Family: Kitaviridae
Pedro L. Ramos-González, Juliana Freitas-Astúa, Jun-Min Li, Antonio Tiberini, Avijit Roy, and Caixia Yang
The citation for this ICTV Report chapter is the summary to be published as Ramos-González et al. (2026): ICTV Virus Taxonomy Profile: Kitaviridae 2026, Journal of General Virology, (in press)
Corresponding author: Pedro L. Ramos-González ([email protected])
Edited by: Luisa Rubino and Evelien Adriaenssens
Posted: April 2026
Summary
The family Kitaviridae comprises plant-infecting viruses with segmented, positive-sense (+), single-stranded RNA genomes that are usually polyadenylated at the 3′-end. Members are classified into different genera based on their genome organization. Virions are bacilliform, ellipsoidal, or quasi-spherical; in some species, they are enveloped. Transmission occurs via Brevipalpus mites (for members of the genera Cilevirus and Higrevirus) and eriophyid mites (for some members of the genus Blunervirus). Infections are typically localized due to the lack of, or inefficient, systemic viral movement. Kitavirids cause economically important plant diseases, including citrus leprosis. Phylogenetically, they are closely related to unclassified arthropod-infecting viruses.
Table 1. Kitaviridae. Characteristics of members of the family Kitaviridae.
| Characteristic | Description |
| Example | Citrus leprosis virus C (RNA1: DQ352194, RNA2: DQ352195), species Cilevirus leprosis |
| Virion | Enveloped bacilliform particles measuring 25–70 nm in width and 100–120 nm in length or shorter bacilliform, ellipsoidal, or quasi-spherical virions |
| Genome | 2–5 segments of positive-sense, single-stranded RNA comprising 11 to 16 kb in total |
| Replication | Cytoplasmic, probably associated with the endoplasmic reticulum, involving the production of dsRNA intermediates |
| Translation | From genomic and sub-genomic RNAs |
| Host range | Plants. Cileviruses and higreviruses are transmitted by Brevipalpus mites; some blunerviruses are transmitted by eriophyid mites |
| Taxonomy | Realm Riboviria, kingdom Orthornavirae, phylum Kitrinoviricota, class Alsuviricetes, order Martellivirales; the family includes three genera and 27 species |
Virion
Morphology
Mature kitavirid particles exhibit bacilliform or ellipsoidal to quasi-spherical shapes and several cileviruses and at least one blunervirus are enveloped (Kitajima et al., 2003, Roy et al., 2013, Kitajima et al., 2022, Ramos-González et al., 2022) (Figure 1 Kitaviridae). Bacilliform virions of some members of the genus Cilevirus, i.e., Citrus leprosis virus C (CiLV-C, Cilevirus leprosis), Citrus leprosis virus C2 (CiLV-C2, Cilevirus colombiaense), passion fruit green spot virus (PfGSV, Cilevirus passiflorae), and Solanum violifolium ring spot virus (SvRSV, Cilevirus solani) measure 40–70 nm in width and 100–120 nm in length (Kitajima et al., 2003, Roy et al., 2013). Particles of the higrevirus Hibiscus green spot virus 2 (HGSV2, Higrevirus waimanalo) isolate WAI 1/1 are bacilliform with variable dimensions (33 to 120 nm in length and 14 to 70 nm in width) (Melzer et al., 2012, Olmedo-Velarde et al., 2024) and those of the blunervirus tomato fruit blotch virus (ToFBV, Blunervirus solani) are slender, short bacilliform of approximately 25 × 100 nm (Kitajima et al., 2022). Virion particles of Ligustrum leprosis virus (LigLV, Cilevirus australis) and Ligustrum chlorotic spot virus (LigCSV, Cilevirus ligustri) are quasi-spherical, measuring 40–53 nm in width and 55–65 nm in length (Ramos-González et al., 2022). In addition, HGSV2 isolate DMs01 (Pereira et al., 2025), Hibiscus yellow blotch virus (HYBV, Cilevirus oahuense) (Olmedo-Velarde et al., 2021), and tea plant necrotic ring blotch virus (TPNRBV, Blunervirus camelliae) (Hao et al., 2018), produce spherical virus-like particles (VLPs) with diameters of 55 nm, 66 nm, and 88 nm, respectively.
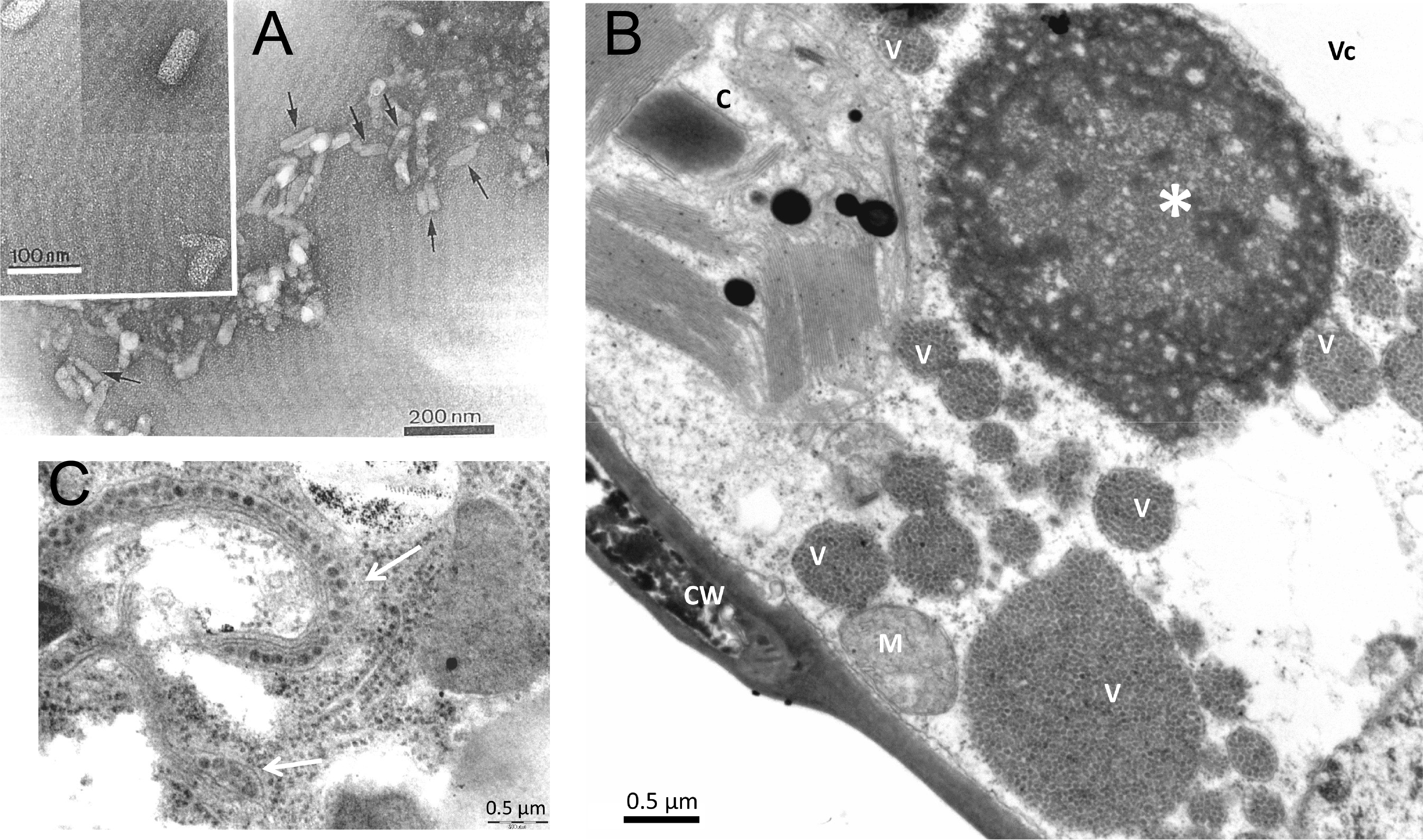 |
| Figure 1 Kitaviridae. (A) Electron micrograph of negatively-stained CiLV-C particles (arrows) in a partially purified preparation from Cleopatra mandarin (Citrus reshni). Insert: magnified particles showing outer surface structure. Courtesy of Dr. Addolorata Colariccio, Instituto Biológico, SP, Brazil. (B) Transmission electron micrograph of a soybean (Glycine max) leaf palisade parenchymal cell, experimentally infected by CiLV-C using viruliferous Brevipalpus mites. Note the large, vacuolated electron-dense viroplasm (*). Several pockets of the endoplasmic reticulum (V) filled with CiLV-C virions are present, scattered in the cytoplasm. A degraded chloroplast (C) is present next to the viroplasm. CW- cell wall; M- mitochondrion; Vc- vacuole. (C) CiLV-C virions (arrows), in the intercellular space between midgut epithelial cells of a viruliferous Brevipalpus yothersi mite (Acari: Tenuipalpidae), collected from an orange leaf with leprotic lesions. Micrographs B and C are courtesy of Dr. Elliot Watanabe Kitajima, Escola Superior de Agricultura Luiz de Queiroz (ESALQ), Universidade de São Paulo, SP, Brazil. |
Physicochemical and physical properties
In plant crude extracts, particles of the cileviruses CiLV-C (Lovisolo et al., 1996) and SvRSV (Ferreira et al., 2007) are inactivated after thermal treatments at 55−60 °C and 40 °C, respectively. CiLV-C-infected leaves stored at −80 °C retained infectivity for approximately four months, while those kept at −15 °C remained infectious for about one month (Lovisolo et al., 1996). Dry leaves can also retain virus infectivity (Lovisolo et al., 1996). The in vitro longevity of CiLV-C is 6 days at 4 °C and three days at room temperature (Lovisolo et al., 1996).
Nucleic acid
All kitavirids possess segmented, linear, positive-sense, single-stranded (ss) RNA genomes, but it remains unclear whether the segments are co-packaged or packaged individually. RNA segments are 3′-terminally polyadenylated for most members of the family.
Proteins
The composition of kitavirid particles has not yet been fully elucidated. In cileviruses, P29 protein takes part in their virions, likely as their coat protein. Immunogold labelling assays using anti-recombinant CiLV-C P29 antibodies detect short bacilliform particles of CiLV-C besides viroplasms in the infected plant cells (Calegario et al., 2013). Orthologs of P29 are restricted to members of the genus Cilevirus (Ramos-González et al., 2020, Olmedo-Velarde et al., 2021, Ramos-González et al., 2022). Transiently expressed P29 proteins of CiLV-C in Nicotiana benthamiana plants form stable icosahedral VLPs with a diameter of 15.8 ± 1.3 nm, a morphology that diverges from that of CiLV-C virions (Ortega-Rivera et al., 2023). The P24 protein (also termed SP24), a membrane protein (Pfam16504) present in all kitavirids, and the putative glycoprotein of 61−65 kDa in cileviruses and higreviruses, are likely virion structural proteins. A protein band of approximately 25 kDa, of unknown identity, but likely corresponding to P24 or P29 proteins, has been detected in semi-purified virus extracts obtained from citrus leprosis-affected tissues (Lovisolo et al., 1996).
Lipids
Kitavirids of the genera Cilevirus and Blunervirus are enveloped in a lipid membrane (Kitajima et al., 2003, Roy et al., 2013, Kitajima et al., 2022, Ramos-González et al., 2022). No information is available for members of the genus Higrevirus.
Genome organization and replication
Kitavirid genomes are split into two to five segments, each of which is generally multicistronic (Figure 2 Kitaviridae). Kitavirids, particularly blunerviruses, are regarded as natural chimeric systems composed of RNA fragments of diverse origin (Morozov and Solovyev 2015). Kitavirid genomes have a minimum of five ORFs, including those encoding an RNA-directed RNA polymerase (RdRP), movement protein (MP), and likely virion structural proteins, such as P29 in cileviruses, and orthologs of the P24 membrane protein (Pfam16504), the latter present in all kitavirids (Ramos-González et al., 2023). In general, genomic RNAs of kitavirids have a poly-A tail at their 3′-end. The expression of the downstream ORFs, at least in cileviruses, is based on the formation of a nested set of 3′-co-terminal sub-genomic RNAs (Pascon et al., 2006, Roy et al., 2013, Ramos-González et al., 2020). In light of the presence of methyltransferase domains in RdRP of kitavirids, the existence of an RNA cap structure at the 5′-end of the viral RNAs is presumed. Double-stranded RNAs, likely to be the intermediary replication molecules, have been detected in plants infected with CiLV-C, SvRSV, HGSV2, and blueberry necrotic ring blotch virus (BNRBV, Blunervirus vaccinii) (Colariccio et al., 2000, Ferreira et al., 2007, Melzer et al., 2012, Quito-Avila et al., 2013). Replication and assembly of kitavirid particles occur in the cytoplasm, and for most species, viral particles accumulate within the lumen of the endoplasmic reticulum (ER) or the perinuclear space (Kitajima et al., 2003, Hao et al., 2018, Kitajima et al., 2022, Ramos-González et al., 2022). Infected plant cells typically contain amorphous, variable-sized electron-dense viroplasms, which may be vacuolated or not (Colariccio et al., 1995, Ramos-González et al., 2022). In cilevirus-infected cells, the budding-like process takes place at the ER, adjacent to viroplasms (Kitajima et al., 2003).
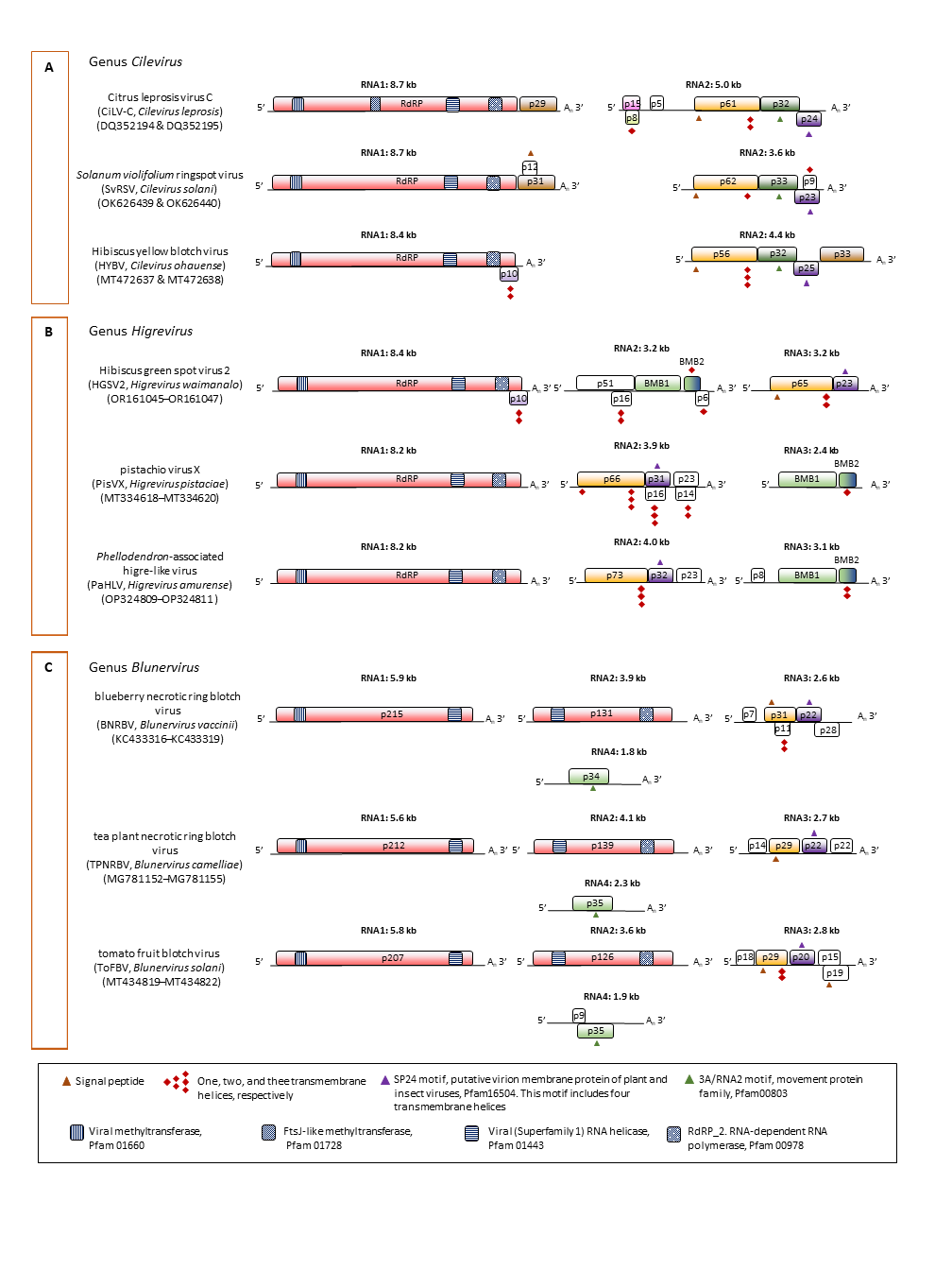 |
| Figure 2 Kitaviridae. Genomic organization of representative members of the family Kitaviridae. A: Genus Cilevirus, B: Genus Higrevirus, and C: Genus Blunervirus. Boxes indicate open reading frames (ORFs); colours denote putative conserved functional or structural relationships across the family. In Citrus leprosis virus C (Cilevirus), ORFs encode RdRP (RNA-directed RNA polymerase), P29 (coat protein), P15 (putative RNA silencing suppressor), P61 (putative glycoprotein, likely a structural protein of the virion), P32 (MP, movement protein), and P24 (putative structural protein of the virion). The RNA2 ORFs p8 and p5 were previously unrecognized. In Hibiscus green spot virus 2 (HGSV2), (Higrevirus), BMB denotes the binary movement block. ORF p16 was previously overlooked (Pereira et al., 2025). Isolates of HGSV2 with different genomic organizations have also been reported (Melzer et al., 2012, Olmedo-Velarde et al., 2024). In blueberry necrotic ring blotch virus (Blunervirus), the RNA3 ORF p11 was previously overlooked (Cantu-Iris et al., 2013); some isolates lack poly-A tails or show minor genomic variation (Quito-Avila et al., 2013). White boxes represent ORFs with unknown functions or without identifiable orthologs. Predicted protein features (signal peptides, transmembrane domains and conserved motifs/domains) are indicated by symbols defined in the legend. |
Biology
Kitavirids are present on six continents, but their distribution is highly uneven. Most kitavirids have been detected in the Americas and Asia, whereas viruses from only one to three species have been identified in Europe, Africa, Australia, and Oceania. Kitavirids naturally infect mono- and dicotyledonous plants, including herbaceous and woody plants comprising weeds, ornamentals, and crops (Ramos-González et al., 2023). Experimentally, several kitavirids have been transmitted to Arabidopsis thaliana plants (Arena et al., 2017). These viruses have limited or no systemic movement in all natural and experimental host plants identified so far. Their infections cause chlorotic to necrotic lesions in which most viruses remain restricted. Slow or inefficient systemic movements of some kitavirids have been described (Ren et al., 2022, Tassi et al., 2025). Cell-to-cell movement of cileviruses and blunerviruses involves 30K movement proteins (Leastro et al., 2021), whereas higreviruses employ a specialized transport mechanism known as the binary movement block (Lazareva et al., 2017a, Lazareva et al., 2017b). In infected cells, virions accumulate in the cytoplasm, although the extent of cytopathic effects can vary among viruses. In citrus leaves infected with CiLV-C, for example, some mesophyll parenchyma cells within lesions undergo marked hypertrophy and hyperplasia, while others, located at lesion margins or scattered within halo regions, experience plasmolysis (Marques et al., 2010). Plant-to-plant transmission of kitavirids is mediated by mites in a persistent manner (Tassi et al., 2017, Bertin et al., 2025, Tassi et al., 2025). Several species of mites from the genus Brevipalpus (Tenuipalpidae, Trombidiformes) are involved in the transmission of cileviruses and higreviruses (de Lillo et al., 2021). For certain viruses, transmission can be mediated by more than one Brevipalpus mite species (Ramos-González et al., 2016, Nunes M.A. et al., 2018, Olmedo-Velarde et al., 2024, Pereira et al., 2025). Virions of cileviruses have been detected in intercellular spaces of the epithelial tissues in mites (Tassi et al., 2022), and CiLV-C and CiLV-C2 seem to propagate in Brevipalpus mites (Roy et al., 2015, Ramos-González et al., 2023). The blunerviruses BNRBV and ToFBV are transmitted, respectively, by eriophyids (Eriophyidae, Trombidiformes) of the species Calacarus corymbosi and Aculops lycopersici (Bertin et al., 2025, Tassi et al., 2025).
Antigenicity
The antigenic properties of kitavirids have been scarcely studied. No serological cross-reactions have been detected between CiLV-C and CiLV-C2 when using polyclonal or monoclonal antibodies generated against recombinant P29 proteins (Calegario et al., 2013, Choudhary et al., 2014, Choudhary et al., 2017).
Derivation of names
Kitaviridae: to honour the Brazilian plant virologist, Elliot Watanabe Kitajima [Academia Brasileira de Ciências: https://www.abc.org.br/membro/elliot-watanabe-kitajima/ ], a pioneer in the discovery and characterization of viruses in the family; the suffix -viridae for family taxa.
Blunervirus: from blueberry necrotic ring blotch virus
Cilevirus: from Citrus leprosis virus C
Higrevirus: from Hibiscus green spot virus 2
Genus demarcation criteria
Genera within the family Kitaviridae are distinguished by the number of genomic segments and the organization and synteny of ORFs across segments of their members, which form monophyletic clades based on the phylogenetic analysis of the deduced amino acid sequence of the RdRP, particularly the RdRP2 domain (Pfam00978).
Relationships within the family
Phylogenetic relationships among kitavirids have been inferred from a maximum likelihood tree generated from phylogenetically informative sequences in their RdRP proteins (Figure 3 Kitaviridae). These relationships can be identified by aligning the RdRP proteins (their RdRP2 domains, Pfam00978, or concatenating proteins involved in the replication process in blunerviruses) using MAFFT with the E-INS-I iterative refinement method, and removing the non-informative, ambiguous sites using alignment trimming tools such as BMGE (Criscuolo and Gribaldo 2010), and Trimal (Capella-Gutiérrez et al., 2009) software. All kitavirids encode orthologs of p24, first identified in CiLV-C, which encodes a putative virion membrane protein containing the conserved domain SP24 (Pfam16504) (Kuchibhatla et al., 2014).
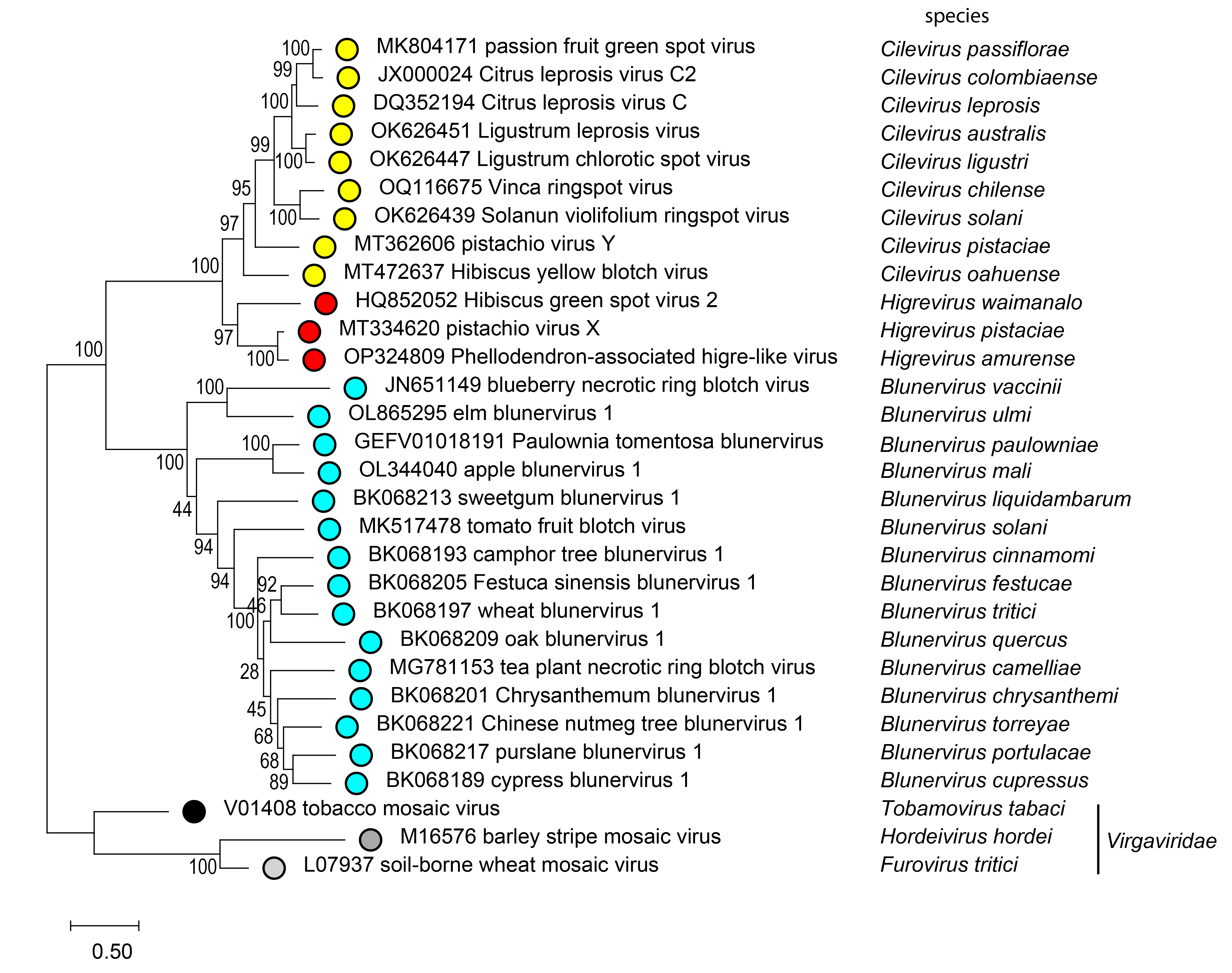 |
| Figure 3 Kitaviridae. Phylogenetic reconstruction for viruses of the family Kitaviridae. The midpoint-rooted maximum-likelihood phylogenetic tree was inferred using IQ-tree software based on the deduced amino acid sequences of RNA-directed (RdRP) proteins. The alignment was constructed in MAFFT using the E-INS-I iterative refinement method. Phylogenetically informative regions from the multiple sequence alignment included 701 residues selected using BMGE software, and the evolutionary history was inferred based on the model LG+F+I+G4. The bootstrap support values (1,000 replications) of branches greater than 50% are indicated next to the corresponding nodes. The scale bar specifies the average number of amino acid substitutions per site, and the tree is drawn to scale. |
Relationships with other taxa
Kitavirids are thought to have arisen from ancestors shared with arthropod-infecting viruses, e.g., of the unclassified taxa negevirus, representing a compelling case of inter-kingdom horizontal virus transfer (Dolja et al., 2020, Ramos-González et al., 2020, Lu et al., 2024). Kitavirids and several groups of unclassified arthropod-infecting viruses are closely related in phylogenetic studies using the RdRP and SP24 protein sequences (Kuchibhatla et al., 2014).

