Family: Barnaviridae
Chapter Version: ICTV Ninth Report; 2009 Taxonomy Release
Since only one genus is currently recognized, the family description corresponds to the genus description.
Genus Barnavirus
Type species Mushroom bacilliform virus
Virion properties
Morphology
Virions are bacilliform, non-enveloped and lack prominent surface projections. Typically, virions are 19×50 nm, but range between 18 and 20 nm in width and 48 and 53 nm in length (Figure 1). Optical diffraction patterns of the virions resemble those of virions of Alfalfa mosaic virus, suggesting a morphological subunit diameter of about 10 nm and a T=1 icosahedral symmetry.
Physicochemical and physical properties
Virion Mr is 7.1×106, buoyant density in Cs2SO4 is 1.32 g cm−3. Virions are stable between pH 6 and 8 and ionic strength of 0.01 to 0.1M phosphate, and are insensitive to chloroform.
Nucleic acid
Virions contain a single linear molecule of a positive sense ssRNA, 4.0 kb in size. The complete 4009 nt sequence of mushroom bacilliform virus (MBV) is available. The RNA has a linked VPg and appears to lack a poly(A) tail. RNA constitutes about 20% of virion weight.
Proteins
Virions contain a single major CP of 21.9 kDa. There are probably 240 molecules in each capsid.
Lipids
None reported.
Carbohydrates
None reported.
Genome organization and replication
The RNA genome (4009 nt) contains four major and three minor ORFs and has 5′- and 3′-UTRs of 60 nt and 250 nt, respectively. ORFs 1 to 4 encode polypeptides of 20, 73, 47 and 22 kDa, respectively. The deduced aa sequence of ORF2 contains putative serine protease motifs related to chymotrypsin. ORF3 encodes a putative RdRp and ORF4 encodes the CP. ORFs 5 to 7 encode 8, 6.5 and 6 kDa polypeptides, respectively. The polypeptides potentially encoded by ORFs 1, 5, 6 and 7 show no homology to known polypeptides (Figure 2).
In a cell-free system, genomic length RNA directs the synthesis of a 21 kDA and a 77 kDa polypeptide and several minor polypeptides of 18–60 kDa. The full-length genomic RNA and a sgRNA (0.9 kb) encoding ORF4 (CP) are found in infected cells. Virions accumulate singly or as aggregates in the cytoplasm.
Antigenic properties
Virions are highly immunogenic.
Biological properties
The virus infects the common cultivated button mushroom (Agaricus bisporus). Bacilliform particles, which are morphologically similar to MBV, have been observed in the field mushroom A. campestris. Transmission is horizontal via mycelium and possibly basidiospores. Distribution of MBV coincides with that of the commercial cultivation of A. bisporus; the virus has been reported to occur in most major mushroom-growing countries. MBV is capable of autonomous replication, but commonly occurs as a double infection with a dsRNA virus (LaFrance isometric virus, LFIV) in mushrooms afflicted with La France disease. MBV is not required in pathogenesis involving LFIV, but it remains to be determined if it is a second, minor causal agent of LaFrance disease, the etiologic agent of an unrecognized pathology or benign. MBV RNA and LFIV dsRNA do not share sequence homology.
Species demarcation criteria in the genus
Not applicable.
List of species in the genus Barnavirus
| Mushroom bacilliform virus |
|
|
| Mushroom bacilliform virus-AUS LF-1 | [U07551] | (MBV- LF1) |
Species names are in italic script; isolate names are in roman script. Sequence accession numbers and assigned abbreviations ( ) are also listed.
List of other related viruses which may be members of the genus Barnavirus but have not been approved as species
None reported.
Phylogenetic relationships within the family
Not applicable.
Similarity with other taxa
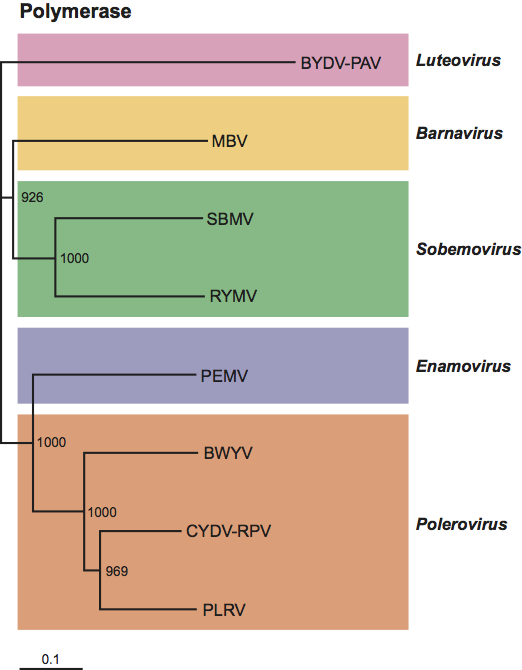
The amino acid sequences of the putative chymotrypsin-related serine protease and RdRp suggest an evolutionary relationship with some ssRNA positive sense plant viruses, particularly poleroviruses, sobemoviruses and enamoviruses (Figure 3).
Derivation of name
Barna: from bacilliform-shaped RNA viruses.
Further reading
Goodin et al., 1992 M.M. Goodin, B. Schlagnhaufer, C.P. Romaine, Encapsidation of the La France disease specific double-stranded RNAs in 36 nm isometric virus-like particles. Phytopathology. 82 (1992) 285–290.
Koonin et al., 2008 E.V. Koonin, Y.I. Wolf, K. Nagasaki, V.V. Dolja, The Big Bang of picorna-like virus evolution antedates the radiation of eukaryotic supergroups. Nat. Rev. Microbiol. 6 (2008) 925–939.
Revill et al., 1994 P.A. Revill, A.D. Davidson, P.J. Wright, The nucleotide sequence and genome organization of mushroom bacilliform virus: a single-stranded RNA virus of Agaricus bisporus (Lange) Imbach. Virology. 202 (1994) 904–911.
Revill et al., 1998 P.A. Revill, A.D. Davidson, P.J. Wright, Mushroom bacilliform virus RNA: the initiation of translation at the 5′-end of the genome and identification of the VPg. Virology. 249 (1998) 231–237.
Revill et al., 1999 P.A. Revill, A.D. Davidson, P.J. Wright, Identification of a subgenomic mRNA encoding the capsid protein of mushroom bacilliform virus, a single-stranded RNA mycovirus. Virology. 260 (1999) 273–276.
Romaine and Schlagnhaufer, 1991 C.P. Romaine, B. Schlagnhaufer, Hybridization analysis of the single-stranded RNA bacilliform virus associated with La France disease of Agaricus bisporus. Phytopathology. 81 (1991) 1336–1340.
Romaine and Schlagnhaufer, 1995 C.P. Romaine, B. Schlagnhaufer, PCR analysis of the viral complex associated with La France disease of Agaricus bisporus. Appl. Environ. Microbiol. 61 (1995) 2322–2325.
Tavantzis et al., 1980 S.M. Tavantzis, C.P. Romaine, S.H. Smith, Purification and partial characterization of a bacilliform virus from Agaricus bisporus: a single-stranded RNA mycovirus. Virology. 105 (1980) 94–102.
Tavantzis et al., 1983 S.M. Tavantzis, C.P. Romaine, S.H. Smith, Mechanism of genome expression in a single-stranded RNA virus from the cultivated mushroom Agaricus bisporus. Phytopathology. 106 (1983) 45–50.
Contributed by
Revill, P.A.
Figures
Figure 1 Negative contrast electron micrograph of particles of an isolate of Mushroom bacilliform virus. The bar represents 100 nm.
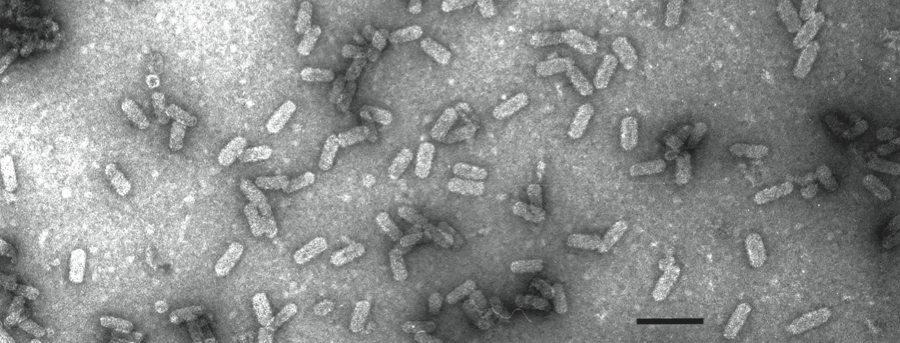
Figure 2 Genome organization of Mushroom bacilliform virus.
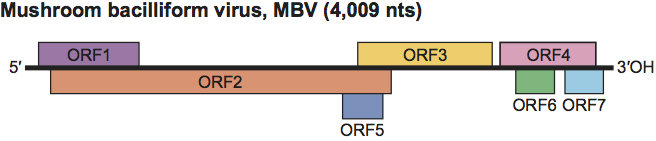
Figure 3 An unrooted neighbour-joining dendrogram (generated in Phylip) showing comparison of the aa sequence of the putative Mushroom bacilliform virus RdRp with those of selected sobemoviruses and poleroviruses, and an enamovirus. The luteovirus BYDV-PAV was included as an outgroup. Amino acid sequences were aligned with Clustal X and the tree was generated with Treeview. Bootstrap values (1000 replicates) are indicated. CYDV-RPV=cereal yellow dwarf virus-RPV; SBMV=southern bean mosaic virus; RYMV=rice yellow mottle virus; PEMV=pea enation mosaic virus-1; BWYV=beet western yellows virus; PLRV=potato leafroll virus. Accession numbers are AF218798 (BYDV-PAV), U07551 (MBV), AF055887 (SBMV), L20893 (RYMV), L04573 (PEMV) X13063 (BWYV), AF020090 (CYDV-RPV), AY138970 (PLRV).