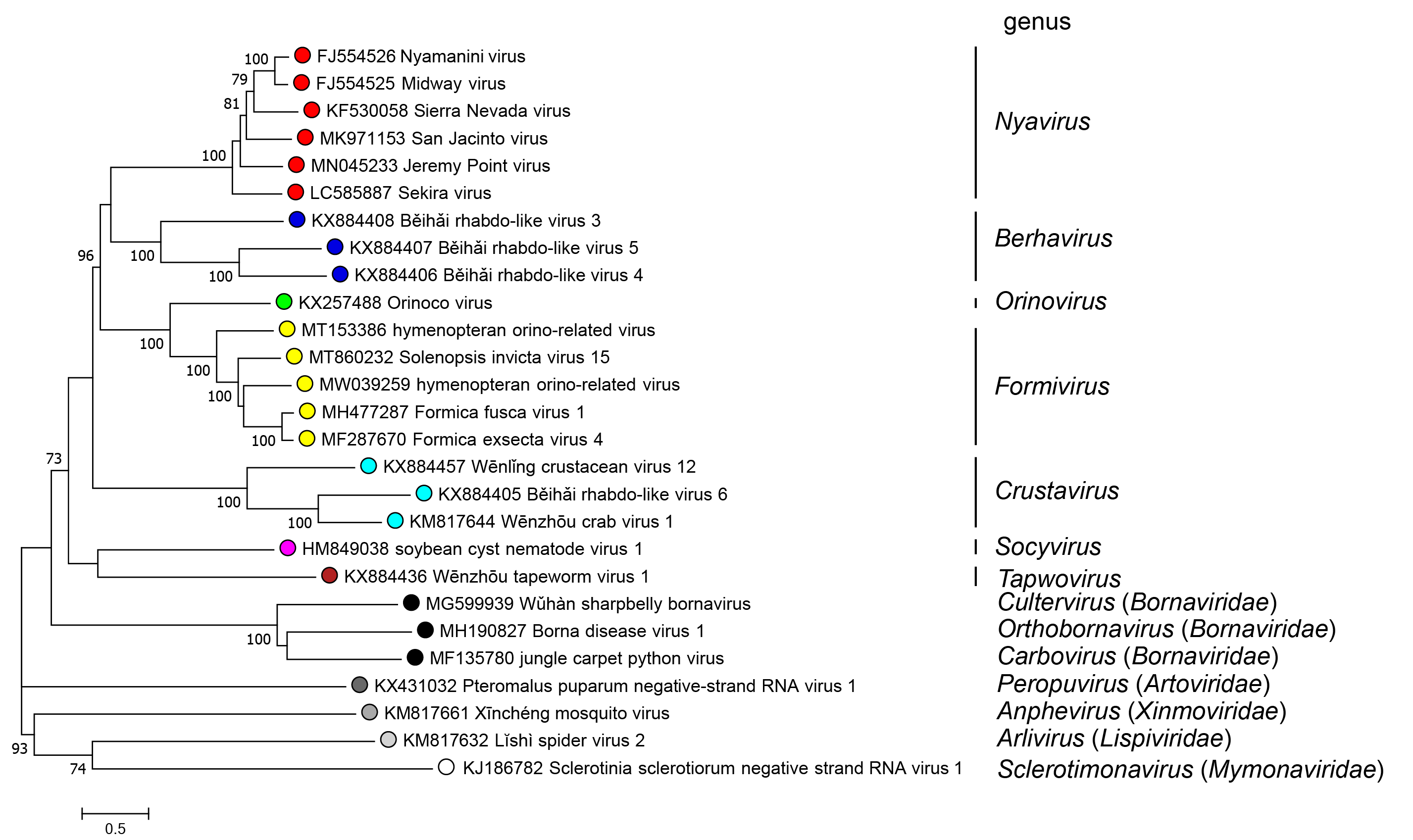

|
| Figure 3.Nyamiviridae. Phylogenetic maximum likelihood tree of mononegavirus L polymerase protein sequences highlighting Nyamiviridae and its genera. Amino acid sequences were aligned using MUSCLE (Edgar 2004). The resulting alignment was used to generate a phylogenetic tree using MEGA7 (Kumar et al., 2016) with the best-fit model JTT + G + I +F. The colors indicate the different mononegavirus families and nyamivirid genera. Numbers at the nodes indicate bootstrap support over 70% (100 replicates). |
Relationships with other taxa
Some biological properties of nyamivirids appear to be similar to those of filovirids (mononegaviral family Filoviridae) and bornavirids (mononegaviral family Bornaviridae).